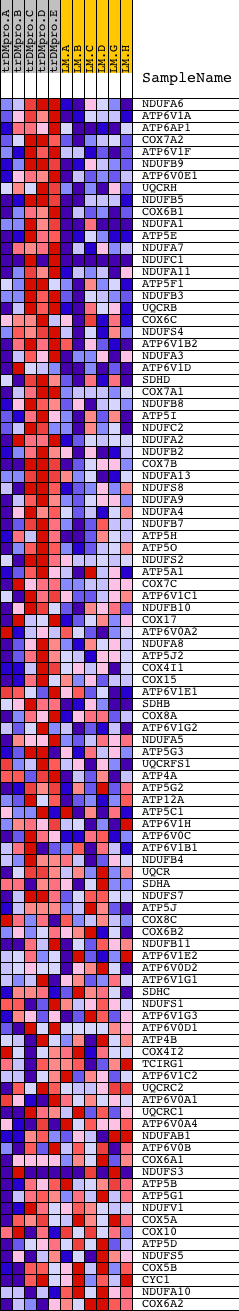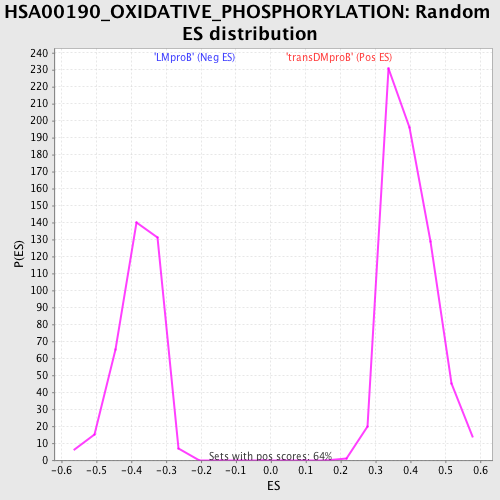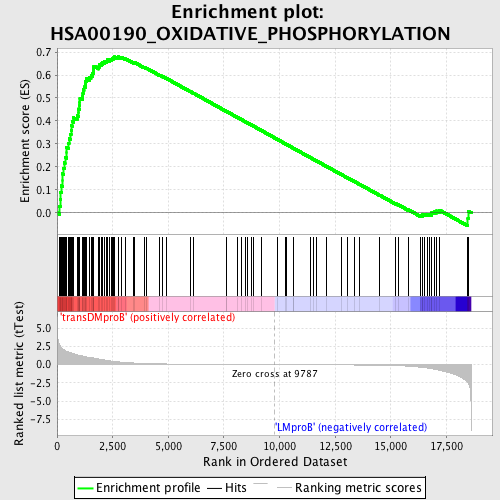
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | HSA00190_OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | 0.6790091 |
| Normalized Enrichment Score (NES) | 1.7151554 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.027461177 |
| FWER p-Value | 0.028 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NDUFA6 | 12473 | 107 | 2.667 | 0.0287 | Yes | ||
| 2 | ATP6V1A | 8638 | 142 | 2.504 | 0.0593 | Yes | ||
| 3 | ATP6AP1 | 24296 | 162 | 2.422 | 0.0897 | Yes | ||
| 4 | COX7A2 | 19053 | 194 | 2.275 | 0.1175 | Yes | ||
| 5 | ATP6V1F | 12291 | 250 | 2.140 | 0.1422 | Yes | ||
| 6 | NDUFB9 | 12306 | 252 | 2.133 | 0.1698 | Yes | ||
| 7 | ATP6V0E1 | 23326 | 296 | 2.045 | 0.1939 | Yes | ||
| 8 | UQCRH | 12378 | 330 | 1.989 | 0.2179 | Yes | ||
| 9 | NDUFB5 | 12277 | 357 | 1.941 | 0.2416 | Yes | ||
| 10 | COX6B1 | 4283 8479 | 425 | 1.813 | 0.2615 | Yes | ||
| 11 | NDUFA1 | 24170 | 427 | 1.811 | 0.2848 | Yes | ||
| 12 | ATP5E | 14321 | 506 | 1.707 | 0.3027 | Yes | ||
| 13 | NDUFA7 | 12347 | 537 | 1.673 | 0.3228 | Yes | ||
| 14 | NDUFC1 | 7287 12339 | 594 | 1.631 | 0.3409 | Yes | ||
| 15 | NDUFA11 | 12832 | 629 | 1.602 | 0.3598 | Yes | ||
| 16 | ATP5F1 | 15212 | 656 | 1.576 | 0.3788 | Yes | ||
| 17 | NDUFB3 | 12362 | 677 | 1.551 | 0.3978 | Yes | ||
| 18 | UQCRB | 12545 | 735 | 1.494 | 0.4140 | Yes | ||
| 19 | COX6C | 8775 | 900 | 1.344 | 0.4226 | Yes | ||
| 20 | NDUFS4 | 9452 5157 3173 | 969 | 1.290 | 0.4356 | Yes | ||
| 21 | ATP6V1B2 | 18599 | 973 | 1.288 | 0.4521 | Yes | ||
| 22 | NDUFA3 | 18410 | 1012 | 1.256 | 0.4664 | Yes | ||
| 23 | ATP6V1D | 7872 | 1021 | 1.251 | 0.4821 | Yes | ||
| 24 | SDHD | 19125 | 1026 | 1.248 | 0.4981 | Yes | ||
| 25 | COX7A1 | 4554 | 1141 | 1.175 | 0.5071 | Yes | ||
| 26 | NDUFB8 | 7418 | 1159 | 1.161 | 0.5212 | Yes | ||
| 27 | ATP5I | 8637 | 1170 | 1.149 | 0.5356 | Yes | ||
| 28 | NDUFC2 | 18183 | 1209 | 1.127 | 0.5481 | Yes | ||
| 29 | NDUFA2 | 9450 | 1265 | 1.089 | 0.5593 | Yes | ||
| 30 | NDUFB2 | 7542 | 1281 | 1.081 | 0.5724 | Yes | ||
| 31 | COX7B | 7260 | 1326 | 1.043 | 0.5836 | Yes | ||
| 32 | NDUFA13 | 7400 | 1451 | 0.990 | 0.5897 | Yes | ||
| 33 | NDUFS8 | 23950 | 1524 | 0.958 | 0.5982 | Yes | ||
| 34 | NDUFA9 | 12288 | 1595 | 0.923 | 0.6064 | Yes | ||
| 35 | NDUFA4 | 9451 | 1624 | 0.903 | 0.6165 | Yes | ||
| 36 | NDUFB7 | 12429 | 1640 | 0.896 | 0.6273 | Yes | ||
| 37 | ATP5H | 12948 | 1651 | 0.890 | 0.6383 | Yes | ||
| 38 | ATP5O | 22539 | 1878 | 0.758 | 0.6359 | Yes | ||
| 39 | NDUFS2 | 4089 13760 | 1905 | 0.742 | 0.6441 | Yes | ||
| 40 | ATP5A1 | 23505 | 1993 | 0.692 | 0.6484 | Yes | ||
| 41 | COX7C | 8776 | 2035 | 0.672 | 0.6549 | Yes | ||
| 42 | ATP6V1C1 | 22487 12329 | 2121 | 0.634 | 0.6585 | Yes | ||
| 43 | NDUFB10 | 23084 | 2227 | 0.572 | 0.6602 | Yes | ||
| 44 | COX17 | 467 | 2255 | 0.557 | 0.6660 | Yes | ||
| 45 | ATP6V0A2 | 5760 | 2360 | 0.518 | 0.6671 | Yes | ||
| 46 | NDUFA8 | 14605 | 2462 | 0.483 | 0.6679 | Yes | ||
| 47 | ATP5J2 | 12186 | 2483 | 0.476 | 0.6730 | Yes | ||
| 48 | COX4I1 | 18444 | 2550 | 0.449 | 0.6752 | Yes | ||
| 49 | COX15 | 5931 | 2588 | 0.431 | 0.6788 | Yes | ||
| 50 | ATP6V1E1 | 4423 | 2737 | 0.382 | 0.6757 | Yes | ||
| 51 | SDHB | 2348 12566 14880 | 2767 | 0.373 | 0.6790 | Yes | ||
| 52 | COX8A | 8777 | 2879 | 0.344 | 0.6775 | No | ||
| 53 | ATP6V1G2 | 23257 1534 | 3086 | 0.289 | 0.6701 | No | ||
| 54 | NDUFA5 | 1076 7544 | 3441 | 0.223 | 0.6538 | No | ||
| 55 | ATP5G3 | 14558 | 3463 | 0.219 | 0.6556 | No | ||
| 56 | UQCRFS1 | 21499 | 3927 | 0.158 | 0.6326 | No | ||
| 57 | ATP4A | 4422 | 4010 | 0.150 | 0.6301 | No | ||
| 58 | ATP5G2 | 12610 | 4623 | 0.105 | 0.5984 | No | ||
| 59 | ATP12A | 16700 | 4755 | 0.097 | 0.5926 | No | ||
| 60 | ATP5C1 | 8635 | 4914 | 0.089 | 0.5852 | No | ||
| 61 | ATP6V1H | 14301 | 5973 | 0.055 | 0.5288 | No | ||
| 62 | ATP6V0C | 8643 | 6122 | 0.051 | 0.5215 | No | ||
| 63 | ATP6V1B1 | 8499 | 7591 | 0.025 | 0.4425 | No | ||
| 64 | NDUFB4 | 22596 7541 | 7626 | 0.025 | 0.4410 | No | ||
| 65 | UQCR | 12382 | 8090 | 0.018 | 0.4162 | No | ||
| 66 | SDHA | 12436 | 8271 | 0.017 | 0.4067 | No | ||
| 67 | NDUFS7 | 19946 | 8476 | 0.014 | 0.3959 | No | ||
| 68 | ATP5J | 880 22555 | 8543 | 0.014 | 0.3925 | No | ||
| 69 | COX8C | 21175 | 8729 | 0.011 | 0.3826 | No | ||
| 70 | COX6B2 | 17983 4110 | 8836 | 0.010 | 0.3771 | No | ||
| 71 | NDUFB11 | 24176 | 9194 | 0.006 | 0.3579 | No | ||
| 72 | ATP6V1E2 | 22880 | 9886 | -0.001 | 0.3206 | No | ||
| 73 | ATP6V0D2 | 15934 | 10264 | -0.005 | 0.3003 | No | ||
| 74 | ATP6V1G1 | 12322 | 10294 | -0.006 | 0.2988 | No | ||
| 75 | SDHC | 7248 12279 | 10612 | -0.009 | 0.2818 | No | ||
| 76 | NDUFS1 | 13940 | 10640 | -0.009 | 0.2804 | No | ||
| 77 | ATP6V1G3 | 14107 | 11399 | -0.018 | 0.2397 | No | ||
| 78 | ATP6V0D1 | 3895 8639 3920 | 11540 | -0.019 | 0.2324 | No | ||
| 79 | ATP4B | 18928 | 11640 | -0.021 | 0.2273 | No | ||
| 80 | COX4I2 | 14791 | 12128 | -0.027 | 0.2014 | No | ||
| 81 | TCIRG1 | 3700 3752 23951 | 12762 | -0.036 | 0.1677 | No | ||
| 82 | ATP6V1C2 | 21098 | 13050 | -0.042 | 0.1527 | No | ||
| 83 | UQCRC2 | 18102 | 13386 | -0.048 | 0.1352 | No | ||
| 84 | ATP6V0A1 | 8640 4424 1197 | 13586 | -0.053 | 0.1252 | No | ||
| 85 | UQCRC1 | 10259 | 14490 | -0.081 | 0.0774 | No | ||
| 86 | ATP6V0A4 | 8934 | 15216 | -0.128 | 0.0400 | No | ||
| 87 | NDUFAB1 | 7667 | 15329 | -0.139 | 0.0357 | No | ||
| 88 | ATP6V0B | 15786 | 15794 | -0.208 | 0.0133 | No | ||
| 89 | COX6A1 | 8774 4553 | 16325 | -0.342 | -0.0109 | No | ||
| 90 | NDUFS3 | 7553 12683 | 16422 | -0.378 | -0.0112 | No | ||
| 91 | ATP5B | 19846 | 16431 | -0.383 | -0.0066 | No | ||
| 92 | ATP5G1 | 8636 | 16499 | -0.407 | -0.0050 | No | ||
| 93 | NDUFV1 | 23953 | 16637 | -0.452 | -0.0065 | No | ||
| 94 | COX5A | 19431 | 16734 | -0.506 | -0.0052 | No | ||
| 95 | COX10 | 20408 | 16825 | -0.545 | -0.0030 | No | ||
| 96 | ATP5D | 19949 | 16843 | -0.556 | 0.0033 | No | ||
| 97 | NDUFS5 | 9296 | 16976 | -0.630 | 0.0043 | No | ||
| 98 | COX5B | 4552 | 17069 | -0.693 | 0.0083 | No | ||
| 99 | CYC1 | 12348 | 17188 | -0.772 | 0.0120 | No | ||
| 100 | NDUFA10 | 7420 | 18465 | -2.476 | -0.0249 | No | ||
| 101 | COX6A2 | 17605 | 18485 | -2.548 | 0.0071 | No |