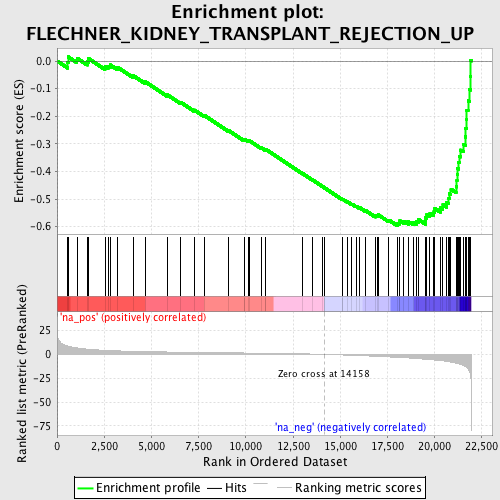
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | FLECHNER_KIDNEY_TRANSPLANT_REJECTION_UP |
| Enrichment Score (ES) | -0.5974916 |
| Normalized Enrichment Score (NES) | -2.2433555 |
| Nominal p-value | 0.0 |
| FDR q-value | 5.6489627E-4 |
| FWER p-Value | 0.0040 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | C1QB | 560 | 8.626 | -0.0045 | No | ||
| 2 | GZMA | 588 | 8.487 | 0.0151 | No | ||
| 3 | FCGR3A | 1053 | 6.726 | 0.0104 | No | ||
| 4 | FCER1G | 1598 | 5.415 | -0.0012 | No | ||
| 5 | ADAM15 | 1674 | 5.286 | 0.0083 | No | ||
| 6 | RAB31 | 2538 | 4.163 | -0.0209 | No | ||
| 7 | EIF2AK2 | 2700 | 4.017 | -0.0184 | No | ||
| 8 | PSMB8 | 2816 | 3.937 | -0.0140 | No | ||
| 9 | DARC | 3196 | 3.659 | -0.0224 | No | ||
| 10 | LGALS9 | 4022 | 3.131 | -0.0524 | No | ||
| 11 | CD14 | 4674 | 2.846 | -0.0752 | No | ||
| 12 | USP9X | 5833 | 2.453 | -0.1221 | No | ||
| 13 | APOC1 | 6560 | 2.234 | -0.1499 | No | ||
| 14 | AOAH | 7272 | 2.044 | -0.1773 | No | ||
| 15 | ECGF1 | 7822 | 1.901 | -0.1978 | No | ||
| 16 | ALDOC | 9076 | 1.605 | -0.2511 | No | ||
| 17 | TYROBP | 9910 | 1.408 | -0.2858 | No | ||
| 18 | VSIG4 | 9917 | 1.407 | -0.2826 | No | ||
| 19 | IFI30 | 10142 | 1.349 | -0.2895 | No | ||
| 20 | S100A4 | 10186 | 1.337 | -0.2882 | No | ||
| 21 | LCP1 | 10823 | 1.181 | -0.3144 | No | ||
| 22 | LY86 | 11019 | 1.130 | -0.3205 | No | ||
| 23 | CD163 | 11063 | 1.117 | -0.3198 | No | ||
| 24 | MOXD1 | 13022 | 0.506 | -0.4080 | No | ||
| 25 | IFITM1 | 13553 | 0.295 | -0.4316 | No | ||
| 26 | CFD | 14060 | 0.046 | -0.4546 | No | ||
| 27 | LST1 | 14171 | -0.014 | -0.4596 | No | ||
| 28 | SFN | 14174 | -0.015 | -0.4596 | No | ||
| 29 | CD2 | 15104 | -0.616 | -0.5006 | No | ||
| 30 | LCN2 | 15126 | -0.632 | -0.5000 | No | ||
| 31 | WARS | 15375 | -0.788 | -0.5094 | No | ||
| 32 | UBE2L6 | 15616 | -0.953 | -0.5180 | No | ||
| 33 | CD48 | 15841 | -1.116 | -0.5255 | No | ||
| 34 | CSTA | 16006 | -1.247 | -0.5300 | No | ||
| 35 | AIF1 | 16322 | -1.506 | -0.5407 | No | ||
| 36 | RAC2 | 16856 | -1.992 | -0.5602 | No | ||
| 37 | PDE6D | 16951 | -2.075 | -0.5594 | No | ||
| 38 | HCLS1 | 17013 | -2.142 | -0.5569 | No | ||
| 39 | MMP9 | 17583 | -2.678 | -0.5764 | No | ||
| 40 | IL10RB | 18046 | -3.152 | -0.5898 | Yes | ||
| 41 | S100A8 | 18123 | -3.243 | -0.5853 | Yes | ||
| 42 | RAB27A | 18166 | -3.290 | -0.5791 | Yes | ||
| 43 | TNFRSF1B | 18378 | -3.515 | -0.5802 | Yes | ||
| 44 | IL10RA | 18621 | -3.773 | -0.5820 | Yes | ||
| 45 | CXCR4 | 18871 | -4.113 | -0.5833 | Yes | ||
| 46 | RGS10 | 19042 | -4.334 | -0.5804 | Yes | ||
| 47 | UBC | 19151 | -4.504 | -0.5743 | Yes | ||
| 48 | PTPRC | 19517 | -5.062 | -0.5786 | Yes | ||
| 49 | GBP2 | 19528 | -5.087 | -0.5666 | Yes | ||
| 50 | ACTB | 19594 | -5.181 | -0.5569 | Yes | ||
| 51 | ISG20 | 19755 | -5.412 | -0.5509 | Yes | ||
| 52 | CD3D | 19969 | -5.819 | -0.5464 | Yes | ||
| 53 | TNFRSF7 | 20015 | -5.900 | -0.5340 | Yes | ||
| 54 | ITGB2 | 20306 | -6.502 | -0.5313 | Yes | ||
| 55 | PIM2 | 20446 | -6.848 | -0.5208 | Yes | ||
| 56 | RHOH | 20639 | -7.326 | -0.5117 | Yes | ||
| 57 | INPP5D | 20758 | -7.665 | -0.4983 | Yes | ||
| 58 | POLR2A | 20787 | -7.778 | -0.4805 | Yes | ||
| 59 | CUGBP2 | 20871 | -8.070 | -0.4645 | Yes | ||
| 60 | GMFG | 21143 | -9.118 | -0.4545 | Yes | ||
| 61 | CD8A | 21185 | -9.290 | -0.4336 | Yes | ||
| 62 | GLUL | 21196 | -9.342 | -0.4112 | Yes | ||
| 63 | IL4R | 21228 | -9.529 | -0.3892 | Yes | ||
| 64 | SLA | 21278 | -9.796 | -0.3674 | Yes | ||
| 65 | CCL5 | 21305 | -9.930 | -0.3443 | Yes | ||
| 66 | DOCK2 | 21381 | -10.369 | -0.3223 | Yes | ||
| 67 | RASSF2 | 21521 | -11.453 | -0.3006 | Yes | ||
| 68 | CORO1A | 21631 | -12.679 | -0.2745 | Yes | ||
| 69 | STAT1 | 21656 | -13.091 | -0.2435 | Yes | ||
| 70 | CD53 | 21704 | -13.844 | -0.2117 | Yes | ||
| 71 | LAPTM5 | 21712 | -14.002 | -0.1777 | Yes | ||
| 72 | LCK | 21778 | -15.166 | -0.1434 | Yes | ||
| 73 | PRKCB1 | 21866 | -17.985 | -0.1033 | Yes | ||
| 74 | CD52 | 21908 | -20.783 | -0.0542 | Yes | ||
| 75 | CCND3 | 21924 | -22.833 | 0.0011 | Yes |
