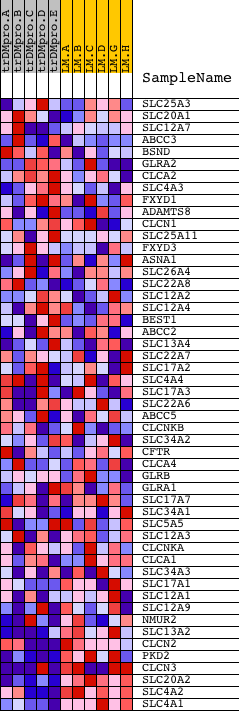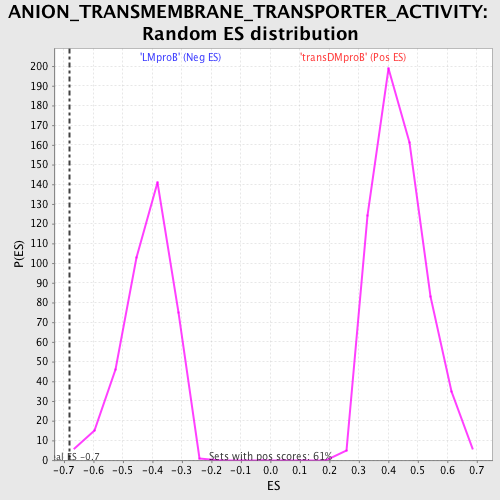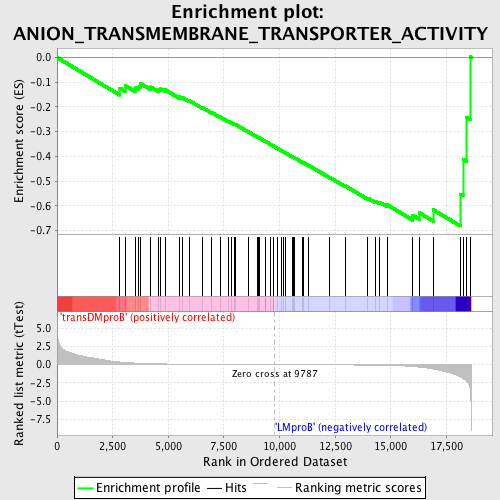
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | ANION_TRANSMEMBRANE_TRANSPORTER_ACTIVITY |
| Enrichment Score (ES) | -0.68158084 |
| Normalized Enrichment Score (NES) | -1.6378837 |
| Nominal p-value | 0.0025839794 |
| FDR q-value | 0.70184934 |
| FWER p-Value | 0.761 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC25A3 | 19644 | 2818 | 0.359 | -0.1238 | No | ||
| 2 | SLC20A1 | 14855 | 3055 | 0.298 | -0.1132 | No | ||
| 3 | SLC12A7 | 21603 | 3523 | 0.211 | -0.1220 | No | ||
| 4 | ABCC3 | 20293 | 3679 | 0.190 | -0.1155 | No | ||
| 5 | BSND | 15821 | 3755 | 0.182 | -0.1054 | No | ||
| 6 | GLRA2 | 24008 | 4200 | 0.132 | -0.1190 | No | ||
| 7 | CLCA2 | 15148 | 4575 | 0.107 | -0.1307 | No | ||
| 8 | SLC4A3 | 5454 14213 9829 | 4653 | 0.103 | -0.1268 | No | ||
| 9 | FXYD1 | 17874 85 | 4860 | 0.092 | -0.1308 | No | ||
| 10 | ADAMTS8 | 6626 | 5484 | 0.068 | -0.1590 | No | ||
| 11 | CLCN1 | 8744 | 5630 | 0.064 | -0.1618 | No | ||
| 12 | SLC25A11 | 20366 1368 | 5952 | 0.056 | -0.1748 | No | ||
| 13 | FXYD3 | 9370 | 6546 | 0.042 | -0.2034 | No | ||
| 14 | ASNA1 | 18810 | 6961 | 0.035 | -0.2230 | No | ||
| 15 | SLC26A4 | 21089 | 7323 | 0.029 | -0.2401 | No | ||
| 16 | SLC22A8 | 23944 | 7683 | 0.024 | -0.2576 | No | ||
| 17 | SLC12A2 | 23547 2026 | 7829 | 0.022 | -0.2637 | No | ||
| 18 | SLC12A4 | 18763 | 7992 | 0.020 | -0.2709 | No | ||
| 19 | BEST1 | 23747 | 8020 | 0.019 | -0.2708 | No | ||
| 20 | ABCC2 | 8755 23844 | 8586 | 0.013 | -0.3002 | No | ||
| 21 | SLC13A4 | 17185 | 8989 | 0.009 | -0.3212 | No | ||
| 22 | SLC22A7 | 22964 | 9042 | 0.008 | -0.3234 | No | ||
| 23 | SLC17A2 | 10163 5744 | 9089 | 0.008 | -0.3253 | No | ||
| 24 | SLC4A4 | 16807 12035 7036 | 9361 | 0.005 | -0.3395 | No | ||
| 25 | SLC17A3 | 3165 21694 | 9375 | 0.005 | -0.3398 | No | ||
| 26 | SLC22A6 | 5213 | 9583 | 0.002 | -0.3508 | No | ||
| 27 | ABCC5 | 22639 351 | 9727 | 0.001 | -0.3585 | No | ||
| 28 | CLCNKB | 7105 | 9730 | 0.001 | -0.3585 | No | ||
| 29 | SLC34A2 | 16845 | 9923 | -0.002 | -0.3687 | No | ||
| 30 | CFTR | 17518 | 10073 | -0.003 | -0.3765 | No | ||
| 31 | CLCA4 | 6041 | 10179 | -0.004 | -0.3818 | No | ||
| 32 | GLRB | 15312 1846 | 10281 | -0.005 | -0.3868 | No | ||
| 33 | GLRA1 | 9021 | 10567 | -0.008 | -0.4015 | No | ||
| 34 | SLC17A7 | 13081 | 10621 | -0.009 | -0.4037 | No | ||
| 35 | SLC34A1 | 9824 | 10655 | -0.009 | -0.4047 | No | ||
| 36 | SLC5A5 | 4322 | 11035 | -0.013 | -0.4241 | No | ||
| 37 | SLC12A3 | 3799 5451 | 11089 | -0.014 | -0.4258 | No | ||
| 38 | CLCNKA | 15690 | 11301 | -0.016 | -0.4359 | No | ||
| 39 | CLCA1 | 15149 | 12264 | -0.029 | -0.4855 | No | ||
| 40 | SLC34A3 | 14664 | 12954 | -0.040 | -0.5195 | No | ||
| 41 | SLC17A1 | 21693 | 13958 | -0.063 | -0.5686 | No | ||
| 42 | SLC12A1 | 14874 2772 2943 | 14300 | -0.074 | -0.5812 | No | ||
| 43 | SLC12A9 | 16332 3654 | 14489 | -0.081 | -0.5850 | No | ||
| 44 | NMUR2 | 20443 | 14837 | -0.099 | -0.5960 | No | ||
| 45 | SLC13A2 | 20336 | 15989 | -0.250 | -0.6385 | Yes | ||
| 46 | CLCN2 | 22637 | 16288 | -0.330 | -0.6288 | Yes | ||
| 47 | PKD2 | 16772 | 16902 | -0.587 | -0.6160 | Yes | ||
| 48 | CLCN3 | 8745 3782 3893 4524 | 18121 | -1.650 | -0.5528 | Yes | ||
| 49 | SLC20A2 | 18654 | 18249 | -1.893 | -0.4119 | Yes | ||
| 50 | SLC4A2 | 9828 | 18416 | -2.271 | -0.2437 | Yes | ||
| 51 | SLC4A1 | 20204 | 18563 | -3.260 | 0.0029 | Yes |