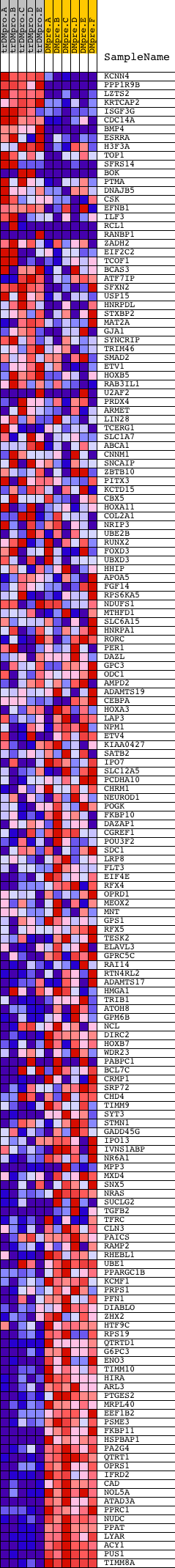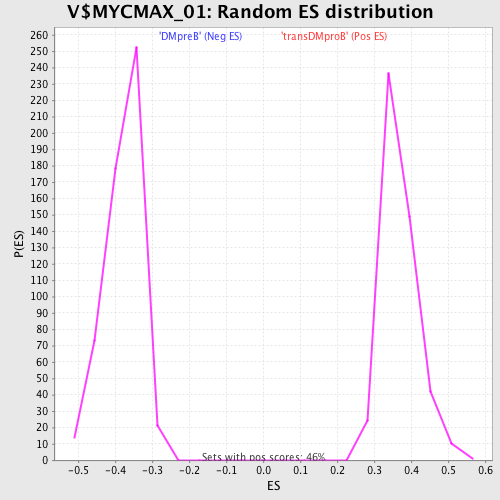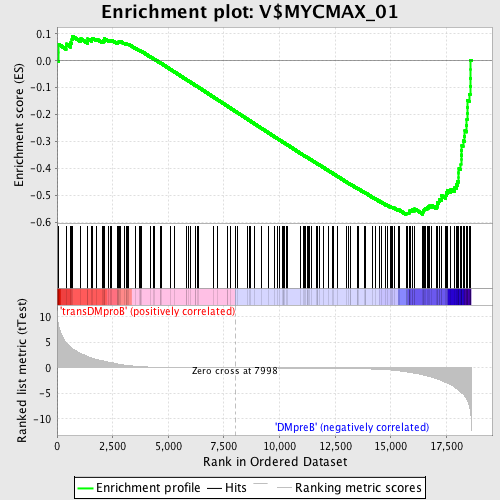
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | V$MYCMAX_01 |
| Enrichment Score (ES) | -0.5711955 |
| Normalized Enrichment Score (NES) | -1.5180612 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.20293266 |
| FWER p-Value | 0.726 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KCNN4 | 18353 | 53 | 8.508 | 0.0297 | No | ||
| 2 | PPP1R9B | 20698 | 54 | 8.508 | 0.0622 | No | ||
| 3 | LZTS2 | 10362 | 401 | 4.974 | 0.0624 | No | ||
| 4 | KRTCAP2 | 15541 | 614 | 4.026 | 0.0663 | No | ||
| 5 | ISGF3G | 2686 9184 | 666 | 3.807 | 0.0781 | No | ||
| 6 | CDC14A | 15184 | 682 | 3.756 | 0.0917 | No | ||
| 7 | BMP4 | 21863 | 1070 | 2.788 | 0.0814 | No | ||
| 8 | ESRRA | 23802 | 1372 | 2.195 | 0.0734 | No | ||
| 9 | H3F3A | 4838 | 1384 | 2.174 | 0.0811 | No | ||
| 10 | TOP1 | 5790 5789 10210 | 1548 | 1.918 | 0.0796 | No | ||
| 11 | SFRS14 | 10622 | 1590 | 1.875 | 0.0846 | No | ||
| 12 | BOK | 11953 14174 | 1776 | 1.639 | 0.0808 | No | ||
| 13 | PTMA | 9657 | 2030 | 1.375 | 0.0724 | No | ||
| 14 | DNAJB5 | 16236 | 2083 | 1.327 | 0.0746 | No | ||
| 15 | CSK | 8805 | 2115 | 1.299 | 0.0779 | No | ||
| 16 | EFNB1 | 24285 | 2130 | 1.288 | 0.0821 | No | ||
| 17 | ILF3 | 3110 3030 9176 | 2324 | 1.096 | 0.0758 | No | ||
| 18 | RCL1 | 3728 3743 23894 | 2393 | 1.037 | 0.0761 | No | ||
| 19 | RANBP1 | 9692 5357 | 2445 | 1.000 | 0.0771 | No | ||
| 20 | ZADH2 | 23501 | 2700 | 0.782 | 0.0663 | No | ||
| 21 | EIF2C2 | 6247 | 2735 | 0.755 | 0.0674 | No | ||
| 22 | TCOF1 | 5660 | 2747 | 0.742 | 0.0696 | No | ||
| 23 | BCAS3 | 5314 | 2779 | 0.711 | 0.0707 | No | ||
| 24 | ATF7IP | 12028 13038 | 2811 | 0.682 | 0.0716 | No | ||
| 25 | SFXN2 | 3707 8255 3678 13588 | 2867 | 0.639 | 0.0711 | No | ||
| 26 | USP15 | 4758 3377 | 3023 | 0.544 | 0.0647 | No | ||
| 27 | HNRPDL | 11947 3551 6954 | 3121 | 0.490 | 0.0614 | No | ||
| 28 | STXBP2 | 18700 | 3131 | 0.482 | 0.0627 | No | ||
| 29 | MAT2A | 10553 10554 6099 | 3178 | 0.458 | 0.0620 | No | ||
| 30 | GJA1 | 20023 | 3216 | 0.444 | 0.0617 | No | ||
| 31 | SYNCRIP | 3078 3035 3107 | 3530 | 0.309 | 0.0459 | No | ||
| 32 | TRIM46 | 1779 15281 | 3685 | 0.260 | 0.0385 | No | ||
| 33 | SMAD2 | 23511 | 3741 | 0.245 | 0.0365 | No | ||
| 34 | ETV1 | 4688 8925 8924 | 3755 | 0.241 | 0.0367 | No | ||
| 35 | HOXB5 | 20687 | 3804 | 0.229 | 0.0350 | No | ||
| 36 | RAB3IL1 | 3723 13229 | 4203 | 0.147 | 0.0139 | No | ||
| 37 | U2AF2 | 5813 5812 5814 | 4319 | 0.132 | 0.0082 | No | ||
| 38 | PRDX4 | 24026 | 4392 | 0.120 | 0.0048 | No | ||
| 39 | ARMET | 19005 | 4660 | 0.090 | -0.0094 | No | ||
| 40 | LIN28 | 15723 | 4666 | 0.090 | -0.0093 | No | ||
| 41 | TCERG1 | 12075 2027 7068 | 4677 | 0.088 | -0.0095 | No | ||
| 42 | SLC1A7 | 16147 | 5106 | 0.063 | -0.0324 | No | ||
| 43 | ABCA1 | 15881 | 5294 | 0.054 | -0.0424 | No | ||
| 44 | CNNM1 | 23846 | 5793 | 0.038 | -0.0692 | No | ||
| 45 | SNCAIP | 12589 | 5815 | 0.037 | -0.0702 | No | ||
| 46 | ZBTB10 | 10468 | 5916 | 0.035 | -0.0755 | No | ||
| 47 | PITX3 | 9571 | 6005 | 0.033 | -0.0802 | No | ||
| 48 | KCTD15 | 17864 | 6209 | 0.028 | -0.0911 | No | ||
| 49 | CBX5 | 22108 4485 | 6228 | 0.028 | -0.0919 | No | ||
| 50 | HOXA11 | 17146 | 6300 | 0.026 | -0.0957 | No | ||
| 51 | COL2A1 | 22145 4542 | 6347 | 0.025 | -0.0981 | No | ||
| 52 | NRIP3 | 17679 | 7020 | 0.014 | -0.1344 | No | ||
| 53 | UBE2B | 10245 | 7199 | 0.011 | -0.1441 | No | ||
| 54 | RUNX2 | 4480 8700 | 7650 | 0.004 | -0.1684 | No | ||
| 55 | FOXD3 | 9088 | 7667 | 0.004 | -0.1693 | No | ||
| 56 | UBXD3 | 15701 | 7805 | 0.003 | -0.1767 | No | ||
| 57 | HHIP | 4849 | 8030 | -0.000 | -0.1888 | No | ||
| 58 | APOA5 | 19469 | 8039 | -0.000 | -0.1893 | No | ||
| 59 | FGF14 | 21716 3005 3463 | 8086 | -0.001 | -0.1918 | No | ||
| 60 | RPS6KA5 | 2076 21005 | 8560 | -0.007 | -0.2174 | No | ||
| 61 | NDUFS1 | 13940 | 8627 | -0.008 | -0.2209 | No | ||
| 62 | MTHFD1 | 2132 21238 | 8706 | -0.010 | -0.2251 | No | ||
| 63 | SLC6A15 | 8335 3321 | 8861 | -0.012 | -0.2334 | No | ||
| 64 | HNRPA1 | 9103 4860 | 9192 | -0.016 | -0.2512 | No | ||
| 65 | RORC | 9732 | 9496 | -0.021 | -0.2676 | No | ||
| 66 | PER1 | 20827 | 9750 | -0.024 | -0.2812 | No | ||
| 67 | DAZL | 22944 | 9900 | -0.027 | -0.2892 | No | ||
| 68 | GPC3 | 24151 | 9984 | -0.028 | -0.2936 | No | ||
| 69 | ODC1 | 9500 | 10123 | -0.031 | -0.3009 | No | ||
| 70 | AMPD2 | 1931 15197 | 10189 | -0.032 | -0.3043 | No | ||
| 71 | ADAMTS19 | 23544 | 10193 | -0.032 | -0.3044 | No | ||
| 72 | CEBPA | 4514 | 10210 | -0.032 | -0.3051 | No | ||
| 73 | HOXA3 | 9108 1075 | 10322 | -0.034 | -0.3110 | No | ||
| 74 | LAP3 | 12442 | 10339 | -0.034 | -0.3118 | No | ||
| 75 | NPM1 | 1196 | 10941 | -0.045 | -0.3442 | No | ||
| 76 | ETV4 | 20212 | 11080 | -0.048 | -0.3515 | No | ||
| 77 | KIAA0427 | 11228 | 11122 | -0.049 | -0.3535 | No | ||
| 78 | SATB2 | 5604 | 11162 | -0.050 | -0.3554 | No | ||
| 79 | IPO7 | 6130 | 11240 | -0.051 | -0.3594 | No | ||
| 80 | SLC12A5 | 14731 | 11255 | -0.052 | -0.3600 | No | ||
| 81 | PCDHA10 | 8792 | 11293 | -0.053 | -0.3618 | No | ||
| 82 | CHRM1 | 23943 | 11350 | -0.054 | -0.3646 | No | ||
| 83 | NEUROD1 | 14550 | 11440 | -0.056 | -0.3692 | No | ||
| 84 | POGK | 7739 | 11666 | -0.062 | -0.3812 | No | ||
| 85 | FKBP10 | 20667 | 11705 | -0.063 | -0.3830 | No | ||
| 86 | DAZAP1 | 19945 | 11720 | -0.063 | -0.3835 | No | ||
| 87 | CGREF1 | 16576 | 11780 | -0.065 | -0.3865 | No | ||
| 88 | POU3F2 | 15931 | 11984 | -0.072 | -0.3972 | No | ||
| 89 | SDC1 | 5560 9953 | 12186 | -0.079 | -0.4078 | No | ||
| 90 | LRP8 | 16149 2425 | 12356 | -0.087 | -0.4166 | No | ||
| 91 | FLT3 | 16288 | 12437 | -0.091 | -0.4206 | No | ||
| 92 | EIF4E | 15403 1827 8890 | 12621 | -0.098 | -0.4301 | No | ||
| 93 | RFX4 | 7721 3410 | 13004 | -0.120 | -0.4504 | No | ||
| 94 | OPRD1 | 15737 | 13108 | -0.125 | -0.4555 | No | ||
| 95 | MEOX2 | 21285 | 13171 | -0.130 | -0.4584 | No | ||
| 96 | MNT | 20784 | 13174 | -0.130 | -0.4580 | No | ||
| 97 | GPS1 | 1392 9943 5550 9944 | 13481 | -0.154 | -0.4740 | No | ||
| 98 | RFX5 | 7018 | 13521 | -0.158 | -0.4755 | No | ||
| 99 | TESK2 | 10504 | 13544 | -0.160 | -0.4761 | No | ||
| 100 | ELAVL3 | 9136 | 13814 | -0.186 | -0.4899 | No | ||
| 101 | GPRC5C | 12864 | 13855 | -0.191 | -0.4914 | No | ||
| 102 | RAI14 | 22327 2191 | 13866 | -0.191 | -0.4912 | No | ||
| 103 | RTN4RL2 | 14544 | 14178 | -0.230 | -0.5072 | No | ||
| 104 | ADAMTS17 | 598 | 14288 | -0.245 | -0.5121 | No | ||
| 105 | HMGA1 | 23323 | 14469 | -0.276 | -0.5208 | No | ||
| 106 | TRIB1 | 22467 | 14563 | -0.295 | -0.5248 | No | ||
| 107 | ATOH8 | 17120 | 14778 | -0.351 | -0.5350 | No | ||
| 108 | GPM6B | 9034 | 14849 | -0.369 | -0.5374 | No | ||
| 109 | NCL | 5153 13899 | 14974 | -0.407 | -0.5426 | No | ||
| 110 | DIRC2 | 22603 | 15024 | -0.423 | -0.5436 | No | ||
| 111 | HOXB7 | 666 9110 | 15082 | -0.446 | -0.5450 | No | ||
| 112 | WDR23 | 22009 | 15153 | -0.475 | -0.5470 | No | ||
| 113 | PABPC1 | 5219 9522 9523 23572 | 15334 | -0.551 | -0.5546 | No | ||
| 114 | BCL7C | 4442 | 15367 | -0.570 | -0.5542 | No | ||
| 115 | CRMP1 | 16860 | 15682 | -0.767 | -0.5683 | Yes | ||
| 116 | SRP72 | 16819 | 15705 | -0.787 | -0.5664 | Yes | ||
| 117 | CHD4 | 8418 17281 4225 | 15754 | -0.826 | -0.5659 | Yes | ||
| 118 | TIMM9 | 19686 | 15817 | -0.873 | -0.5659 | Yes | ||
| 119 | SYT3 | 18264 | 15819 | -0.874 | -0.5626 | Yes | ||
| 120 | STMN1 | 9261 | 15852 | -0.895 | -0.5609 | Yes | ||
| 121 | GADD45G | 21637 | 15858 | -0.901 | -0.5578 | Yes | ||
| 122 | IPO13 | 15784 | 15881 | -0.920 | -0.5554 | Yes | ||
| 123 | IVNS1ABP | 4384 4050 | 15957 | -0.984 | -0.5558 | Yes | ||
| 124 | NR6A1 | 14596 | 15971 | -0.993 | -0.5527 | Yes | ||
| 125 | MPP3 | 8853 4630 5334 | 16077 | -1.038 | -0.5544 | Yes | ||
| 126 | MXD4 | 9346 | 16082 | -1.040 | -0.5506 | Yes | ||
| 127 | SNX5 | 12783 14410 | 16441 | -1.378 | -0.5648 | Yes | ||
| 128 | NRAS | 5191 | 16449 | -1.395 | -0.5598 | Yes | ||
| 129 | SUCLG2 | 17062 5545 | 16480 | -1.436 | -0.5559 | Yes | ||
| 130 | TGFB2 | 5743 | 16495 | -1.457 | -0.5511 | Yes | ||
| 131 | TFRC | 22785 | 16555 | -1.516 | -0.5485 | Yes | ||
| 132 | CLN3 | 17635 | 16628 | -1.574 | -0.5464 | Yes | ||
| 133 | PAICS | 16820 | 16678 | -1.613 | -0.5429 | Yes | ||
| 134 | RAMP2 | 20660 | 16733 | -1.668 | -0.5395 | Yes | ||
| 135 | RHEBL1 | 12781 | 16845 | -1.806 | -0.5386 | Yes | ||
| 136 | UBE1 | 24370 2551 | 17042 | -2.087 | -0.5412 | Yes | ||
| 137 | PPARGC1B | 9318 | 17086 | -2.155 | -0.5353 | Yes | ||
| 138 | KCMF1 | 17112 | 17100 | -2.175 | -0.5277 | Yes | ||
| 139 | PRPS1 | 24233 | 17165 | -2.238 | -0.5226 | Yes | ||
| 140 | PFN1 | 9555 | 17197 | -2.289 | -0.5155 | Yes | ||
| 141 | DIABLO | 16375 | 17272 | -2.465 | -0.5101 | Yes | ||
| 142 | ZHX2 | 11842 | 17273 | -2.466 | -0.5007 | Yes | ||
| 143 | HTF9C | 4886 1678 | 17479 | -2.864 | -0.5009 | Yes | ||
| 144 | RPS19 | 5398 | 17490 | -2.882 | -0.4904 | Yes | ||
| 145 | QTRTD1 | 22590 | 17555 | -2.998 | -0.4824 | Yes | ||
| 146 | G6PC3 | 1221 20645 | 17699 | -3.328 | -0.4774 | Yes | ||
| 147 | ENO3 | 8905 | 17874 | -3.773 | -0.4724 | Yes | ||
| 148 | TIMM10 | 11390 2158 | 17944 | -4.026 | -0.4608 | Yes | ||
| 149 | HIRA | 4852 9090 | 18002 | -4.201 | -0.4478 | Yes | ||
| 150 | ARL3 | 23652 | 18057 | -4.429 | -0.4338 | Yes | ||
| 151 | PTGES2 | 15038 | 18059 | -4.433 | -0.4169 | Yes | ||
| 152 | MRPL40 | 22641 | 18062 | -4.439 | -0.4001 | Yes | ||
| 153 | EEF1B2 | 4131 12063 | 18152 | -4.785 | -0.3866 | Yes | ||
| 154 | PSME3 | 20657 | 18155 | -4.800 | -0.3683 | Yes | ||
| 155 | FKBP11 | 22136 | 18158 | -4.815 | -0.3500 | Yes | ||
| 156 | HSPBAP1 | 7321 | 18172 | -4.865 | -0.3322 | Yes | ||
| 157 | PA2G4 | 5265 | 18195 | -4.947 | -0.3144 | Yes | ||
| 158 | QTRT1 | 19540 | 18247 | -5.085 | -0.2978 | Yes | ||
| 159 | OPRS1 | 9515 2402 | 18303 | -5.345 | -0.2803 | Yes | ||
| 160 | IFRD2 | 19320 | 18331 | -5.544 | -0.2606 | Yes | ||
| 161 | CAD | 16886 | 18408 | -6.035 | -0.2416 | Yes | ||
| 162 | NOL5A | 12474 | 18420 | -6.093 | -0.2189 | Yes | ||
| 163 | ATAD3A | 8440 | 18432 | -6.199 | -0.1958 | Yes | ||
| 164 | PPRC1 | 23808 | 18439 | -6.263 | -0.1722 | Yes | ||
| 165 | NUDC | 9495 | 18448 | -6.381 | -0.1482 | Yes | ||
| 166 | PPAT | 6081 | 18522 | -7.211 | -0.1246 | Yes | ||
| 167 | LYAR | 16856 | 18565 | -8.120 | -0.0959 | Yes | ||
| 168 | ACY1 | 19012 2996 | 18576 | -8.328 | -0.0646 | Yes | ||
| 169 | PUS1 | 12103 | 18585 | -8.611 | -0.0321 | Yes | ||
| 170 | TIMM8A | 24062 | 18588 | -8.817 | 0.0015 | Yes |