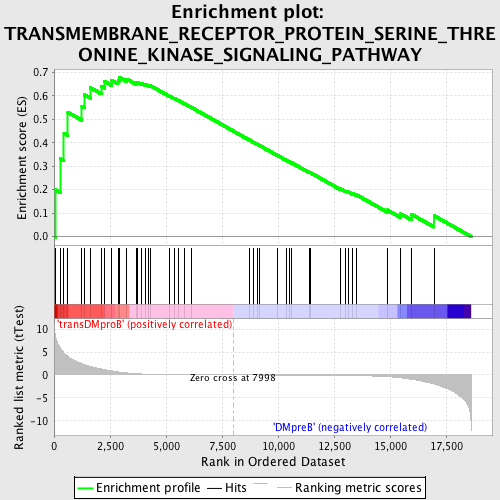
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | TRANSMEMBRANE_RECEPTOR_PROTEIN_SERINE_THREONINE_KINASE_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.6778209 |
| Normalized Enrichment Score (NES) | 1.5380864 |
| Nominal p-value | 0.008333334 |
| FDR q-value | 0.3962696 |
| FWER p-Value | 0.993 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ENG | 4667 | 66 | 8.260 | 0.1993 | Yes | ||
| 2 | SMAD5 | 21621 | 269 | 5.871 | 0.3327 | Yes | ||
| 3 | SMAD3 | 19084 | 440 | 4.748 | 0.4401 | Yes | ||
| 4 | TGFB1 | 18332 | 616 | 4.024 | 0.5296 | Yes | ||
| 5 | SMAD4 | 5058 | 1242 | 2.437 | 0.5558 | Yes | ||
| 6 | LEFTY1 | 8877 | 1356 | 2.217 | 0.6041 | Yes | ||
| 7 | ACVR1 | 4334 | 1627 | 1.818 | 0.6343 | Yes | ||
| 8 | CDKN1C | 17546 | 2111 | 1.301 | 0.6402 | Yes | ||
| 9 | FNTA | 18904 | 2254 | 1.160 | 0.6611 | Yes | ||
| 10 | HIPK2 | 17180 | 2574 | 0.899 | 0.6660 | Yes | ||
| 11 | ACVR1B | 4335 22353 | 2853 | 0.647 | 0.6669 | Yes | ||
| 12 | MAP3K7 | 16255 | 2925 | 0.600 | 0.6778 | Yes | ||
| 13 | ZFYVE9 | 10501 2449 | 3239 | 0.432 | 0.6716 | No | ||
| 14 | ACVR2B | 19276 | 3662 | 0.266 | 0.6554 | No | ||
| 15 | SMAD2 | 23511 | 3741 | 0.245 | 0.6572 | No | ||
| 16 | GDF9 | 20887 | 3908 | 0.202 | 0.6533 | No | ||
| 17 | ACVR2A | 8542 | 4076 | 0.167 | 0.6484 | No | ||
| 18 | BMPR2 | 14241 4081 | 4206 | 0.147 | 0.6450 | No | ||
| 19 | CIDEA | 23529 | 4299 | 0.134 | 0.6434 | No | ||
| 20 | ACVRL1 | 22354 | 5145 | 0.061 | 0.5994 | No | ||
| 21 | TGFBR1 | 16213 10165 5745 | 5354 | 0.052 | 0.5895 | No | ||
| 22 | TGFBR3 | 16453 | 5532 | 0.045 | 0.5810 | No | ||
| 23 | KLF10 | 22308 | 5836 | 0.037 | 0.5656 | No | ||
| 24 | SOST | 20210 | 6128 | 0.030 | 0.5507 | No | ||
| 25 | FMOD | 14133 | 8740 | -0.010 | 0.4104 | No | ||
| 26 | SMAD7 | 23513 | 8883 | -0.012 | 0.4030 | No | ||
| 27 | RGMB | 12736 7585 | 9058 | -0.015 | 0.3940 | No | ||
| 28 | IL17F | 13992 | 9150 | -0.016 | 0.3895 | No | ||
| 29 | GDF5 | 4768 | 9949 | -0.028 | 0.3472 | No | ||
| 30 | LEFTY2 | 6735 | 10359 | -0.035 | 0.3260 | No | ||
| 31 | FOXH1 | 22238 | 10491 | -0.037 | 0.3199 | No | ||
| 32 | CITED2 | 5118 14477 | 10616 | -0.039 | 0.3142 | No | ||
| 33 | SMURF1 | 13289 7989 | 11408 | -0.055 | 0.2729 | No | ||
| 34 | BMPR1A | 8660 4450 | 11434 | -0.056 | 0.2730 | No | ||
| 35 | GDF10 | 4767 | 12772 | -0.106 | 0.2036 | No | ||
| 36 | GDF15 | 10721 | 13018 | -0.120 | 0.1933 | No | ||
| 37 | SNX6 | 21070 | 13131 | -0.127 | 0.1904 | No | ||
| 38 | EID2 | 11815 | 13306 | -0.140 | 0.1845 | No | ||
| 39 | HPGD | 4867 | 13497 | -0.156 | 0.1781 | No | ||
| 40 | SMAD1 | 18831 922 | 14865 | -0.374 | 0.1137 | No | ||
| 41 | STUB1 | 23070 | 15472 | -0.640 | 0.0968 | No | ||
| 42 | LTBP2 | 5007 | 15951 | -0.981 | 0.0951 | No | ||
| 43 | CDKN2B | 15840 | 16958 | -1.965 | 0.0893 | No |

