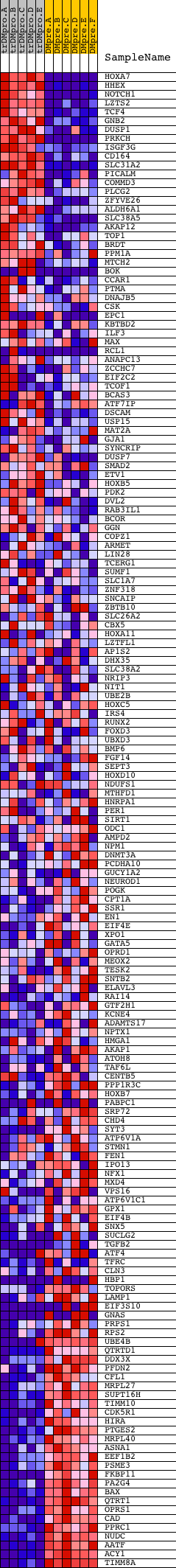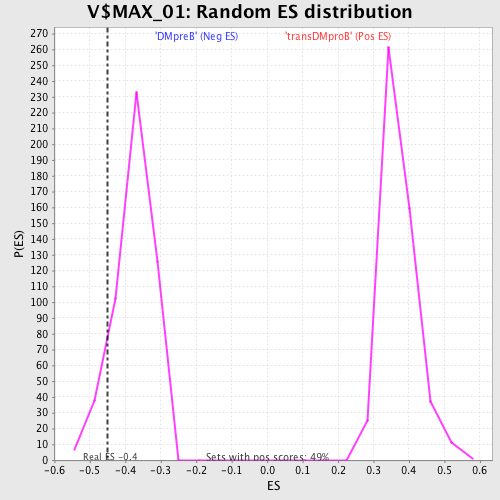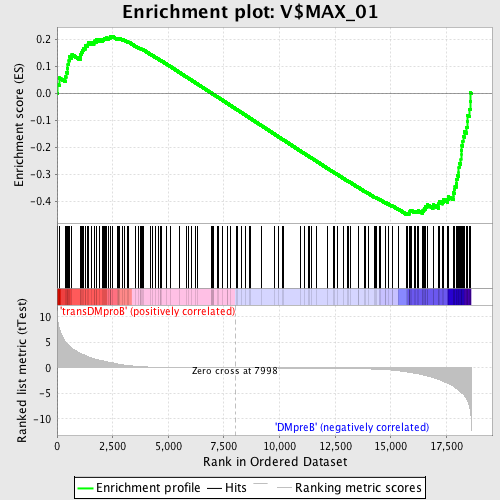
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | V$MAX_01 |
| Enrichment Score (ES) | -0.44957414 |
| Normalized Enrichment Score (NES) | -1.1977459 |
| Nominal p-value | 0.104743086 |
| FDR q-value | 0.8175155 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HOXA7 | 4862 | 35 | 9.076 | 0.0317 | No | ||
| 2 | HHEX | 23872 | 100 | 7.768 | 0.0569 | No | ||
| 3 | NOTCH1 | 14649 | 381 | 5.091 | 0.0606 | No | ||
| 4 | LZTS2 | 10362 | 401 | 4.974 | 0.0779 | No | ||
| 5 | TCF4 | 5642 | 463 | 4.620 | 0.0917 | No | ||
| 6 | GNB2 | 9026 | 487 | 4.520 | 0.1072 | No | ||
| 7 | DUSP1 | 23061 | 532 | 4.376 | 0.1210 | No | ||
| 8 | PRKCH | 21246 | 560 | 4.274 | 0.1353 | No | ||
| 9 | ISGF3G | 2686 9184 | 666 | 3.807 | 0.1437 | No | ||
| 10 | CD164 | 20046 | 1042 | 2.850 | 0.1339 | No | ||
| 11 | SLC31A2 | 9827 | 1063 | 2.803 | 0.1432 | No | ||
| 12 | PICALM | 18191 | 1089 | 2.731 | 0.1519 | No | ||
| 13 | COMMD3 | 73 | 1139 | 2.643 | 0.1590 | No | ||
| 14 | PLCG2 | 18453 | 1181 | 2.559 | 0.1663 | No | ||
| 15 | ZFYVE26 | 9999 | 1257 | 2.405 | 0.1711 | No | ||
| 16 | ALDH6A1 | 21025 | 1280 | 2.351 | 0.1786 | No | ||
| 17 | SLC38A5 | 24388 | 1387 | 2.170 | 0.1809 | No | ||
| 18 | AKAP12 | 17600 | 1392 | 2.167 | 0.1887 | No | ||
| 19 | TOP1 | 5790 5789 10210 | 1548 | 1.918 | 0.1874 | No | ||
| 20 | BRDT | 4327 | 1659 | 1.779 | 0.1880 | No | ||
| 21 | PPM1A | 5279 9607 | 1673 | 1.756 | 0.1938 | No | ||
| 22 | MTCH2 | 14954 7120 | 1761 | 1.653 | 0.1952 | No | ||
| 23 | BOK | 11953 14174 | 1776 | 1.639 | 0.2005 | No | ||
| 24 | CCAR1 | 7454 | 1917 | 1.496 | 0.1984 | No | ||
| 25 | PTMA | 9657 | 2030 | 1.375 | 0.1974 | No | ||
| 26 | DNAJB5 | 16236 | 2083 | 1.327 | 0.1995 | No | ||
| 27 | CSK | 8805 | 2115 | 1.299 | 0.2026 | No | ||
| 28 | EPC1 | 2039 8907 2033 | 2166 | 1.256 | 0.2046 | No | ||
| 29 | KBTBD2 | 17137 | 2234 | 1.184 | 0.2053 | No | ||
| 30 | ILF3 | 3110 3030 9176 | 2324 | 1.096 | 0.2045 | No | ||
| 31 | MAX | 21034 | 2381 | 1.044 | 0.2054 | No | ||
| 32 | RCL1 | 3728 3743 23894 | 2393 | 1.037 | 0.2086 | No | ||
| 33 | ANAPC13 | 12761 | 2405 | 1.025 | 0.2118 | No | ||
| 34 | ZCCHC7 | 16225 | 2506 | 0.962 | 0.2099 | No | ||
| 35 | EIF2C2 | 6247 | 2735 | 0.755 | 0.2004 | No | ||
| 36 | TCOF1 | 5660 | 2747 | 0.742 | 0.2025 | No | ||
| 37 | BCAS3 | 5314 | 2779 | 0.711 | 0.2035 | No | ||
| 38 | ATF7IP | 12028 13038 | 2811 | 0.682 | 0.2043 | No | ||
| 39 | DSCAM | 1667 4641 22530 | 2958 | 0.578 | 0.1985 | No | ||
| 40 | USP15 | 4758 3377 | 3023 | 0.544 | 0.1971 | No | ||
| 41 | MAT2A | 10553 10554 6099 | 3178 | 0.458 | 0.1904 | No | ||
| 42 | GJA1 | 20023 | 3216 | 0.444 | 0.1900 | No | ||
| 43 | SYNCRIP | 3078 3035 3107 | 3530 | 0.309 | 0.1742 | No | ||
| 44 | DUSP7 | 10661 | 3639 | 0.273 | 0.1694 | No | ||
| 45 | SMAD2 | 23511 | 3741 | 0.245 | 0.1648 | No | ||
| 46 | ETV1 | 4688 8925 8924 | 3755 | 0.241 | 0.1650 | No | ||
| 47 | HOXB5 | 20687 | 3804 | 0.229 | 0.1632 | No | ||
| 48 | PDK2 | 20285 | 3835 | 0.222 | 0.1624 | No | ||
| 49 | DVL2 | 20813 | 3883 | 0.212 | 0.1607 | No | ||
| 50 | RAB3IL1 | 3723 13229 | 4203 | 0.147 | 0.1439 | No | ||
| 51 | BCOR | 12934 2561 | 4278 | 0.138 | 0.1404 | No | ||
| 52 | GGN | 2472 1171 18312 | 4435 | 0.115 | 0.1324 | No | ||
| 53 | COPZ1 | 22338 | 4541 | 0.101 | 0.1271 | No | ||
| 54 | ARMET | 19005 | 4660 | 0.090 | 0.1210 | No | ||
| 55 | LIN28 | 15723 | 4666 | 0.090 | 0.1211 | No | ||
| 56 | TCERG1 | 12075 2027 7068 | 4677 | 0.088 | 0.1208 | No | ||
| 57 | SUMF1 | 17049 | 4898 | 0.074 | 0.1092 | No | ||
| 58 | SLC1A7 | 16147 | 5106 | 0.063 | 0.0982 | No | ||
| 59 | ZNF318 | 12198 7179 | 5495 | 0.046 | 0.0773 | No | ||
| 60 | SNCAIP | 12589 | 5815 | 0.037 | 0.0602 | No | ||
| 61 | ZBTB10 | 10468 | 5916 | 0.035 | 0.0549 | No | ||
| 62 | SLC26A2 | 23427 | 6029 | 0.032 | 0.0489 | No | ||
| 63 | CBX5 | 22108 4485 | 6228 | 0.028 | 0.0383 | No | ||
| 64 | HOXA11 | 17146 | 6300 | 0.026 | 0.0346 | No | ||
| 65 | LZTFL1 | 8210 | 6920 | 0.015 | 0.0011 | No | ||
| 66 | AP1S2 | 8423 4228 | 6926 | 0.015 | 0.0009 | No | ||
| 67 | DHX35 | 14754 | 6979 | 0.014 | -0.0019 | No | ||
| 68 | SLC38A2 | 22149 2219 | 6982 | 0.014 | -0.0020 | No | ||
| 69 | NRIP3 | 17679 | 7020 | 0.014 | -0.0039 | No | ||
| 70 | NIT1 | 11293 933 | 7039 | 0.013 | -0.0049 | No | ||
| 71 | UBE2B | 10245 | 7199 | 0.011 | -0.0134 | No | ||
| 72 | HOXC5 | 22340 | 7250 | 0.010 | -0.0161 | No | ||
| 73 | IRS4 | 9183 4926 | 7411 | 0.008 | -0.0247 | No | ||
| 74 | RUNX2 | 4480 8700 | 7650 | 0.004 | -0.0376 | No | ||
| 75 | FOXD3 | 9088 | 7667 | 0.004 | -0.0385 | No | ||
| 76 | UBXD3 | 15701 | 7805 | 0.003 | -0.0459 | No | ||
| 77 | BMP6 | 8659 | 8051 | -0.001 | -0.0592 | No | ||
| 78 | FGF14 | 21716 3005 3463 | 8086 | -0.001 | -0.0610 | No | ||
| 79 | SEPT3 | 6275 10775 | 8303 | -0.004 | -0.0727 | No | ||
| 80 | HOXD10 | 14983 | 8488 | -0.007 | -0.0827 | No | ||
| 81 | NDUFS1 | 13940 | 8627 | -0.008 | -0.0901 | No | ||
| 82 | MTHFD1 | 2132 21238 | 8706 | -0.010 | -0.0943 | No | ||
| 83 | HNRPA1 | 9103 4860 | 9192 | -0.016 | -0.1205 | No | ||
| 84 | PER1 | 20827 | 9750 | -0.024 | -0.1506 | No | ||
| 85 | SIRT1 | 19745 | 9971 | -0.028 | -0.1625 | No | ||
| 86 | ODC1 | 9500 | 10123 | -0.031 | -0.1705 | No | ||
| 87 | AMPD2 | 1931 15197 | 10189 | -0.032 | -0.1739 | No | ||
| 88 | NPM1 | 1196 | 10941 | -0.045 | -0.2145 | No | ||
| 89 | DNMT3A | 2167 21330 | 11106 | -0.048 | -0.2232 | No | ||
| 90 | PCDHA10 | 8792 | 11293 | -0.053 | -0.2331 | No | ||
| 91 | GUCY1A2 | 19580 | 11338 | -0.054 | -0.2353 | No | ||
| 92 | NEUROD1 | 14550 | 11440 | -0.056 | -0.2405 | No | ||
| 93 | POGK | 7739 | 11666 | -0.062 | -0.2525 | No | ||
| 94 | CPT1A | 23756 | 12142 | -0.077 | -0.2780 | No | ||
| 95 | SSR1 | 4211 | 12408 | -0.089 | -0.2920 | No | ||
| 96 | EN1 | 387 14155 | 12477 | -0.092 | -0.2953 | No | ||
| 97 | EIF4E | 15403 1827 8890 | 12621 | -0.098 | -0.3027 | No | ||
| 98 | XPO1 | 4172 | 12880 | -0.112 | -0.3163 | No | ||
| 99 | GATA5 | 14315 | 13062 | -0.123 | -0.3257 | No | ||
| 100 | OPRD1 | 15737 | 13108 | -0.125 | -0.3276 | No | ||
| 101 | MEOX2 | 21285 | 13171 | -0.130 | -0.3305 | No | ||
| 102 | TESK2 | 10504 | 13544 | -0.160 | -0.3501 | No | ||
| 103 | SNTB2 | 3887 18476 | 13549 | -0.161 | -0.3497 | No | ||
| 104 | ELAVL3 | 9136 | 13814 | -0.186 | -0.3633 | No | ||
| 105 | RAI14 | 22327 2191 | 13866 | -0.191 | -0.3654 | No | ||
| 106 | GTF2H1 | 4069 18236 | 13978 | -0.204 | -0.3707 | No | ||
| 107 | KCNE4 | 12196 | 14270 | -0.243 | -0.3855 | No | ||
| 108 | ADAMTS17 | 598 | 14288 | -0.245 | -0.3855 | No | ||
| 109 | NPTX1 | 20126 | 14362 | -0.257 | -0.3886 | No | ||
| 110 | HMGA1 | 23323 | 14469 | -0.276 | -0.3933 | No | ||
| 111 | AKAP1 | 4364 1326 1286 20299 | 14520 | -0.287 | -0.3949 | No | ||
| 112 | ATOH8 | 17120 | 14778 | -0.351 | -0.4076 | No | ||
| 113 | TAF6L | 5923 | 14780 | -0.351 | -0.4063 | No | ||
| 114 | CENTB5 | 15961 | 14876 | -0.378 | -0.4101 | No | ||
| 115 | PPP1R3C | 23688 | 15077 | -0.444 | -0.4193 | No | ||
| 116 | HOXB7 | 666 9110 | 15082 | -0.446 | -0.4178 | No | ||
| 117 | PABPC1 | 5219 9522 9523 23572 | 15334 | -0.551 | -0.4294 | No | ||
| 118 | SRP72 | 16819 | 15705 | -0.787 | -0.4466 | Yes | ||
| 119 | CHD4 | 8418 17281 4225 | 15754 | -0.826 | -0.4461 | Yes | ||
| 120 | SYT3 | 18264 | 15819 | -0.874 | -0.4463 | Yes | ||
| 121 | ATP6V1A | 8638 | 15844 | -0.888 | -0.4444 | Yes | ||
| 122 | STMN1 | 9261 | 15852 | -0.895 | -0.4414 | Yes | ||
| 123 | FEN1 | 8961 | 15855 | -0.898 | -0.4382 | Yes | ||
| 124 | IPO13 | 15784 | 15881 | -0.920 | -0.4362 | Yes | ||
| 125 | NFX1 | 2475 | 15926 | -0.961 | -0.4350 | Yes | ||
| 126 | MXD4 | 9346 | 16082 | -1.040 | -0.4396 | Yes | ||
| 127 | VPS16 | 2927 14845 2825 | 16105 | -1.061 | -0.4368 | Yes | ||
| 128 | ATP6V1C1 | 22487 12329 | 16220 | -1.155 | -0.4387 | Yes | ||
| 129 | GPX1 | 19310 | 16224 | -1.160 | -0.4346 | Yes | ||
| 130 | EIF4B | 13279 7979 | 16431 | -1.360 | -0.4407 | Yes | ||
| 131 | SNX5 | 12783 14410 | 16441 | -1.378 | -0.4361 | Yes | ||
| 132 | SUCLG2 | 17062 5545 | 16480 | -1.436 | -0.4329 | Yes | ||
| 133 | TGFB2 | 5743 | 16495 | -1.457 | -0.4283 | Yes | ||
| 134 | ATF4 | 22417 2239 | 16551 | -1.513 | -0.4256 | Yes | ||
| 135 | TFRC | 22785 | 16555 | -1.516 | -0.4202 | Yes | ||
| 136 | CLN3 | 17635 | 16628 | -1.574 | -0.4183 | Yes | ||
| 137 | HBP1 | 2048 2162 21086 | 16634 | -1.578 | -0.4127 | Yes | ||
| 138 | TOPORS | 15920 | 16898 | -1.882 | -0.4200 | Yes | ||
| 139 | LAMP1 | 18679 | 16912 | -1.905 | -0.4137 | Yes | ||
| 140 | EIF3S10 | 4659 8887 | 17154 | -2.229 | -0.4185 | Yes | ||
| 141 | GNAS | 9025 2963 2752 | 17156 | -2.230 | -0.4103 | Yes | ||
| 142 | PRPS1 | 24233 | 17165 | -2.238 | -0.4025 | Yes | ||
| 143 | RPS2 | 9279 | 17318 | -2.573 | -0.4012 | Yes | ||
| 144 | UBE4B | 2433 15672 | 17367 | -2.645 | -0.3940 | Yes | ||
| 145 | QTRTD1 | 22590 | 17555 | -2.998 | -0.3931 | Yes | ||
| 146 | DDX3X | 24379 | 17590 | -3.059 | -0.3836 | Yes | ||
| 147 | PFDN2 | 9551 | 17813 | -3.599 | -0.3823 | Yes | ||
| 148 | CFL1 | 4516 | 17815 | -3.604 | -0.3691 | Yes | ||
| 149 | MRPL27 | 13573 | 17873 | -3.766 | -0.3582 | Yes | ||
| 150 | SUPT16H | 8541 | 17882 | -3.805 | -0.3446 | Yes | ||
| 151 | TIMM10 | 11390 2158 | 17944 | -4.026 | -0.3330 | Yes | ||
| 152 | CDK5R1 | 20745 | 17957 | -4.064 | -0.3186 | Yes | ||
| 153 | HIRA | 4852 9090 | 18002 | -4.201 | -0.3055 | Yes | ||
| 154 | PTGES2 | 15038 | 18059 | -4.433 | -0.2921 | Yes | ||
| 155 | MRPL40 | 22641 | 18062 | -4.439 | -0.2758 | Yes | ||
| 156 | ASNA1 | 18810 | 18070 | -4.491 | -0.2596 | Yes | ||
| 157 | EEF1B2 | 4131 12063 | 18152 | -4.785 | -0.2463 | Yes | ||
| 158 | PSME3 | 20657 | 18155 | -4.800 | -0.2287 | Yes | ||
| 159 | FKBP11 | 22136 | 18158 | -4.815 | -0.2110 | Yes | ||
| 160 | PA2G4 | 5265 | 18195 | -4.947 | -0.1946 | Yes | ||
| 161 | BAX | 17832 | 18237 | -5.057 | -0.1781 | Yes | ||
| 162 | QTRT1 | 19540 | 18247 | -5.085 | -0.1598 | Yes | ||
| 163 | OPRS1 | 9515 2402 | 18303 | -5.345 | -0.1430 | Yes | ||
| 164 | CAD | 16886 | 18408 | -6.035 | -0.1264 | Yes | ||
| 165 | PPRC1 | 23808 | 18439 | -6.263 | -0.1048 | Yes | ||
| 166 | NUDC | 9495 | 18448 | -6.381 | -0.0817 | Yes | ||
| 167 | AATF | 20315 | 18534 | -7.364 | -0.0591 | Yes | ||
| 168 | ACY1 | 19012 2996 | 18576 | -8.328 | -0.0305 | Yes | ||
| 169 | TIMM8A | 24062 | 18588 | -8.817 | 0.0015 | Yes |