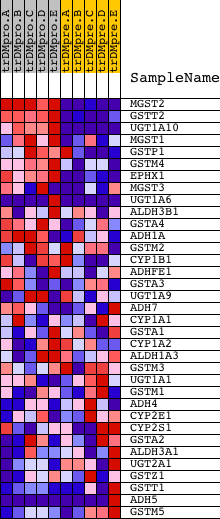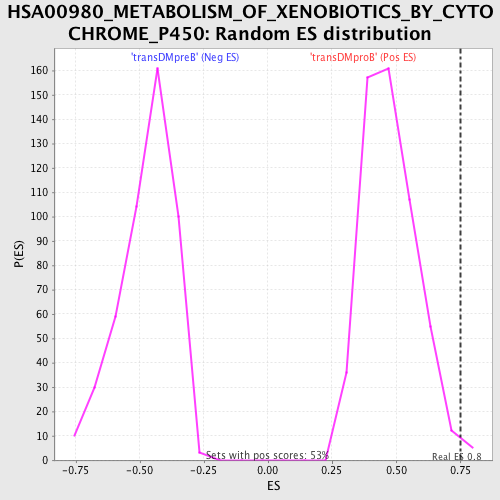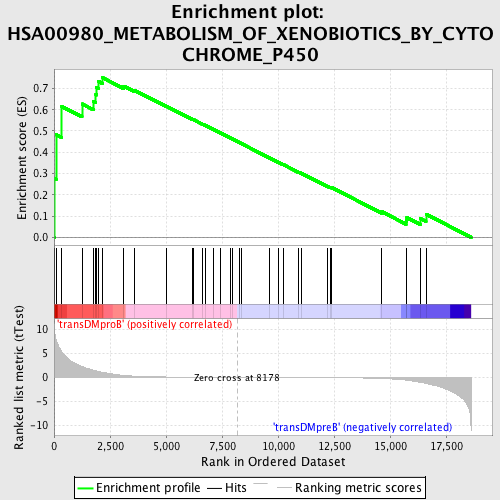
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | HSA00980_METABOLISM_OF_XENOBIOTICS_BY_CYTOCHROME_P450 |
| Enrichment Score (ES) | 0.75111234 |
| Normalized Enrichment Score (NES) | 1.5750744 |
| Nominal p-value | 0.009380863 |
| FDR q-value | 0.43283486 |
| FWER p-Value | 0.646 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MGST2 | 15597 | 12 | 10.104 | 0.2757 | Yes | ||
| 2 | GSTT2 | 9052 | 91 | 7.735 | 0.4831 | Yes | ||
| 3 | UGT1A10 | 6908 | 342 | 5.321 | 0.6151 | Yes | ||
| 4 | MGST1 | 17254 | 1246 | 2.214 | 0.6271 | Yes | ||
| 5 | GSTP1 | 9051 4812 4811 | 1764 | 1.471 | 0.6395 | Yes | ||
| 6 | GSTM4 | 15199 | 1861 | 1.335 | 0.6709 | Yes | ||
| 7 | EPHX1 | 13734 | 1894 | 1.300 | 0.7047 | Yes | ||
| 8 | MGST3 | 13771 12349 | 1991 | 1.179 | 0.7318 | Yes | ||
| 9 | UGT1A6 | 3969 4079 6911 13591 | 2151 | 1.019 | 0.7511 | Yes | ||
| 10 | ALDH3B1 | 12569 23949 | 3108 | 0.403 | 0.7107 | No | ||
| 11 | GSTA4 | 19371 | 3603 | 0.234 | 0.6905 | No | ||
| 12 | ADH1A | 15406 | 5008 | 0.063 | 0.6167 | No | ||
| 13 | GSTM2 | 9049 4810 | 6190 | 0.030 | 0.5539 | No | ||
| 14 | CYP1B1 | 4581 | 6202 | 0.030 | 0.5542 | No | ||
| 15 | ADHFE1 | 4049 8008 | 6629 | 0.023 | 0.5319 | No | ||
| 16 | GSTA3 | 14287 | 6741 | 0.021 | 0.5265 | No | ||
| 17 | UGT1A9 | 11849 11850 | 7126 | 0.015 | 0.5062 | No | ||
| 18 | ADH7 | 15408 | 7430 | 0.011 | 0.4902 | No | ||
| 19 | CYP1A1 | 19428 | 7874 | 0.004 | 0.4665 | No | ||
| 20 | GSTA1 | 4807 | 7979 | 0.003 | 0.4609 | No | ||
| 21 | CYP1A2 | 19097 | 8280 | -0.001 | 0.4448 | No | ||
| 22 | ALDH1A3 | 17802 | 8375 | -0.003 | 0.4398 | No | ||
| 23 | GSTM3 | 9050 | 9635 | -0.018 | 0.3726 | No | ||
| 24 | UGT1A1 | 11851 | 9994 | -0.023 | 0.3540 | No | ||
| 25 | GSTM1 | 4809 9048 | 10235 | -0.026 | 0.3418 | No | ||
| 26 | ADH4 | 15405 | 10918 | -0.037 | 0.3061 | No | ||
| 27 | CYP2E1 | 18025 | 11025 | -0.039 | 0.3015 | No | ||
| 28 | CYP2S1 | 17921 | 12203 | -0.067 | 0.2399 | No | ||
| 29 | GSTA2 | 4808 6173 | 12345 | -0.071 | 0.2343 | No | ||
| 30 | ALDH3A1 | 20854 | 12392 | -0.072 | 0.2338 | No | ||
| 31 | UGT2A1 | 8247 | 14635 | -0.246 | 0.1199 | No | ||
| 32 | GSTZ1 | 21193 | 15711 | -0.584 | 0.0780 | No | ||
| 33 | GSTT1 | 19730 | 15746 | -0.603 | 0.0926 | No | ||
| 34 | ADH5 | 8554 15404 | 16344 | -1.012 | 0.0882 | No | ||
| 35 | GSTM5 | 15455 | 16603 | -1.244 | 0.1083 | No |