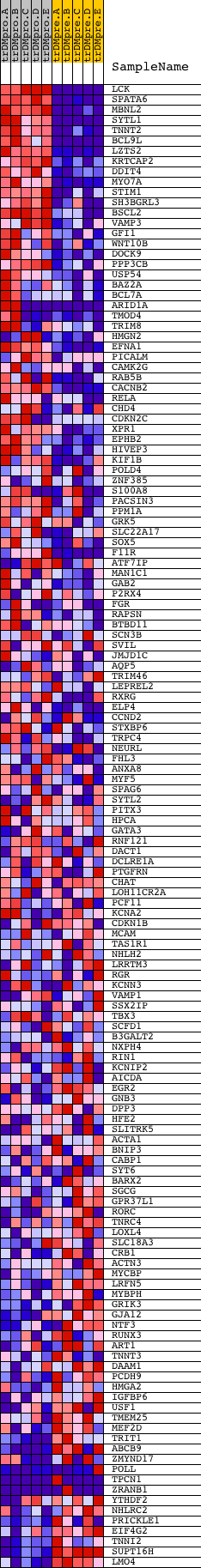
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$LBP1_Q6 |
| Enrichment Score (ES) | 0.63263124 |
| Normalized Enrichment Score (NES) | 1.6034867 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06345621 |
| FWER p-Value | 0.258 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LCK | 15746 | 27 | 9.283 | 0.0674 | Yes | ||
| 2 | SPATA6 | 16134 | 50 | 8.522 | 0.1295 | Yes | ||
| 3 | MBNL2 | 21932 | 197 | 6.415 | 0.1692 | Yes | ||
| 4 | SYTL1 | 15728 | 242 | 6.022 | 0.2115 | Yes | ||
| 5 | TNNT2 | 14113 | 262 | 5.908 | 0.2543 | Yes | ||
| 6 | BCL9L | 19475 | 267 | 5.879 | 0.2978 | Yes | ||
| 7 | LZTS2 | 10362 | 356 | 5.232 | 0.3318 | Yes | ||
| 8 | KRTCAP2 | 15541 | 494 | 4.444 | 0.3574 | Yes | ||
| 9 | DDIT4 | 19762 | 566 | 4.118 | 0.3841 | Yes | ||
| 10 | MYO7A | 9439 | 575 | 4.072 | 0.4139 | Yes | ||
| 11 | STIM1 | 18164 | 689 | 3.617 | 0.4346 | Yes | ||
| 12 | SH3BGRL3 | 15722 | 691 | 3.612 | 0.4614 | Yes | ||
| 13 | BSCL2 | 23940 | 752 | 3.379 | 0.4832 | Yes | ||
| 14 | VAMP3 | 15660 | 898 | 3.007 | 0.4977 | Yes | ||
| 15 | GFI1 | 16451 | 1010 | 2.707 | 0.5118 | Yes | ||
| 16 | WNT10B | 5874 | 1081 | 2.548 | 0.5269 | Yes | ||
| 17 | DOCK9 | 8369 | 1109 | 2.488 | 0.5439 | Yes | ||
| 18 | PPP3CB | 5285 | 1113 | 2.475 | 0.5621 | Yes | ||
| 19 | USP54 | 21910 | 1165 | 2.387 | 0.5771 | Yes | ||
| 20 | BAZ2A | 4371 | 1422 | 1.937 | 0.5776 | Yes | ||
| 21 | BCL7A | 8063 | 1550 | 1.750 | 0.5837 | Yes | ||
| 22 | ARID1A | 8214 2501 13554 | 1644 | 1.632 | 0.5908 | Yes | ||
| 23 | TMOD4 | 1816 1884 15509 | 1860 | 1.335 | 0.5891 | Yes | ||
| 24 | TRIM8 | 23824 | 1898 | 1.293 | 0.5967 | Yes | ||
| 25 | HMGN2 | 9095 | 1929 | 1.257 | 0.6044 | Yes | ||
| 26 | EFNA1 | 15279 | 2031 | 1.136 | 0.6073 | Yes | ||
| 27 | PICALM | 18191 | 2059 | 1.107 | 0.6141 | Yes | ||
| 28 | CAMK2G | 21905 | 2068 | 1.101 | 0.6218 | Yes | ||
| 29 | RAB5B | 359 19591 3430 | 2126 | 1.041 | 0.6265 | Yes | ||
| 30 | CACNB2 | 4466 8678 | 2250 | 0.982 | 0.6271 | Yes | ||
| 31 | RELA | 23783 | 2280 | 0.956 | 0.6326 | Yes | ||
| 32 | CHD4 | 8418 17281 4225 | 2472 | 0.796 | 0.6282 | No | ||
| 33 | CDKN2C | 15806 2321 | 2761 | 0.583 | 0.6169 | No | ||
| 34 | XPR1 | 5383 | 2890 | 0.507 | 0.6138 | No | ||
| 35 | EPHB2 | 4675 2440 8910 | 2951 | 0.476 | 0.6141 | No | ||
| 36 | HIVEP3 | 4973 | 2968 | 0.468 | 0.6167 | No | ||
| 37 | KIF1B | 15671 2476 4950 15673 | 3014 | 0.449 | 0.6176 | No | ||
| 38 | POLD4 | 12822 | 3141 | 0.386 | 0.6136 | No | ||
| 39 | ZNF385 | 22106 2297 | 3145 | 0.386 | 0.6163 | No | ||
| 40 | S100A8 | 15527 | 3362 | 0.301 | 0.6068 | No | ||
| 41 | PACSIN3 | 14948 8149 2776 2665 | 3386 | 0.294 | 0.6078 | No | ||
| 42 | PPM1A | 5279 9607 | 3450 | 0.272 | 0.6064 | No | ||
| 43 | GRK5 | 9036 23814 | 3495 | 0.261 | 0.6059 | No | ||
| 44 | SLC22A17 | 21830 3096 | 3599 | 0.235 | 0.6021 | No | ||
| 45 | SOX5 | 9850 1044 5480 16937 | 3645 | 0.223 | 0.6013 | No | ||
| 46 | F11R | 9199 | 3669 | 0.218 | 0.6017 | No | ||
| 47 | ATF7IP | 12028 13038 | 3684 | 0.215 | 0.6025 | No | ||
| 48 | MAN1C1 | 10514 6064 15714 | 3715 | 0.207 | 0.6025 | No | ||
| 49 | GAB2 | 1821 18184 2025 | 3956 | 0.160 | 0.5907 | No | ||
| 50 | P2RX4 | 3520 16711 | 4031 | 0.148 | 0.5877 | No | ||
| 51 | FGR | 4723 | 4353 | 0.105 | 0.5711 | No | ||
| 52 | RAPSN | 2879 14951 | 4503 | 0.092 | 0.5638 | No | ||
| 53 | BTBD11 | 13147 | 4712 | 0.078 | 0.5531 | No | ||
| 54 | SCN3B | 39 3117 6163 10646 | 4850 | 0.070 | 0.5462 | No | ||
| 55 | SVIL | 1990 1942 10334 1976 23626 1946 1945 2041 | 4916 | 0.067 | 0.5432 | No | ||
| 56 | JMJD1C | 11531 19996 | 4975 | 0.064 | 0.5405 | No | ||
| 57 | AQP5 | 22365 | 5098 | 0.060 | 0.5343 | No | ||
| 58 | TRIM46 | 1779 15281 | 5111 | 0.059 | 0.5341 | No | ||
| 59 | LEPREL2 | 17001 | 5142 | 0.058 | 0.5329 | No | ||
| 60 | RXRG | 14063 | 5238 | 0.055 | 0.5282 | No | ||
| 61 | ELP4 | 8094 | 5241 | 0.055 | 0.5285 | No | ||
| 62 | CCND2 | 16987 | 5335 | 0.052 | 0.5239 | No | ||
| 63 | STXBP6 | 21076 | 5457 | 0.048 | 0.5177 | No | ||
| 64 | TRPC4 | 15593 | 5527 | 0.046 | 0.5143 | No | ||
| 65 | NEURL | 5164 | 5986 | 0.035 | 0.4897 | No | ||
| 66 | FHL3 | 16092 2457 | 6240 | 0.029 | 0.4762 | No | ||
| 67 | ANXA8 | 22046 | 6532 | 0.024 | 0.4607 | No | ||
| 68 | MYF5 | 19632 | 6743 | 0.021 | 0.4495 | No | ||
| 69 | SPAG6 | 22650 | 6889 | 0.018 | 0.4417 | No | ||
| 70 | SYTL2 | 18190 1539 3730 | 6951 | 0.017 | 0.4386 | No | ||
| 71 | PITX3 | 9571 | 7034 | 0.016 | 0.4343 | No | ||
| 72 | HPCA | 9118 | 7228 | 0.013 | 0.4239 | No | ||
| 73 | GATA3 | 9004 | 7615 | 0.008 | 0.4031 | No | ||
| 74 | RNF121 | 7952 373 | 7616 | 0.008 | 0.4031 | No | ||
| 75 | DACT1 | 7203 | 7715 | 0.007 | 0.3979 | No | ||
| 76 | DCLRE1A | 23641 | 7804 | 0.005 | 0.3931 | No | ||
| 77 | PTGFRN | 15224 | 7822 | 0.005 | 0.3923 | No | ||
| 78 | CHAT | 21882 | 8042 | 0.002 | 0.3804 | No | ||
| 79 | LOH11CR2A | 19498 | 8194 | -0.000 | 0.3723 | No | ||
| 80 | PCF11 | 121 | 8489 | -0.004 | 0.3564 | No | ||
| 81 | KCNA2 | 15458 | 8727 | -0.007 | 0.3436 | No | ||
| 82 | CDKN1B | 4512 8730 | 8756 | -0.007 | 0.3421 | No | ||
| 83 | MCAM | 3109 19479 | 8833 | -0.008 | 0.3381 | No | ||
| 84 | TAS1R1 | 15657 | 8934 | -0.010 | 0.3327 | No | ||
| 85 | NHLH2 | 9462 | 9251 | -0.013 | 0.3157 | No | ||
| 86 | LRRTM3 | 19744 | 9280 | -0.014 | 0.3143 | No | ||
| 87 | RGR | 21874 3042 | 9437 | -0.016 | 0.3060 | No | ||
| 88 | KCNN3 | 15538 1913 | 9580 | -0.018 | 0.2984 | No | ||
| 89 | VAMP1 | 5846 1043 | 9721 | -0.020 | 0.2910 | No | ||
| 90 | SSX2IP | 13617 | 9940 | -0.022 | 0.2794 | No | ||
| 91 | TBX3 | 16723 | 10010 | -0.023 | 0.2758 | No | ||
| 92 | SCFD1 | 21278 2157 | 10151 | -0.025 | 0.2684 | No | ||
| 93 | B3GALT2 | 6498 | 10395 | -0.029 | 0.2555 | No | ||
| 94 | NXPH4 | 19602 | 10425 | -0.029 | 0.2541 | No | ||
| 95 | RIN1 | 10349 | 10552 | -0.031 | 0.2475 | No | ||
| 96 | KCNIP2 | 23655 3725 | 10757 | -0.035 | 0.2367 | No | ||
| 97 | AICDA | 17295 1087 | 10954 | -0.038 | 0.2264 | No | ||
| 98 | EGR2 | 8886 | 10983 | -0.039 | 0.2252 | No | ||
| 99 | GNB3 | 9027 | 11208 | -0.043 | 0.2134 | No | ||
| 100 | DPP3 | 7954 3746 | 11227 | -0.043 | 0.2127 | No | ||
| 101 | HFE2 | 1777 15495 | 11262 | -0.044 | 0.2112 | No | ||
| 102 | SLITRK5 | 21939 | 11394 | -0.047 | 0.2045 | No | ||
| 103 | ACTA1 | 18715 | 11547 | -0.050 | 0.1966 | No | ||
| 104 | BNIP3 | 17581 | 11567 | -0.050 | 0.1960 | No | ||
| 105 | CABP1 | 16412 | 11578 | -0.051 | 0.1958 | No | ||
| 106 | SYT6 | 13437 7042 12040 | 11590 | -0.051 | 0.1956 | No | ||
| 107 | BARX2 | 8649 | 11747 | -0.055 | 0.1875 | No | ||
| 108 | SGCG | 21798 | 11882 | -0.058 | 0.1807 | No | ||
| 109 | GPR37L1 | 13825 | 11950 | -0.059 | 0.1775 | No | ||
| 110 | RORC | 9732 | 12036 | -0.062 | 0.1734 | No | ||
| 111 | TNRC4 | 15514 | 12081 | -0.063 | 0.1715 | No | ||
| 112 | LOXL4 | 23672 | 12089 | -0.063 | 0.1715 | No | ||
| 113 | SLC18A3 | 21883 | 12327 | -0.070 | 0.1592 | No | ||
| 114 | CRB1 | 5031 | 12664 | -0.082 | 0.1417 | No | ||
| 115 | ACTN3 | 23967 8540 | 12742 | -0.085 | 0.1381 | No | ||
| 116 | MYCBP | 7093 | 12974 | -0.094 | 0.1263 | No | ||
| 117 | LRFN5 | 6213 | 13219 | -0.106 | 0.1139 | No | ||
| 118 | MYBPH | 14130 | 13434 | -0.118 | 0.1032 | No | ||
| 119 | GRIK3 | 16083 | 13491 | -0.121 | 0.1011 | No | ||
| 120 | GJA12 | 20433 1323 | 13629 | -0.132 | 0.0946 | No | ||
| 121 | NTF3 | 16991 | 13693 | -0.137 | 0.0922 | No | ||
| 122 | RUNX3 | 8702 | 13786 | -0.144 | 0.0883 | No | ||
| 123 | ART1 | 660 4414 | 13975 | -0.162 | 0.0793 | No | ||
| 124 | TNNT3 | 18004 | 14319 | -0.199 | 0.0623 | No | ||
| 125 | DAAM1 | 5536 | 14391 | -0.208 | 0.0600 | No | ||
| 126 | PCDH9 | 21736 | 14458 | -0.218 | 0.0580 | No | ||
| 127 | HMGA2 | 4857 9098 3341 3444 | 14557 | -0.234 | 0.0544 | No | ||
| 128 | IGFBP6 | 9161 | 14619 | -0.243 | 0.0529 | No | ||
| 129 | USF1 | 5832 10260 10261 | 14870 | -0.287 | 0.0415 | No | ||
| 130 | TMEM25 | 2992 19141 | 14887 | -0.293 | 0.0428 | No | ||
| 131 | MEF2D | 9379 | 15241 | -0.390 | 0.0266 | No | ||
| 132 | TRIT1 | 16098 | 15398 | -0.439 | 0.0214 | No | ||
| 133 | ABCB9 | 3576 3546 7096 | 15412 | -0.445 | 0.0240 | No | ||
| 134 | ZMYND17 | 21909 | 16075 | -0.818 | -0.0057 | No | ||
| 135 | POLL | 23658 3688 | 16240 | -0.961 | -0.0075 | No | ||
| 136 | TPCN1 | 16398 6383 10915 | 16268 | -0.978 | -0.0017 | No | ||
| 137 | ZRANB1 | 6884 11702 | 16325 | -1.004 | 0.0027 | No | ||
| 138 | YTHDF2 | 5623 5624 | 16611 | -1.249 | -0.0034 | No | ||
| 139 | NHLRC2 | 23828 | 16845 | -1.540 | -0.0046 | No | ||
| 140 | PRICKLE1 | 8379 | 16882 | -1.584 | 0.0052 | No | ||
| 141 | EIF4G2 | 1908 8892 | 17159 | -1.893 | 0.0043 | No | ||
| 142 | TNNI2 | 18005 | 17174 | -1.914 | 0.0178 | No | ||
| 143 | SUPT16H | 8541 | 17944 | -3.485 | 0.0020 | No | ||
| 144 | LMO4 | 15151 | 18276 | -4.622 | 0.0184 | No |