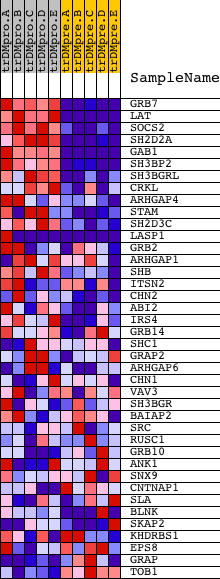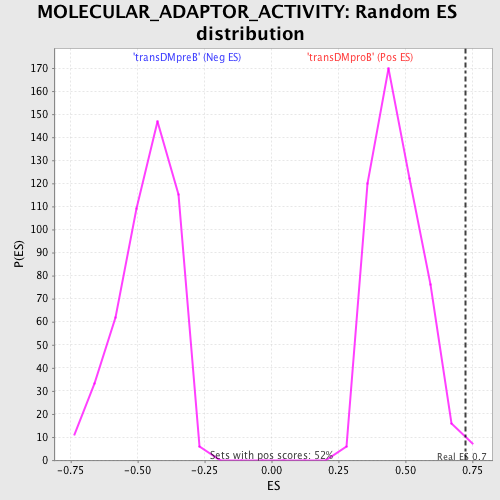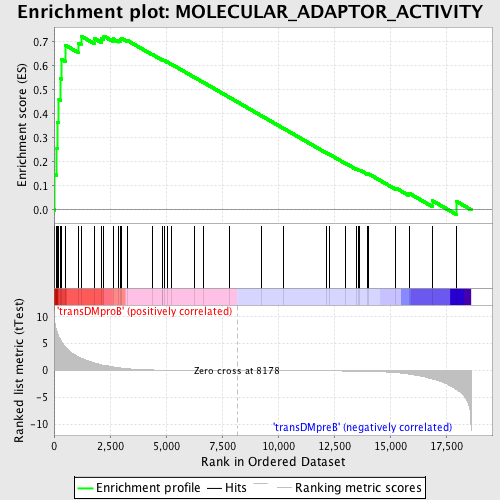
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | MOLECULAR_ADAPTOR_ACTIVITY |
| Enrichment Score (ES) | 0.72386765 |
| Normalized Enrichment Score (NES) | 1.5434953 |
| Nominal p-value | 0.00967118 |
| FDR q-value | 0.30228844 |
| FWER p-Value | 0.972 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GRB7 | 20673 | 22 | 9.600 | 0.1486 | Yes | ||
| 2 | LAT | 17643 | 124 | 7.317 | 0.2573 | Yes | ||
| 3 | SOCS2 | 5694 | 148 | 7.026 | 0.3656 | Yes | ||
| 4 | SH2D2A | 6569 | 215 | 6.241 | 0.4594 | Yes | ||
| 5 | GAB1 | 18828 | 280 | 5.825 | 0.5469 | Yes | ||
| 6 | SH3BP2 | 16874 | 345 | 5.295 | 0.6260 | Yes | ||
| 7 | SH3BGRL | 24267 | 508 | 4.382 | 0.6856 | Yes | ||
| 8 | CRKL | 4560 | 1082 | 2.544 | 0.6945 | Yes | ||
| 9 | ARHGAP4 | 9343 | 1204 | 2.301 | 0.7239 | Yes | ||
| 10 | STAM | 2912 15117 | 1795 | 1.426 | 0.7144 | No | ||
| 11 | SH2D3C | 2864 | 2100 | 1.072 | 0.7147 | No | ||
| 12 | LASP1 | 4986 4985 1248 | 2224 | 0.997 | 0.7236 | No | ||
| 13 | GRB2 | 20149 | 2633 | 0.675 | 0.7122 | No | ||
| 14 | ARHGAP1 | 6001 10448 | 2873 | 0.518 | 0.7074 | No | ||
| 15 | SHB | 10493 | 2962 | 0.469 | 0.7100 | No | ||
| 16 | ITSN2 | 21327 | 2996 | 0.457 | 0.7154 | No | ||
| 17 | CHN2 | 7652 | 3280 | 0.329 | 0.7053 | No | ||
| 18 | ABI2 | 4037 6830 | 4391 | 0.102 | 0.6471 | No | ||
| 19 | IRS4 | 9183 4926 | 4827 | 0.071 | 0.6248 | No | ||
| 20 | GRB14 | 14574 2719 | 4858 | 0.070 | 0.6243 | No | ||
| 21 | SHC1 | 9813 9812 5430 | 4928 | 0.066 | 0.6216 | No | ||
| 22 | GRAP2 | 5113 9398 | 5041 | 0.061 | 0.6165 | No | ||
| 23 | ARHGAP6 | 24207 | 5261 | 0.055 | 0.6056 | No | ||
| 24 | CHN1 | 4246 | 6264 | 0.029 | 0.5521 | No | ||
| 25 | VAV3 | 1774 1848 15443 | 6652 | 0.022 | 0.5316 | No | ||
| 26 | SH3BGR | 11937 | 7842 | 0.004 | 0.4677 | No | ||
| 27 | BAIAP2 | 449 4236 | 9266 | -0.014 | 0.3913 | No | ||
| 28 | SRC | 5507 | 10226 | -0.026 | 0.3401 | No | ||
| 29 | RUSC1 | 1853 15283 | 12151 | -0.065 | 0.2375 | No | ||
| 30 | GRB10 | 4799 | 12295 | -0.070 | 0.2309 | No | ||
| 31 | ANK1 | 22501 18650 4385 8590 18649 | 12997 | -0.095 | 0.1946 | No | ||
| 32 | SNX9 | 23132 | 13488 | -0.121 | 0.1702 | No | ||
| 33 | CNTNAP1 | 11987 | 13605 | -0.130 | 0.1659 | No | ||
| 34 | SLA | 5449 | 13654 | -0.134 | 0.1654 | No | ||
| 35 | BLNK | 23681 3691 | 13987 | -0.164 | 0.1501 | No | ||
| 36 | SKAP2 | 17153 | 14049 | -0.170 | 0.1495 | No | ||
| 37 | KHDRBS1 | 5405 9778 | 15253 | -0.393 | 0.0909 | No | ||
| 38 | EPS8 | 16948 | 15842 | -0.652 | 0.0694 | No | ||
| 39 | GRAP | 20851 | 16894 | -1.600 | 0.0378 | No | ||
| 40 | TOB1 | 20703 | 17958 | -3.518 | 0.0354 | No |