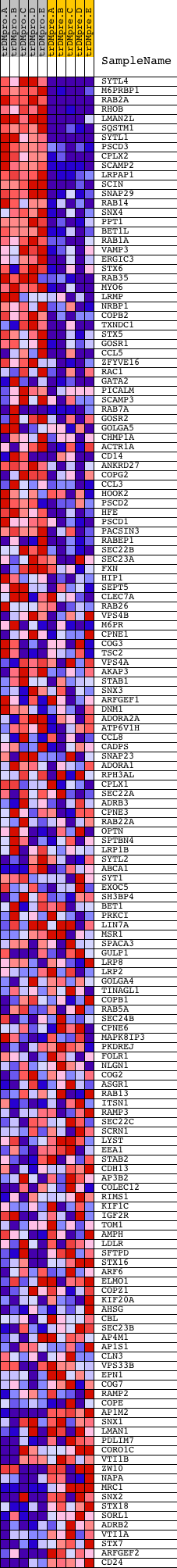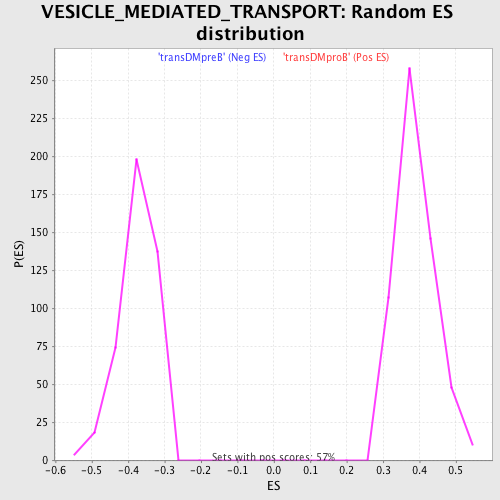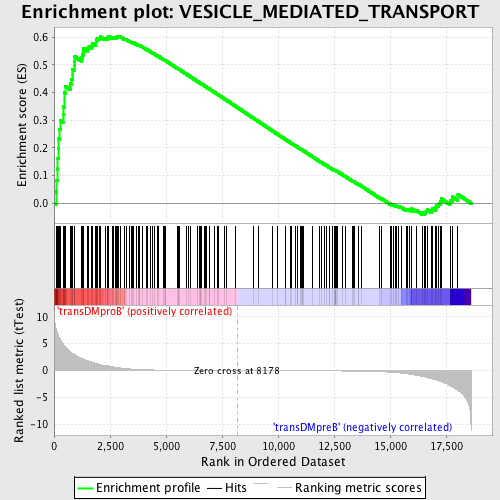
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | VESICLE_MEDIATED_TRANSPORT |
| Enrichment Score (ES) | 0.6035487 |
| Normalized Enrichment Score (NES) | 1.5487984 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.29611704 |
| FWER p-Value | 0.961 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SYTL4 | 24067 | 86 | 7.816 | 0.0410 | Yes | ||
| 2 | M6PRBP1 | 22930 | 118 | 7.404 | 0.0825 | Yes | ||
| 3 | RAB2A | 16279 | 137 | 7.207 | 0.1237 | Yes | ||
| 4 | RHOB | 4411 | 168 | 6.852 | 0.1620 | Yes | ||
| 5 | LMAN2L | 5670 10070 5669 | 205 | 6.304 | 0.1969 | Yes | ||
| 6 | SQSTM1 | 9517 | 213 | 6.243 | 0.2330 | Yes | ||
| 7 | SYTL1 | 15728 | 242 | 6.022 | 0.2666 | Yes | ||
| 8 | PSCD3 | 16633 | 276 | 5.842 | 0.2990 | Yes | ||
| 9 | CPLX2 | 8779 | 399 | 4.954 | 0.3213 | Yes | ||
| 10 | SCAMP2 | 19430 | 423 | 4.810 | 0.3481 | Yes | ||
| 11 | LRPAP1 | 5005 | 470 | 4.556 | 0.3722 | Yes | ||
| 12 | SCIN | 21079 | 471 | 4.551 | 0.3988 | Yes | ||
| 13 | SNAP29 | 22837 | 500 | 4.425 | 0.4231 | Yes | ||
| 14 | RAB14 | 12685 | 726 | 3.475 | 0.4312 | Yes | ||
| 15 | SNX4 | 12 | 785 | 3.275 | 0.4472 | Yes | ||
| 16 | PPT1 | 9613 | 827 | 3.178 | 0.4635 | Yes | ||
| 17 | BET1L | 17570 | 829 | 3.174 | 0.4820 | Yes | ||
| 18 | RAB1A | 20942 | 893 | 3.022 | 0.4963 | Yes | ||
| 19 | VAMP3 | 15660 | 898 | 3.007 | 0.5136 | Yes | ||
| 20 | ERGIC3 | 14770 12336 | 925 | 2.943 | 0.5294 | Yes | ||
| 21 | STX6 | 14091 12213 | 1235 | 2.237 | 0.5257 | Yes | ||
| 22 | RAB35 | 13386 | 1274 | 2.173 | 0.5363 | Yes | ||
| 23 | MYO6 | 19368 | 1300 | 2.140 | 0.5475 | Yes | ||
| 24 | LRMP | 17244 | 1319 | 2.100 | 0.5588 | Yes | ||
| 25 | NRBP1 | 9656 | 1504 | 1.807 | 0.5594 | Yes | ||
| 26 | COPB2 | 19346 | 1554 | 1.742 | 0.5669 | Yes | ||
| 27 | TXNDC1 | 13067 | 1688 | 1.580 | 0.5689 | Yes | ||
| 28 | STX5 | 23942 | 1724 | 1.532 | 0.5759 | Yes | ||
| 29 | GOSR1 | 20345 | 1849 | 1.349 | 0.5771 | Yes | ||
| 30 | CCL5 | 20320 | 1869 | 1.327 | 0.5838 | Yes | ||
| 31 | ZFYVE16 | 21395 | 1891 | 1.307 | 0.5903 | Yes | ||
| 32 | RAC1 | 16302 | 1922 | 1.265 | 0.5961 | Yes | ||
| 33 | GATA2 | 17370 | 2040 | 1.121 | 0.5963 | Yes | ||
| 34 | PICALM | 18191 | 2059 | 1.107 | 0.6018 | Yes | ||
| 35 | SCAMP3 | 15543 | 2277 | 0.958 | 0.5956 | Yes | ||
| 36 | RAB7A | 9684 5347 9683 | 2380 | 0.862 | 0.5951 | Yes | ||
| 37 | GOSR2 | 20194 1372 1360 | 2393 | 0.856 | 0.5995 | Yes | ||
| 38 | GOLGA5 | 21179 | 2410 | 0.847 | 0.6035 | Yes | ||
| 39 | CHMP1A | 10634 | 2589 | 0.706 | 0.5980 | No | ||
| 40 | ACTR1A | 23653 | 2654 | 0.665 | 0.5984 | No | ||
| 41 | CD14 | 23452 | 2751 | 0.590 | 0.5967 | No | ||
| 42 | ANKRD27 | 18290 10893 6368 | 2789 | 0.558 | 0.5979 | No | ||
| 43 | COPG2 | 17193 | 2804 | 0.552 | 0.6004 | No | ||
| 44 | CCL3 | 20317 | 2845 | 0.528 | 0.6013 | No | ||
| 45 | HOOK2 | 18541 | 2888 | 0.508 | 0.6020 | No | ||
| 46 | PSCD2 | 17826 | 2952 | 0.475 | 0.6014 | No | ||
| 47 | HFE | 21519 | 3158 | 0.379 | 0.5925 | No | ||
| 48 | PSCD1 | 9629 5296 | 3212 | 0.357 | 0.5917 | No | ||
| 49 | PACSIN3 | 14948 8149 2776 2665 | 3386 | 0.294 | 0.5840 | No | ||
| 50 | RABEP1 | 12016 1344 | 3471 | 0.268 | 0.5811 | No | ||
| 51 | SEC22B | 15486 | 3519 | 0.253 | 0.5800 | No | ||
| 52 | SEC23A | 5420 | 3554 | 0.244 | 0.5796 | No | ||
| 53 | FXN | 23709 | 3690 | 0.214 | 0.5735 | No | ||
| 54 | HIP1 | 5676 | 3764 | 0.197 | 0.5707 | No | ||
| 55 | SEPT5 | 9598 | 3805 | 0.190 | 0.5696 | No | ||
| 56 | CLEC7A | 7142 16977 12157 1111 | 3807 | 0.190 | 0.5707 | No | ||
| 57 | RAB26 | 11609 | 3947 | 0.162 | 0.5641 | No | ||
| 58 | VPS4B | 5446 | 4126 | 0.133 | 0.5552 | No | ||
| 59 | M6PR | 1053 5044 9328 | 4159 | 0.129 | 0.5543 | No | ||
| 60 | CPNE1 | 11186 | 4320 | 0.109 | 0.5462 | No | ||
| 61 | COG3 | 21751 | 4398 | 0.101 | 0.5426 | No | ||
| 62 | TSC2 | 23090 | 4467 | 0.095 | 0.5395 | No | ||
| 63 | VPS4A | 18475 | 4622 | 0.083 | 0.5317 | No | ||
| 64 | AKAP3 | 8566 | 4641 | 0.082 | 0.5312 | No | ||
| 65 | STAB1 | 21892 | 4867 | 0.069 | 0.5194 | No | ||
| 66 | SNX3 | 20044 3416 | 4937 | 0.066 | 0.5160 | No | ||
| 67 | ARFGEF1 | 9995 | 4952 | 0.065 | 0.5156 | No | ||
| 68 | DNM1 | 2777 8857 | 4970 | 0.064 | 0.5151 | No | ||
| 69 | ADORA2A | 8557 | 5510 | 0.046 | 0.4862 | No | ||
| 70 | ATP6V1H | 14301 | 5552 | 0.045 | 0.4842 | No | ||
| 71 | CCL8 | 20740 | 5559 | 0.045 | 0.4841 | No | ||
| 72 | CADPS | 11298 | 5603 | 0.044 | 0.4821 | No | ||
| 73 | SNAP23 | 2853 14893 | 5888 | 0.037 | 0.4669 | No | ||
| 74 | ADORA1 | 13831 | 6013 | 0.034 | 0.4604 | No | ||
| 75 | RPH3AL | 11725 | 6068 | 0.033 | 0.4576 | No | ||
| 76 | CPLX1 | 16447 | 6411 | 0.026 | 0.4393 | No | ||
| 77 | SEC22A | 1710 1731 1728 22605 | 6498 | 0.025 | 0.4348 | No | ||
| 78 | ADRB3 | 18901 | 6518 | 0.024 | 0.4339 | No | ||
| 79 | CPNE3 | 7689 | 6549 | 0.024 | 0.4324 | No | ||
| 80 | RAB22A | 2888 5343 | 6573 | 0.023 | 0.4313 | No | ||
| 81 | OPTN | 14693 | 6722 | 0.021 | 0.4234 | No | ||
| 82 | SPTBN4 | 13471 1104 | 6776 | 0.020 | 0.4206 | No | ||
| 83 | LRP1B | 8249 | 6794 | 0.020 | 0.4198 | No | ||
| 84 | SYTL2 | 18190 1539 3730 | 6951 | 0.017 | 0.4115 | No | ||
| 85 | ABCA1 | 15881 | 6956 | 0.017 | 0.4114 | No | ||
| 86 | SYT1 | 5565 | 7160 | 0.014 | 0.4004 | No | ||
| 87 | EXOC5 | 8370 | 7299 | 0.012 | 0.3930 | No | ||
| 88 | SH3BP4 | 8270 | 7331 | 0.012 | 0.3914 | No | ||
| 89 | BET1 | 17226 | 7622 | 0.008 | 0.3758 | No | ||
| 90 | PRKCI | 9576 | 7682 | 0.007 | 0.3726 | No | ||
| 91 | LIN7A | 8424 4230 | 8091 | 0.001 | 0.3505 | No | ||
| 92 | MSR1 | 5417 | 8921 | -0.009 | 0.3056 | No | ||
| 93 | SPACA3 | 13680 | 9111 | -0.012 | 0.2955 | No | ||
| 94 | GULP1 | 14254 3960 3932 | 9765 | -0.020 | 0.2602 | No | ||
| 95 | LRP8 | 16149 2425 | 9967 | -0.023 | 0.2495 | No | ||
| 96 | LRP2 | 14566 14567 | 10308 | -0.027 | 0.2312 | No | ||
| 97 | GOLGA4 | 12022 | 10330 | -0.028 | 0.2302 | No | ||
| 98 | TINAGL1 | 13587 | 10532 | -0.031 | 0.2195 | No | ||
| 99 | COPB1 | 17670 | 10612 | -0.032 | 0.2154 | No | ||
| 100 | RAB5A | 11310 | 10797 | -0.035 | 0.2057 | No | ||
| 101 | SEC24B | 13622 1910 | 10842 | -0.036 | 0.2035 | No | ||
| 102 | CPNE6 | 22010 | 10883 | -0.037 | 0.2015 | No | ||
| 103 | MAPK8IP3 | 11402 23080 | 11012 | -0.039 | 0.1948 | No | ||
| 104 | PKDREJ | 22170 | 11052 | -0.040 | 0.1930 | No | ||
| 105 | FOLR1 | 17731 | 11068 | -0.040 | 0.1924 | No | ||
| 106 | NLGN1 | 15368 | 11081 | -0.040 | 0.1920 | No | ||
| 107 | COG2 | 18423 | 11123 | -0.041 | 0.1900 | No | ||
| 108 | ASGR1 | 20812 | 11541 | -0.050 | 0.1677 | No | ||
| 109 | RAB13 | 15535 | 11854 | -0.057 | 0.1511 | No | ||
| 110 | ITSN1 | 9196 | 11918 | -0.059 | 0.1480 | No | ||
| 111 | RAMP3 | 20949 | 12062 | -0.063 | 0.1407 | No | ||
| 112 | SEC22C | 10084 5680 | 12168 | -0.066 | 0.1354 | No | ||
| 113 | SCRN1 | 7649 12840 | 12273 | -0.069 | 0.1301 | No | ||
| 114 | LYST | 9325 5040 | 12289 | -0.069 | 0.1297 | No | ||
| 115 | EEA1 | 5695 10097 | 12414 | -0.073 | 0.1234 | No | ||
| 116 | STAB2 | 19656 | 12508 | -0.076 | 0.1188 | No | ||
| 117 | CDH13 | 4506 3826 | 12518 | -0.077 | 0.1188 | No | ||
| 118 | AP3B2 | 4394 | 12548 | -0.078 | 0.1177 | No | ||
| 119 | COLEC12 | 23624 8946 | 12550 | -0.078 | 0.1181 | No | ||
| 120 | RIMS1 | 8575 | 12570 | -0.078 | 0.1175 | No | ||
| 121 | KIF1C | 4951 | 12588 | -0.079 | 0.1170 | No | ||
| 122 | IGF2R | 23123 | 12662 | -0.082 | 0.1136 | No | ||
| 123 | TOM1 | 3817 18566 | 12879 | -0.090 | 0.1024 | No | ||
| 124 | AMPH | 3256 21700 | 13004 | -0.095 | 0.0962 | No | ||
| 125 | LDLR | 4990 | 13326 | -0.111 | 0.0795 | No | ||
| 126 | SFTPD | 21867 | 13350 | -0.112 | 0.0789 | No | ||
| 127 | STX16 | 10464 | 13392 | -0.115 | 0.0773 | No | ||
| 128 | ARF6 | 21252 | 13600 | -0.129 | 0.0669 | No | ||
| 129 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 13606 | -0.130 | 0.0674 | No | ||
| 130 | COPZ1 | 22338 | 13743 | -0.141 | 0.0608 | No | ||
| 131 | KIF20A | 23602 | 14535 | -0.230 | 0.0193 | No | ||
| 132 | AHSG | 22812 | 14630 | -0.245 | 0.0157 | No | ||
| 133 | CBL | 19154 | 15001 | -0.315 | -0.0026 | No | ||
| 134 | SEC23B | 14821 2709 | 15079 | -0.342 | -0.0047 | No | ||
| 135 | AP4M1 | 3499 16656 | 15134 | -0.359 | -0.0056 | No | ||
| 136 | AP1S1 | 3500 3453 16335 | 15247 | -0.392 | -0.0093 | No | ||
| 137 | CLN3 | 17635 | 15299 | -0.410 | -0.0097 | No | ||
| 138 | VPS33B | 18204 | 15383 | -0.436 | -0.0117 | No | ||
| 139 | EPN1 | 18401 | 15486 | -0.476 | -0.0144 | No | ||
| 140 | COG7 | 10605 | 15731 | -0.595 | -0.0242 | No | ||
| 141 | RAMP2 | 20660 | 15761 | -0.611 | -0.0222 | No | ||
| 142 | COPE | 12228 7204 12229 | 15868 | -0.673 | -0.0240 | No | ||
| 143 | AP1M2 | 19209 8600 4393 | 15963 | -0.736 | -0.0248 | No | ||
| 144 | SNX1 | 19076 | 15966 | -0.740 | -0.0205 | No | ||
| 145 | LMAN1 | 12867 | 16157 | -0.887 | -0.0257 | No | ||
| 146 | PDLIM7 | 7434 | 16436 | -1.078 | -0.0344 | No | ||
| 147 | CORO1C | 16425 | 16551 | -1.200 | -0.0336 | No | ||
| 148 | VTI1B | 21032 957 | 16600 | -1.241 | -0.0289 | No | ||
| 149 | ZW10 | 3114 19464 | 16651 | -1.297 | -0.0241 | No | ||
| 150 | NAPA | 2150 18386 | 16853 | -1.553 | -0.0259 | No | ||
| 151 | MRC1 | 15116 | 16879 | -1.582 | -0.0180 | No | ||
| 152 | SNX2 | 1974 23554 12585 | 17020 | -1.722 | -0.0155 | No | ||
| 153 | STX18 | 16857 | 17067 | -1.775 | -0.0077 | No | ||
| 154 | SORL1 | 5474 | 17178 | -1.917 | -0.0024 | No | ||
| 155 | ADRB2 | 23422 | 17243 | -2.032 | 0.0060 | No | ||
| 156 | VTI1A | 11994 7009 12908 | 17276 | -2.077 | 0.0164 | No | ||
| 157 | STX7 | 20070 | 17673 | -2.805 | 0.0113 | No | ||
| 158 | ARFGEF2 | 14728 | 17794 | -3.073 | 0.0227 | No | ||
| 159 | CD24 | 8711 | 18026 | -3.725 | 0.0320 | No |