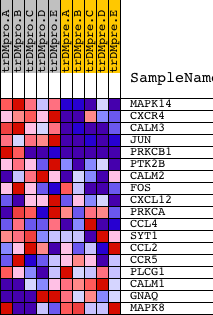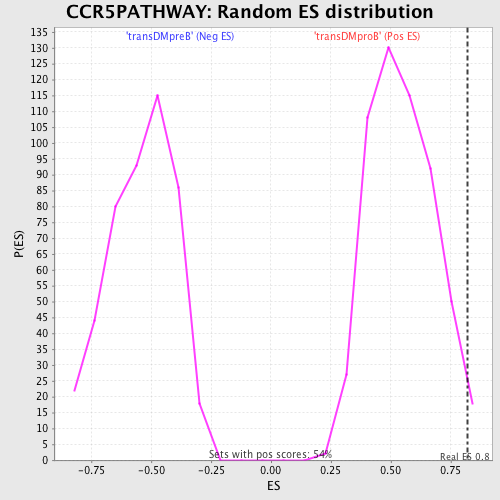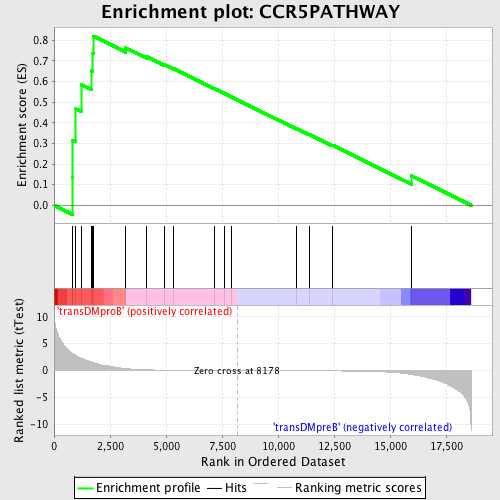
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | CCR5PATHWAY |
| Enrichment Score (ES) | 0.82012683 |
| Normalized Enrichment Score (NES) | 1.4955472 |
| Nominal p-value | 0.016605167 |
| FDR q-value | 0.24949393 |
| FWER p-Value | 0.979 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MAPK14 | 23313 | 828 | 3.176 | 0.1356 | Yes | ||
| 2 | CXCR4 | 13844 | 833 | 3.161 | 0.3147 | Yes | ||
| 3 | CALM3 | 8682 | 973 | 2.818 | 0.4670 | Yes | ||
| 4 | JUN | 15832 | 1203 | 2.302 | 0.5853 | Yes | ||
| 5 | PRKCB1 | 1693 9574 | 1672 | 1.605 | 0.6512 | Yes | ||
| 6 | PTK2B | 21776 | 1704 | 1.562 | 0.7381 | Yes | ||
| 7 | CALM2 | 8681 | 1752 | 1.490 | 0.8201 | Yes | ||
| 8 | FOS | 21202 | 3200 | 0.360 | 0.7628 | No | ||
| 9 | CXCL12 | 9792 1182 1031 | 4120 | 0.133 | 0.7209 | No | ||
| 10 | PRKCA | 20174 | 4927 | 0.066 | 0.6814 | No | ||
| 11 | CCL4 | 1362 20730 | 5320 | 0.052 | 0.6632 | No | ||
| 12 | SYT1 | 5565 | 7160 | 0.014 | 0.5652 | No | ||
| 13 | CCL2 | 9788 | 7586 | 0.009 | 0.5428 | No | ||
| 14 | CCR5 | 8754 4534 | 7927 | 0.004 | 0.5247 | No | ||
| 15 | PLCG1 | 14753 | 10828 | -0.036 | 0.3708 | No | ||
| 16 | CALM1 | 21184 | 11385 | -0.046 | 0.3436 | No | ||
| 17 | GNAQ | 4786 23909 3685 | 12422 | -0.073 | 0.2920 | No | ||
| 18 | MAPK8 | 6459 | 15940 | -0.722 | 0.1439 | No |