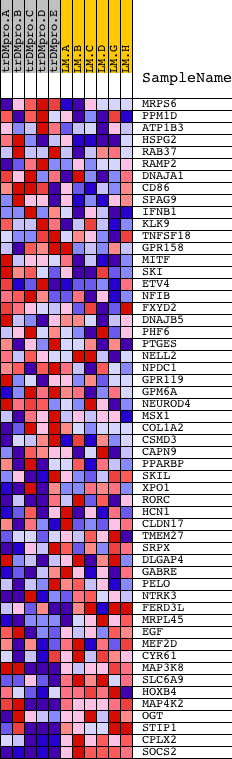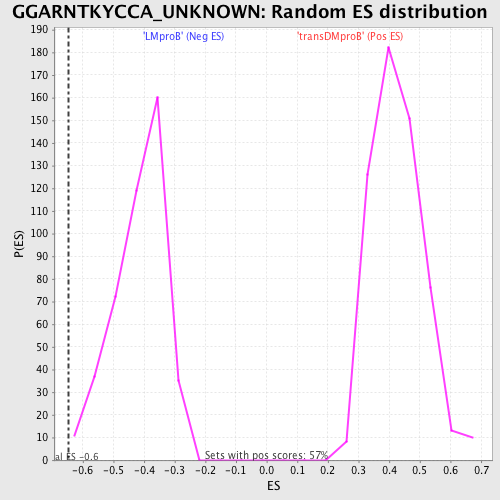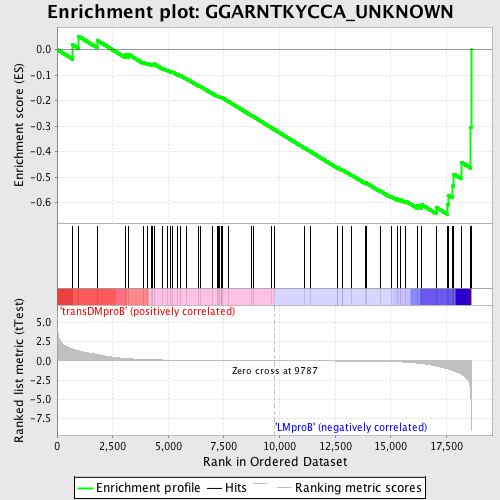
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | GGARNTKYCCA_UNKNOWN |
| Enrichment Score (ES) | -0.64613557 |
| Normalized Enrichment Score (NES) | -1.5600419 |
| Nominal p-value | 0.004608295 |
| FDR q-value | 0.5120605 |
| FWER p-Value | 0.568 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MRPS6 | 8656 | 703 | 1.529 | 0.0183 | No | ||
| 2 | PPM1D | 20721 | 956 | 1.299 | 0.0525 | No | ||
| 3 | ATP1B3 | 19037 | 1813 | 0.792 | 0.0356 | No | ||
| 4 | HSPG2 | 4884 16023 | 3064 | 0.295 | -0.0209 | No | ||
| 5 | RAB37 | 20602 | 3224 | 0.261 | -0.0199 | No | ||
| 6 | RAMP2 | 20660 | 3901 | 0.161 | -0.0504 | No | ||
| 7 | DNAJA1 | 4878 16242 2345 | 4080 | 0.144 | -0.0547 | No | ||
| 8 | CD86 | 1660 22601 | 4254 | 0.127 | -0.0593 | No | ||
| 9 | SPAG9 | 1322 12904 7707 | 4304 | 0.123 | -0.0574 | No | ||
| 10 | IFNB1 | 15846 | 4367 | 0.119 | -0.0564 | No | ||
| 11 | KLK9 | 18267 | 4753 | 0.097 | -0.0736 | No | ||
| 12 | TNFSF18 | 10789 6290 | 4941 | 0.088 | -0.0804 | No | ||
| 13 | GPR158 | 149 | 5116 | 0.081 | -0.0868 | No | ||
| 14 | MITF | 17349 | 5192 | 0.078 | -0.0880 | No | ||
| 15 | SKI | 5448 | 5426 | 0.070 | -0.0979 | No | ||
| 16 | ETV4 | 20212 | 5543 | 0.066 | -0.1018 | No | ||
| 17 | NFIB | 15855 | 5817 | 0.059 | -0.1143 | No | ||
| 18 | FXYD2 | 3038 19470 3068 3039 | 6365 | 0.046 | -0.1421 | No | ||
| 19 | DNAJB5 | 16236 | 6445 | 0.044 | -0.1447 | No | ||
| 20 | PHF6 | 2648 24340 | 6970 | 0.035 | -0.1717 | No | ||
| 21 | PTGES | 7224 | 7189 | 0.032 | -0.1822 | No | ||
| 22 | NELL2 | 12006 7020 22151 | 7233 | 0.031 | -0.1834 | No | ||
| 23 | NPDC1 | 9479 | 7302 | 0.030 | -0.1860 | No | ||
| 24 | GPR119 | 24159 | 7370 | 0.029 | -0.1886 | No | ||
| 25 | GPM6A | 3834 10618 | 7382 | 0.028 | -0.1881 | No | ||
| 26 | NEUROD4 | 19582 | 7421 | 0.028 | -0.1891 | No | ||
| 27 | MSX1 | 16547 | 7714 | 0.024 | -0.2040 | No | ||
| 28 | COL1A2 | 8772 | 8717 | 0.011 | -0.2576 | No | ||
| 29 | CSMD3 | 10735 7938 | 8823 | 0.010 | -0.2628 | No | ||
| 30 | CAPN9 | 3772 18422 | 9638 | 0.002 | -0.3066 | No | ||
| 31 | PPARBP | 1203 20263 1195 | 9774 | 0.000 | -0.3139 | No | ||
| 32 | SKIL | 15617 | 11130 | -0.015 | -0.3863 | No | ||
| 33 | XPO1 | 4172 | 11401 | -0.018 | -0.4002 | No | ||
| 34 | RORC | 9732 | 12608 | -0.034 | -0.4640 | No | ||
| 35 | HCN1 | 21557 | 12616 | -0.034 | -0.4631 | No | ||
| 36 | CLDN17 | 10757 | 12809 | -0.037 | -0.4721 | No | ||
| 37 | TMEM27 | 24214 | 12839 | -0.038 | -0.4723 | No | ||
| 38 | SRPX | 11952 2630 | 13236 | -0.045 | -0.4919 | No | ||
| 39 | DLGAP4 | 2950 10461 | 13872 | -0.060 | -0.5239 | No | ||
| 40 | GABRE | 24140 | 13884 | -0.061 | -0.5223 | No | ||
| 41 | PELO | 8360 | 14521 | -0.082 | -0.5535 | No | ||
| 42 | NTRK3 | 2069 3652 3705 9492 | 15039 | -0.114 | -0.5772 | No | ||
| 43 | FERD3L | 21292 | 15279 | -0.134 | -0.5851 | No | ||
| 44 | MRPL45 | 20680 | 15436 | -0.151 | -0.5880 | No | ||
| 45 | EGF | 15169 | 15676 | -0.185 | -0.5941 | No | ||
| 46 | MEF2D | 9379 | 16193 | -0.303 | -0.6107 | No | ||
| 47 | CYR61 | 9159 15145 9311 5024 | 16397 | -0.368 | -0.6081 | No | ||
| 48 | MAP3K8 | 23495 | 17035 | -0.667 | -0.6179 | Yes | ||
| 49 | SLC6A9 | 4783 | 17560 | -1.042 | -0.6078 | Yes | ||
| 50 | HOXB4 | 20686 | 17584 | -1.062 | -0.5700 | Yes | ||
| 51 | MAP4K2 | 6457 | 17791 | -1.274 | -0.5343 | Yes | ||
| 52 | OGT | 4241 24274 | 17835 | -1.316 | -0.4882 | Yes | ||
| 53 | STIP1 | 23798 | 18183 | -1.756 | -0.4423 | Yes | ||
| 54 | CPLX2 | 8779 | 18596 | -4.366 | -0.3040 | Yes | ||
| 55 | SOCS2 | 5694 | 18614 | -8.294 | 0.0001 | Yes |