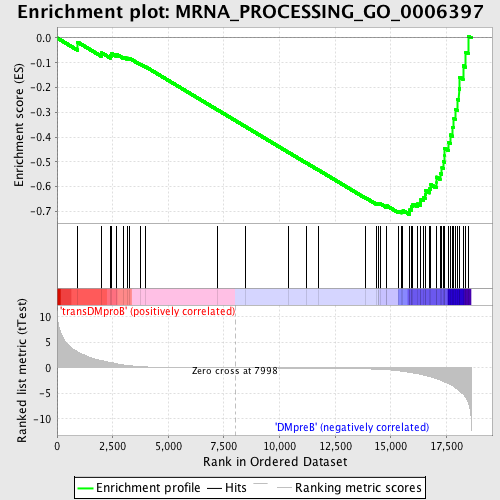
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | MRNA_PROCESSING_GO_0006397 |
| Enrichment Score (ES) | -0.7120557 |
| Normalized Enrichment Score (NES) | -1.6536324 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.021176387 |
| FWER p-Value | 0.32 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPARGC1A | 16533 | 925 | 3.119 | -0.0170 | No | ||
| 2 | SFRS5 | 9808 2062 | 1983 | 1.425 | -0.0589 | No | ||
| 3 | SFRS7 | 22889 | 2401 | 1.027 | -0.0705 | No | ||
| 4 | SFRS10 | 9818 1722 5441 | 2457 | 0.997 | -0.0630 | No | ||
| 5 | SFPQ | 12936 | 2659 | 0.820 | -0.0652 | No | ||
| 6 | KIN | 15120 | 2993 | 0.557 | -0.0773 | No | ||
| 7 | SFRS8 | 10543 6089 | 3152 | 0.468 | -0.0809 | No | ||
| 8 | SPOP | 5498 | 3270 | 0.417 | -0.0828 | No | ||
| 9 | RNGTT | 10769 2354 6272 | 3737 | 0.247 | -0.1053 | No | ||
| 10 | ELAVL4 | 15805 4889 9137 | 3984 | 0.185 | -0.1166 | No | ||
| 11 | CSTF2 | 24257 | 7209 | 0.011 | -0.2902 | No | ||
| 12 | CUGBP1 | 2805 8819 4576 2924 | 8484 | -0.007 | -0.3587 | No | ||
| 13 | LSM5 | 7286 | 10421 | -0.036 | -0.4627 | No | ||
| 14 | CPSF3 | 7038 | 11212 | -0.051 | -0.5047 | No | ||
| 15 | SLBP | 9822 | 11754 | -0.064 | -0.5332 | No | ||
| 16 | SFRS1 | 8492 | 13847 | -0.190 | -0.6439 | No | ||
| 17 | SRPK2 | 5513 | 14351 | -0.255 | -0.6683 | No | ||
| 18 | SFRS2IP | 7794 13009 | 14427 | -0.269 | -0.6695 | No | ||
| 19 | NONO | 11993 | 14460 | -0.275 | -0.6683 | No | ||
| 20 | KHDRBS1 | 5405 9778 | 14513 | -0.285 | -0.6681 | No | ||
| 21 | PTBP1 | 5303 | 14798 | -0.355 | -0.6797 | No | ||
| 22 | SFRS2 | 9807 20136 | 14825 | -0.363 | -0.6773 | No | ||
| 23 | PABPC1 | 5219 9522 9523 23572 | 15334 | -0.551 | -0.6988 | No | ||
| 24 | CDC2L2 | 15964 2419 | 15470 | -0.639 | -0.6994 | No | ||
| 25 | PRPF31 | 7594 | 15546 | -0.677 | -0.6963 | No | ||
| 26 | SFRS9 | 16731 | 15840 | -0.887 | -0.7027 | Yes | ||
| 27 | CRNKL1 | 14407 | 15850 | -0.893 | -0.6938 | Yes | ||
| 28 | CSTF3 | 10449 6002 | 15917 | -0.952 | -0.6873 | Yes | ||
| 29 | SNW1 | 7282 | 15941 | -0.972 | -0.6783 | Yes | ||
| 30 | GEMIN6 | 12492 7415 | 15989 | -0.998 | -0.6703 | Yes | ||
| 31 | DHX38 | 48 | 16196 | -1.137 | -0.6694 | Yes | ||
| 32 | NUDT21 | 12665 | 16330 | -1.253 | -0.6634 | Yes | ||
| 33 | GEMIN7 | 17944 | 16349 | -1.275 | -0.6510 | Yes | ||
| 34 | DHX15 | 8842 | 16474 | -1.430 | -0.6426 | Yes | ||
| 35 | RNMT | 7501 | 16547 | -1.511 | -0.6305 | Yes | ||
| 36 | SF3A1 | 7450 | 16568 | -1.527 | -0.6155 | Yes | ||
| 37 | TXNL4A | 6567 11329 | 16753 | -1.689 | -0.6076 | Yes | ||
| 38 | SF3A3 | 16091 | 16797 | -1.744 | -0.5916 | Yes | ||
| 39 | CPSF1 | 22241 | 17044 | -2.094 | -0.5828 | Yes | ||
| 40 | CSTF1 | 12515 | 17048 | -2.098 | -0.5608 | Yes | ||
| 41 | SIP1 | 21263 | 17216 | -2.327 | -0.5453 | Yes | ||
| 42 | SF3A2 | 19938 | 17297 | -2.514 | -0.5231 | Yes | ||
| 43 | DDX39 | 18551 | 17356 | -2.632 | -0.4985 | Yes | ||
| 44 | DDX20 | 15213 | 17409 | -2.718 | -0.4727 | Yes | ||
| 45 | SFRS6 | 14751 | 17415 | -2.724 | -0.4442 | Yes | ||
| 46 | PHF5A | 22194 | 17601 | -3.077 | -0.4218 | Yes | ||
| 47 | FUSIP1 | 4715 16036 | 17671 | -3.254 | -0.3912 | Yes | ||
| 48 | GEMIN5 | 20439 | 17765 | -3.478 | -0.3596 | Yes | ||
| 49 | LSM3 | 12565 | 17832 | -3.644 | -0.3247 | Yes | ||
| 50 | SF3B3 | 18746 | 17915 | -3.934 | -0.2877 | Yes | ||
| 51 | SNRPD1 | 23622 | 17992 | -4.171 | -0.2478 | Yes | ||
| 52 | SRPK1 | 23049 | 18064 | -4.455 | -0.2047 | Yes | ||
| 53 | SNRPD2 | 8412 | 18097 | -4.591 | -0.1581 | Yes | ||
| 54 | USP39 | 1116 1083 11373 | 18250 | -5.095 | -0.1126 | Yes | ||
| 55 | EFTUD2 | 1219 20203 | 18369 | -5.754 | -0.0583 | Yes | ||
| 56 | GRSF1 | 16487 | 18479 | -6.791 | 0.0074 | Yes |

