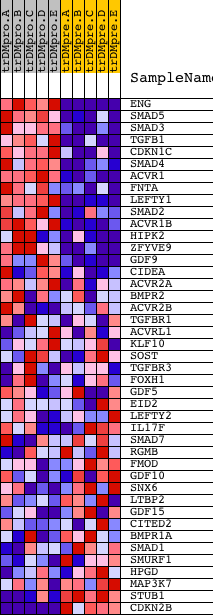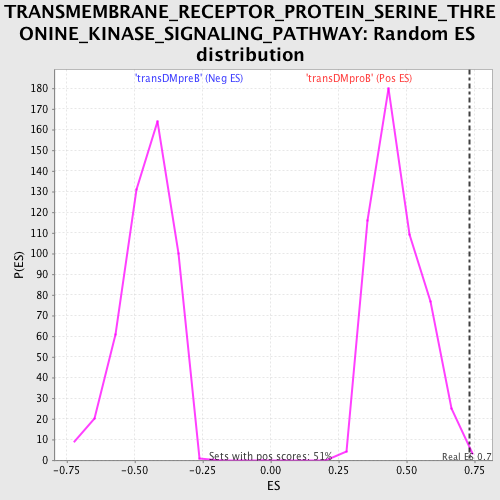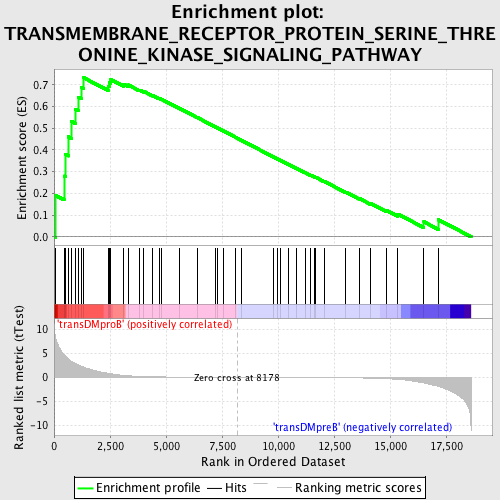
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | TRANSMEMBRANE_RECEPTOR_PROTEIN_SERINE_THREONINE_KINASE_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.7335013 |
| Normalized Enrichment Score (NES) | 1.5728809 |
| Nominal p-value | 0.0019455253 |
| FDR q-value | 0.2772625 |
| FWER p-Value | 0.866 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ENG | 4667 | 56 | 8.263 | 0.1914 | Yes | ||
| 2 | SMAD5 | 21621 | 447 | 4.668 | 0.2802 | Yes | ||
| 3 | SMAD3 | 19084 | 511 | 4.367 | 0.3795 | Yes | ||
| 4 | TGFB1 | 18332 | 647 | 3.751 | 0.4605 | Yes | ||
| 5 | CDKN1C | 17546 | 779 | 3.287 | 0.5307 | Yes | ||
| 6 | SMAD4 | 5058 | 952 | 2.856 | 0.5887 | Yes | ||
| 7 | ACVR1 | 4334 | 1076 | 2.564 | 0.6424 | Yes | ||
| 8 | FNTA | 18904 | 1211 | 2.275 | 0.6887 | Yes | ||
| 9 | LEFTY1 | 8877 | 1307 | 2.123 | 0.7335 | Yes | ||
| 10 | SMAD2 | 23511 | 2417 | 0.844 | 0.6937 | No | ||
| 11 | ACVR1B | 4335 22353 | 2478 | 0.793 | 0.7091 | No | ||
| 12 | HIPK2 | 17180 | 2530 | 0.756 | 0.7241 | No | ||
| 13 | ZFYVE9 | 10501 2449 | 3100 | 0.408 | 0.7031 | No | ||
| 14 | GDF9 | 20887 | 3306 | 0.319 | 0.6996 | No | ||
| 15 | CIDEA | 23529 | 3822 | 0.186 | 0.6762 | No | ||
| 16 | ACVR2A | 8542 | 3998 | 0.153 | 0.6704 | No | ||
| 17 | BMPR2 | 14241 4081 | 4406 | 0.100 | 0.6509 | No | ||
| 18 | ACVR2B | 19276 | 4695 | 0.079 | 0.6372 | No | ||
| 19 | TGFBR1 | 16213 10165 5745 | 4776 | 0.074 | 0.6346 | No | ||
| 20 | ACVRL1 | 22354 | 5579 | 0.045 | 0.5925 | No | ||
| 21 | KLF10 | 22308 | 6387 | 0.026 | 0.5497 | No | ||
| 22 | SOST | 20210 | 7206 | 0.014 | 0.5060 | No | ||
| 23 | TGFBR3 | 16453 | 7294 | 0.012 | 0.5016 | No | ||
| 24 | FOXH1 | 22238 | 7539 | 0.009 | 0.4887 | No | ||
| 25 | GDF5 | 4768 | 8111 | 0.001 | 0.4579 | No | ||
| 26 | EID2 | 11815 | 8379 | -0.003 | 0.4436 | No | ||
| 27 | LEFTY2 | 6735 | 9783 | -0.020 | 0.3686 | No | ||
| 28 | IL17F | 13992 | 9961 | -0.023 | 0.3596 | No | ||
| 29 | SMAD7 | 23513 | 10122 | -0.025 | 0.3515 | No | ||
| 30 | RGMB | 12736 7585 | 10481 | -0.030 | 0.3330 | No | ||
| 31 | FMOD | 14133 | 10807 | -0.035 | 0.3163 | No | ||
| 32 | GDF10 | 4767 | 11221 | -0.043 | 0.2951 | No | ||
| 33 | SNX6 | 21070 | 11459 | -0.048 | 0.2835 | No | ||
| 34 | LTBP2 | 5007 | 11604 | -0.051 | 0.2769 | No | ||
| 35 | GDF15 | 10721 | 11659 | -0.052 | 0.2752 | No | ||
| 36 | CITED2 | 5118 14477 | 12078 | -0.063 | 0.2542 | No | ||
| 37 | BMPR1A | 8660 4450 | 13026 | -0.096 | 0.2055 | No | ||
| 38 | SMAD1 | 18831 922 | 13610 | -0.131 | 0.1772 | No | ||
| 39 | SMURF1 | 13289 7989 | 14136 | -0.177 | 0.1531 | No | ||
| 40 | HPGD | 4867 | 14846 | -0.281 | 0.1215 | No | ||
| 41 | MAP3K7 | 16255 | 15341 | -0.423 | 0.1048 | No | ||
| 42 | STUB1 | 23070 | 16508 | -1.140 | 0.0689 | No | ||
| 43 | CDKN2B | 15840 | 17160 | -1.894 | 0.0784 | No |