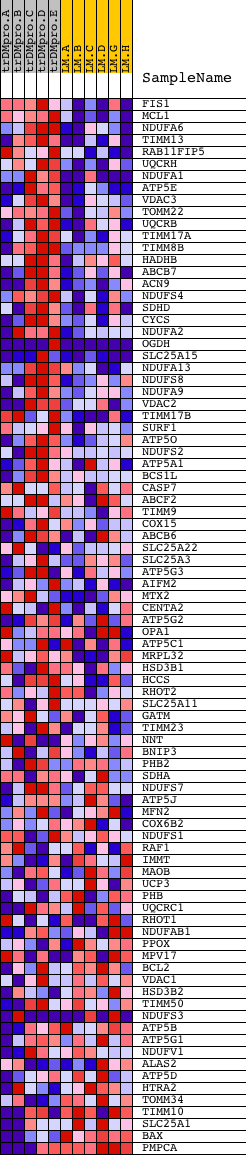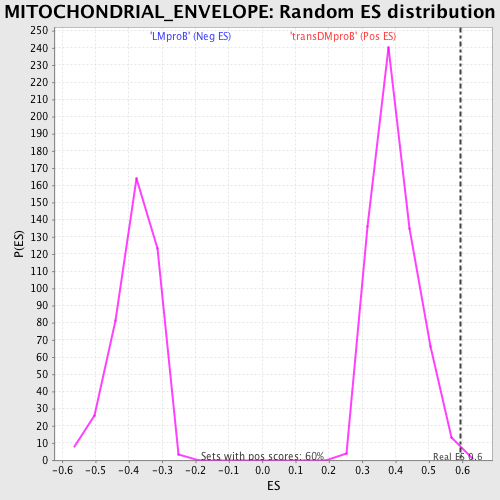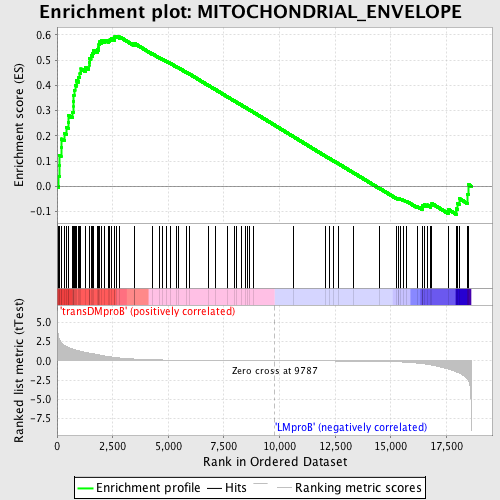
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | MITOCHONDRIAL_ENVELOPE |
| Enrichment Score (ES) | 0.59512204 |
| Normalized Enrichment Score (NES) | 1.5002552 |
| Nominal p-value | 0.0016806723 |
| FDR q-value | 0.46122253 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FIS1 | 16669 3579 | 81 | 2.887 | 0.0398 | Yes | ||
| 2 | MCL1 | 15502 | 99 | 2.742 | 0.0808 | Yes | ||
| 3 | NDUFA6 | 12473 | 107 | 2.667 | 0.1212 | Yes | ||
| 4 | TIMM13 | 14974 | 179 | 2.310 | 0.1527 | Yes | ||
| 5 | RAB11FIP5 | 17089 | 187 | 2.296 | 0.1874 | Yes | ||
| 6 | UQCRH | 12378 | 330 | 1.989 | 0.2102 | Yes | ||
| 7 | NDUFA1 | 24170 | 427 | 1.811 | 0.2327 | Yes | ||
| 8 | ATP5E | 14321 | 506 | 1.707 | 0.2546 | Yes | ||
| 9 | VDAC3 | 10284 | 520 | 1.693 | 0.2798 | Yes | ||
| 10 | TOMM22 | 5863 | 689 | 1.540 | 0.2943 | Yes | ||
| 11 | UQCRB | 12545 | 735 | 1.494 | 0.3147 | Yes | ||
| 12 | TIMM17A | 13822 | 739 | 1.490 | 0.3373 | Yes | ||
| 13 | TIMM8B | 11391 | 749 | 1.481 | 0.3595 | Yes | ||
| 14 | HADHB | 3514 10527 | 764 | 1.463 | 0.3811 | Yes | ||
| 15 | ABCB7 | 24085 | 805 | 1.427 | 0.4008 | Yes | ||
| 16 | ACN9 | 12920 | 873 | 1.366 | 0.4181 | Yes | ||
| 17 | NDUFS4 | 9452 5157 3173 | 969 | 1.290 | 0.4327 | Yes | ||
| 18 | SDHD | 19125 | 1026 | 1.248 | 0.4487 | Yes | ||
| 19 | CYCS | 8821 | 1071 | 1.216 | 0.4650 | Yes | ||
| 20 | NDUFA2 | 9450 | 1265 | 1.089 | 0.4712 | Yes | ||
| 21 | OGDH | 9502 1412 | 1433 | 0.996 | 0.4774 | Yes | ||
| 22 | SLC25A15 | 356 3819 | 1436 | 0.995 | 0.4925 | Yes | ||
| 23 | NDUFA13 | 7400 | 1451 | 0.990 | 0.5069 | Yes | ||
| 24 | NDUFS8 | 23950 | 1524 | 0.958 | 0.5177 | Yes | ||
| 25 | NDUFA9 | 12288 | 1595 | 0.923 | 0.5280 | Yes | ||
| 26 | VDAC2 | 22081 | 1629 | 0.901 | 0.5400 | Yes | ||
| 27 | TIMM17B | 24390 | 1806 | 0.797 | 0.5427 | Yes | ||
| 28 | SURF1 | 14645 | 1872 | 0.761 | 0.5508 | Yes | ||
| 29 | ATP5O | 22539 | 1878 | 0.758 | 0.5621 | Yes | ||
| 30 | NDUFS2 | 4089 13760 | 1905 | 0.742 | 0.5721 | Yes | ||
| 31 | ATP5A1 | 23505 | 1993 | 0.692 | 0.5780 | Yes | ||
| 32 | BCS1L | 14223 | 2140 | 0.622 | 0.5796 | Yes | ||
| 33 | CASP7 | 23826 | 2307 | 0.540 | 0.5789 | Yes | ||
| 34 | ABCF2 | 11338 6580 6579 | 2376 | 0.512 | 0.5831 | Yes | ||
| 35 | TIMM9 | 19686 | 2450 | 0.488 | 0.5866 | Yes | ||
| 36 | COX15 | 5931 | 2588 | 0.431 | 0.5858 | Yes | ||
| 37 | ABCB6 | 13162 | 2597 | 0.429 | 0.5919 | Yes | ||
| 38 | SLC25A22 | 12671 | 2655 | 0.411 | 0.5951 | Yes | ||
| 39 | SLC25A3 | 19644 | 2818 | 0.359 | 0.5919 | No | ||
| 40 | ATP5G3 | 14558 | 3463 | 0.219 | 0.5605 | No | ||
| 41 | AIFM2 | 3320 3407 20006 3317 | 3478 | 0.218 | 0.5630 | No | ||
| 42 | MTX2 | 14980 | 3489 | 0.215 | 0.5658 | No | ||
| 43 | CENTA2 | 20748 | 4276 | 0.126 | 0.5253 | No | ||
| 44 | ATP5G2 | 12610 | 4623 | 0.105 | 0.5082 | No | ||
| 45 | OPA1 | 13169 22800 7894 | 4726 | 0.099 | 0.5042 | No | ||
| 46 | ATP5C1 | 8635 | 4914 | 0.089 | 0.4955 | No | ||
| 47 | MRPL32 | 7961 | 5111 | 0.081 | 0.4862 | No | ||
| 48 | HSD3B1 | 15227 | 5362 | 0.072 | 0.4738 | No | ||
| 49 | HCCS | 9079 4842 | 5438 | 0.070 | 0.4708 | No | ||
| 50 | RHOT2 | 1533 10073 | 5836 | 0.058 | 0.4503 | No | ||
| 51 | SLC25A11 | 20366 1368 | 5952 | 0.056 | 0.4449 | No | ||
| 52 | GATM | 14451 | 6815 | 0.037 | 0.3990 | No | ||
| 53 | TIMM23 | 7004 5768 | 7129 | 0.032 | 0.3826 | No | ||
| 54 | NNT | 3273 5181 9471 3238 | 7663 | 0.024 | 0.3542 | No | ||
| 55 | BNIP3 | 17581 | 7980 | 0.020 | 0.3374 | No | ||
| 56 | PHB2 | 17286 | 8063 | 0.019 | 0.3333 | No | ||
| 57 | SDHA | 12436 | 8271 | 0.017 | 0.3224 | No | ||
| 58 | NDUFS7 | 19946 | 8476 | 0.014 | 0.3116 | No | ||
| 59 | ATP5J | 880 22555 | 8543 | 0.014 | 0.3082 | No | ||
| 60 | MFN2 | 15678 2417 | 8669 | 0.012 | 0.3017 | No | ||
| 61 | COX6B2 | 17983 4110 | 8836 | 0.010 | 0.2929 | No | ||
| 62 | NDUFS1 | 13940 | 10640 | -0.009 | 0.1957 | No | ||
| 63 | RAF1 | 17035 | 12066 | -0.026 | 0.1192 | No | ||
| 64 | IMMT | 367 8029 | 12249 | -0.029 | 0.1098 | No | ||
| 65 | MAOB | 24182 2556 | 12439 | -0.031 | 0.1001 | No | ||
| 66 | UCP3 | 3972 18175 | 12653 | -0.035 | 0.0891 | No | ||
| 67 | PHB | 9558 | 13331 | -0.047 | 0.0533 | No | ||
| 68 | UQCRC1 | 10259 | 14490 | -0.081 | -0.0080 | No | ||
| 69 | RHOT1 | 12227 | 15265 | -0.132 | -0.0477 | No | ||
| 70 | NDUFAB1 | 7667 | 15329 | -0.139 | -0.0490 | No | ||
| 71 | PPOX | 13761 4104 | 15352 | -0.141 | -0.0480 | No | ||
| 72 | MPV17 | 9407 3505 3543 | 15441 | -0.152 | -0.0505 | No | ||
| 73 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 15568 | -0.167 | -0.0547 | No | ||
| 74 | VDAC1 | 10282 | 15698 | -0.188 | -0.0588 | No | ||
| 75 | HSD3B2 | 4873 4874 15229 4875 9126 | 16196 | -0.304 | -0.0810 | No | ||
| 76 | TIMM50 | 17911 | 16420 | -0.377 | -0.0872 | No | ||
| 77 | NDUFS3 | 7553 12683 | 16422 | -0.378 | -0.0815 | No | ||
| 78 | ATP5B | 19846 | 16431 | -0.383 | -0.0761 | No | ||
| 79 | ATP5G1 | 8636 | 16499 | -0.407 | -0.0735 | No | ||
| 80 | NDUFV1 | 23953 | 16637 | -0.452 | -0.0740 | No | ||
| 81 | ALAS2 | 24229 | 16801 | -0.536 | -0.0746 | No | ||
| 82 | ATP5D | 19949 | 16843 | -0.556 | -0.0683 | No | ||
| 83 | HTRA2 | 1122 17103 | 17598 | -1.073 | -0.0926 | No | ||
| 84 | TOMM34 | 14359 | 17955 | -1.452 | -0.0896 | No | ||
| 85 | TIMM10 | 11390 2158 | 17991 | -1.495 | -0.0686 | No | ||
| 86 | SLC25A1 | 22645 | 18089 | -1.611 | -0.0492 | No | ||
| 87 | BAX | 17832 | 18451 | -2.419 | -0.0317 | No | ||
| 88 | PMPCA | 12421 7350 | 18502 | -2.649 | 0.0062 | No |