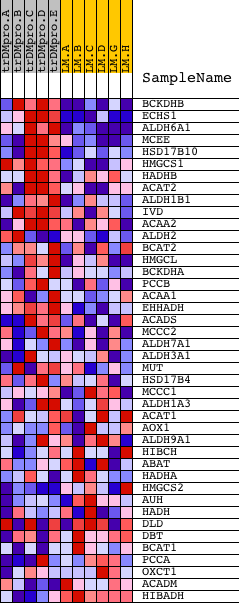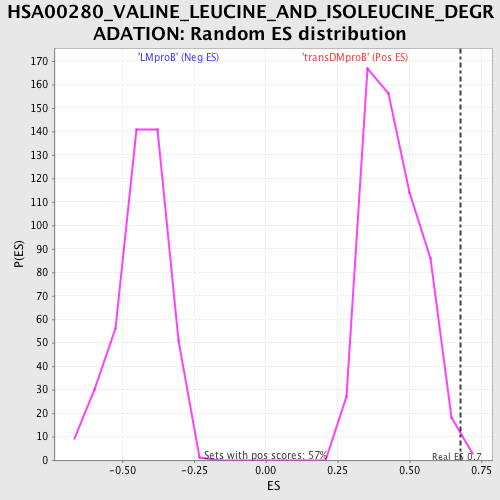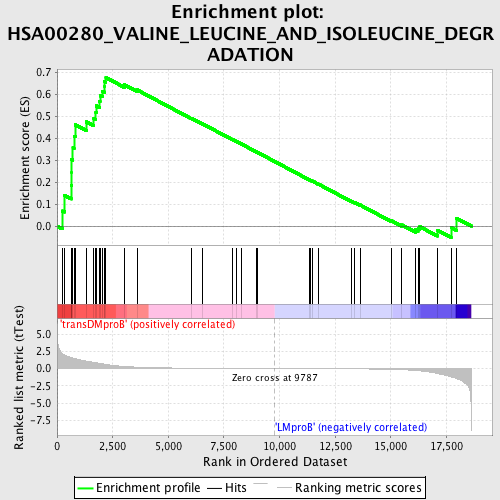
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | HSA00280_VALINE_LEUCINE_AND_ISOLEUCINE_DEGRADATION |
| Enrichment Score (ES) | 0.6762625 |
| Normalized Enrichment Score (NES) | 1.5235244 |
| Nominal p-value | 0.0052539404 |
| FDR q-value | 0.3751395 |
| FWER p-Value | 0.93 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | BCKDHB | 19365 | 242 | 2.161 | 0.0689 | Yes | ||
| 2 | ECHS1 | 17573 | 332 | 1.987 | 0.1395 | Yes | ||
| 3 | ALDH6A1 | 21025 | 631 | 1.601 | 0.1841 | Yes | ||
| 4 | MCEE | 18219 | 647 | 1.586 | 0.2435 | Yes | ||
| 5 | HSD17B10 | 24226 | 652 | 1.580 | 0.3032 | Yes | ||
| 6 | HMGCS1 | 9917 | 713 | 1.516 | 0.3575 | Yes | ||
| 7 | HADHB | 3514 10527 | 764 | 1.463 | 0.4102 | Yes | ||
| 8 | ACAT2 | 8484 1537 | 816 | 1.417 | 0.4612 | Yes | ||
| 9 | ALDH1B1 | 16219 | 1313 | 1.055 | 0.4746 | Yes | ||
| 10 | IVD | 7102 | 1650 | 0.891 | 0.4903 | Yes | ||
| 11 | ACAA2 | 1956 1981 23515 | 1741 | 0.833 | 0.5170 | Yes | ||
| 12 | ALDH2 | 16384 | 1758 | 0.825 | 0.5475 | Yes | ||
| 13 | BCAT2 | 18244 | 1900 | 0.746 | 0.5682 | Yes | ||
| 14 | HMGCL | 2538 16033 | 1931 | 0.734 | 0.5944 | Yes | ||
| 15 | BCKDHA | 17925 | 2027 | 0.678 | 0.6150 | Yes | ||
| 16 | PCCB | 19026 | 2120 | 0.634 | 0.6341 | Yes | ||
| 17 | ACAA1 | 4310 | 2127 | 0.628 | 0.6576 | Yes | ||
| 18 | EHHADH | 22635 | 2195 | 0.587 | 0.6763 | Yes | ||
| 19 | ACADS | 16415 | 3018 | 0.307 | 0.6436 | No | ||
| 20 | MCCC2 | 13422 | 3593 | 0.200 | 0.6203 | No | ||
| 21 | ALDH7A1 | 23435 | 6018 | 0.054 | 0.4918 | No | ||
| 22 | ALDH3A1 | 20854 | 6528 | 0.043 | 0.4661 | No | ||
| 23 | MUT | 9431 | 7889 | 0.021 | 0.3937 | No | ||
| 24 | HSD17B4 | 23560 | 8078 | 0.019 | 0.3842 | No | ||
| 25 | MCCC1 | 15359 | 8274 | 0.017 | 0.3744 | No | ||
| 26 | ALDH1A3 | 17802 | 8953 | 0.009 | 0.3382 | No | ||
| 27 | ACAT1 | 19114 10019 | 9011 | 0.008 | 0.3355 | No | ||
| 28 | AOX1 | 14246 4090 | 11327 | -0.017 | 0.2115 | No | ||
| 29 | ALDH9A1 | 14064 | 11384 | -0.017 | 0.2091 | No | ||
| 30 | HIBCH | 14250 | 11464 | -0.018 | 0.2056 | No | ||
| 31 | ABAT | 22864 1713 22669 | 11736 | -0.022 | 0.1918 | No | ||
| 32 | HADHA | 16579 | 13234 | -0.045 | 0.1129 | No | ||
| 33 | HMGCS2 | 15483 | 13380 | -0.048 | 0.1069 | No | ||
| 34 | AUH | 21464 | 13614 | -0.054 | 0.0964 | No | ||
| 35 | HADH | 9075 15165 | 15018 | -0.112 | 0.0251 | No | ||
| 36 | DLD | 2097 21090 | 15499 | -0.158 | 0.0053 | No | ||
| 37 | DBT | 1770 4599 | 16108 | -0.279 | -0.0168 | No | ||
| 38 | BCAT1 | 1042 4434 | 16223 | -0.312 | -0.0112 | No | ||
| 39 | PCCA | 21926 | 16306 | -0.335 | -0.0029 | No | ||
| 40 | OXCT1 | 22523 | 17092 | -0.707 | -0.0183 | No | ||
| 41 | ACADM | 15133 1800 | 17712 | -1.192 | -0.0064 | No | ||
| 42 | HIBADH | 17144 | 17956 | -1.452 | 0.0355 | No |