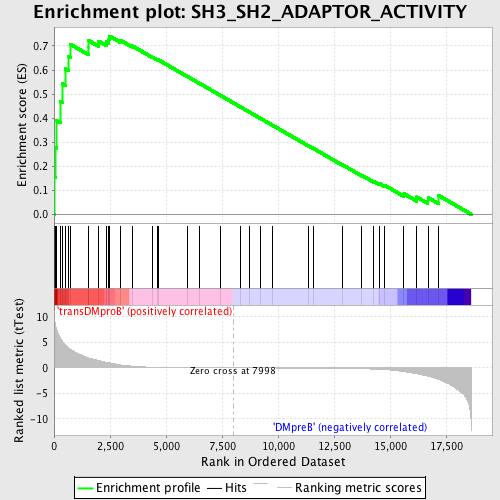
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | SH3_SH2_ADAPTOR_ACTIVITY |
| Enrichment Score (ES) | 0.7409671 |
| Normalized Enrichment Score (NES) | 1.6376777 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.30632693 |
| FWER p-Value | 0.558 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GRB7 | 20673 | 14 | 10.160 | 0.1542 | Yes | ||
| 2 | SOCS2 | 5694 | 65 | 8.301 | 0.2781 | Yes | ||
| 3 | LAT | 17643 | 116 | 7.434 | 0.3888 | Yes | ||
| 4 | GAB1 | 18828 | 276 | 5.806 | 0.4687 | Yes | ||
| 5 | SH3BP2 | 16874 | 360 | 5.199 | 0.5436 | Yes | ||
| 6 | SH2D2A | 6569 | 501 | 4.488 | 0.6045 | Yes | ||
| 7 | SH3BGRL | 24267 | 654 | 3.864 | 0.6552 | Yes | ||
| 8 | ARHGAP4 | 9343 | 716 | 3.651 | 0.7076 | Yes | ||
| 9 | CRKL | 4560 | 1532 | 1.945 | 0.6934 | Yes | ||
| 10 | GRB2 | 20149 | 1534 | 1.943 | 0.7230 | Yes | ||
| 11 | STAM | 2912 15117 | 1995 | 1.404 | 0.7196 | Yes | ||
| 12 | ITSN2 | 21327 | 2346 | 1.082 | 0.7173 | Yes | ||
| 13 | SH2D3C | 2864 | 2446 | 1.000 | 0.7272 | Yes | ||
| 14 | LASP1 | 4986 4985 1248 | 2473 | 0.995 | 0.7410 | Yes | ||
| 15 | SHB | 10493 | 2944 | 0.587 | 0.7246 | No | ||
| 16 | CHN2 | 7652 | 3493 | 0.323 | 0.7001 | No | ||
| 17 | GRB14 | 14574 2719 | 4412 | 0.117 | 0.6524 | No | ||
| 18 | ARHGAP6 | 24207 | 4626 | 0.093 | 0.6424 | No | ||
| 19 | ABI2 | 4037 6830 | 4651 | 0.091 | 0.6425 | No | ||
| 20 | GRAP2 | 5113 9398 | 5961 | 0.034 | 0.5726 | No | ||
| 21 | CHN1 | 4246 | 6478 | 0.023 | 0.5451 | No | ||
| 22 | IRS4 | 9183 4926 | 7411 | 0.008 | 0.4951 | No | ||
| 23 | SH3BGR | 11937 | 8311 | -0.004 | 0.4468 | No | ||
| 24 | SKAP2 | 17153 | 8741 | -0.010 | 0.4238 | No | ||
| 25 | SLA | 5449 | 9234 | -0.017 | 0.3976 | No | ||
| 26 | RUSC1 | 1853 15283 | 9739 | -0.024 | 0.3709 | No | ||
| 27 | SRC | 5507 | 11343 | -0.054 | 0.2854 | No | ||
| 28 | GRB10 | 4799 | 11590 | -0.060 | 0.2731 | No | ||
| 29 | SNX9 | 23132 | 12867 | -0.111 | 0.2061 | No | ||
| 30 | ARHGAP1 | 6001 10448 | 13740 | -0.179 | 0.1619 | No | ||
| 31 | CNTNAP1 | 11987 | 14259 | -0.242 | 0.1377 | No | ||
| 32 | KHDRBS1 | 5405 9778 | 14513 | -0.285 | 0.1284 | No | ||
| 33 | VAV3 | 1774 1848 15443 | 14752 | -0.346 | 0.1209 | No | ||
| 34 | GRAP | 20851 | 15618 | -0.722 | 0.0854 | No | ||
| 35 | BLNK | 23681 3691 | 16184 | -1.124 | 0.0721 | No | ||
| 36 | EPS8 | 16948 | 16690 | -1.625 | 0.0697 | No | ||
| 37 | TOB1 | 20703 | 17150 | -2.224 | 0.0789 | No |

