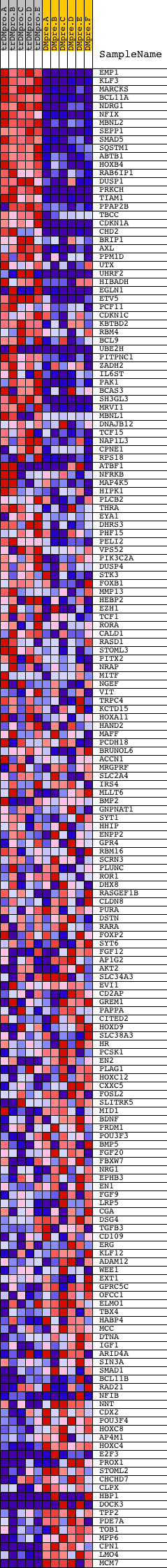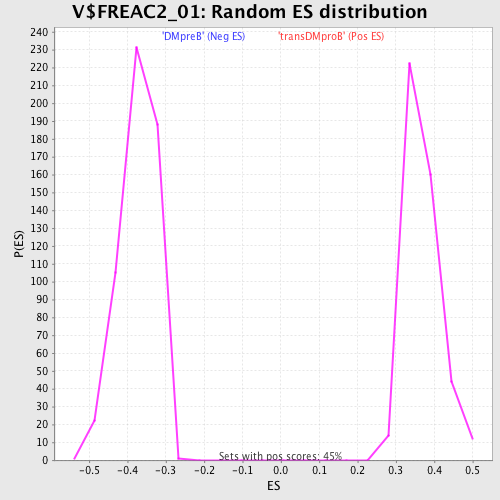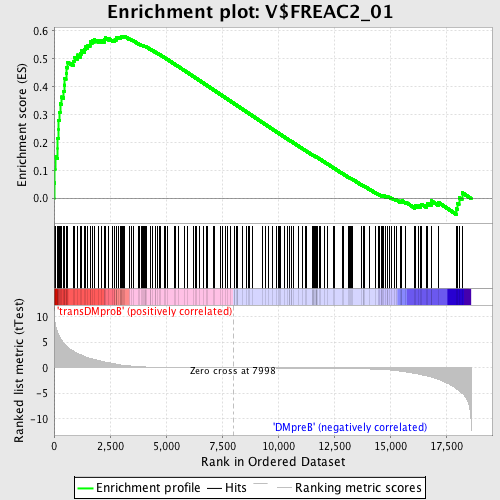
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$FREAC2_01 |
| Enrichment Score (ES) | 0.5810969 |
| Normalized Enrichment Score (NES) | 1.5863699 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04996974 |
| FWER p-Value | 0.372 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EMP1 | 17260 | 2 | 10.887 | 0.0565 | Yes | ||
| 2 | KLF3 | 4961 | 29 | 9.444 | 0.1042 | Yes | ||
| 3 | MARCKS | 9331 | 41 | 8.902 | 0.1499 | Yes | ||
| 4 | BCL11A | 4691 | 159 | 6.951 | 0.1797 | Yes | ||
| 5 | NDRG1 | 22276 | 161 | 6.937 | 0.2157 | Yes | ||
| 6 | NFIX | 9458 | 199 | 6.474 | 0.2473 | Yes | ||
| 7 | MBNL2 | 21932 | 205 | 6.438 | 0.2805 | Yes | ||
| 8 | SEPP1 | 22524 | 249 | 6.088 | 0.3098 | Yes | ||
| 9 | SMAD5 | 21621 | 269 | 5.871 | 0.3393 | Yes | ||
| 10 | SQSTM1 | 9517 | 338 | 5.381 | 0.3636 | Yes | ||
| 11 | ABTB1 | 17075 | 431 | 4.793 | 0.3835 | Yes | ||
| 12 | HOXB4 | 20686 | 456 | 4.654 | 0.4064 | Yes | ||
| 13 | RAB6IP1 | 17676 | 473 | 4.566 | 0.4293 | Yes | ||
| 14 | DUSP1 | 23061 | 532 | 4.376 | 0.4489 | Yes | ||
| 15 | PRKCH | 21246 | 560 | 4.274 | 0.4697 | Yes | ||
| 16 | TIAM1 | 22547 | 609 | 4.034 | 0.4880 | Yes | ||
| 17 | PPAP2B | 7503 16161 | 856 | 3.293 | 0.4918 | Yes | ||
| 18 | TBCC | 23215 | 897 | 3.189 | 0.5062 | Yes | ||
| 19 | CDKN1A | 4511 8729 | 1034 | 2.865 | 0.5137 | Yes | ||
| 20 | CHD2 | 10847 | 1177 | 2.569 | 0.5194 | Yes | ||
| 21 | BRIP1 | 20311 | 1224 | 2.480 | 0.5298 | Yes | ||
| 22 | AXL | 17922 | 1358 | 2.215 | 0.5341 | Yes | ||
| 23 | PPM1D | 20721 | 1407 | 2.132 | 0.5426 | Yes | ||
| 24 | UTX | 10266 2574 | 1503 | 1.992 | 0.5478 | Yes | ||
| 25 | UHRF2 | 3731 4262 8452 23887 | 1605 | 1.854 | 0.5519 | Yes | ||
| 26 | HIBADH | 17144 | 1610 | 1.847 | 0.5613 | Yes | ||
| 27 | EGLN1 | 8504 18707 4298 | 1705 | 1.718 | 0.5652 | Yes | ||
| 28 | ETV5 | 22630 | 1802 | 1.629 | 0.5684 | Yes | ||
| 29 | PCF11 | 121 | 1970 | 1.442 | 0.5669 | Yes | ||
| 30 | CDKN1C | 17546 | 2111 | 1.301 | 0.5660 | Yes | ||
| 31 | KBTBD2 | 17137 | 2234 | 1.184 | 0.5656 | Yes | ||
| 32 | RBM4 | 9706 3693 | 2253 | 1.162 | 0.5706 | Yes | ||
| 33 | BCL9 | 15232 | 2272 | 1.139 | 0.5756 | Yes | ||
| 34 | UBE2H | 5823 5822 | 2433 | 1.005 | 0.5721 | Yes | ||
| 35 | PITPNC1 | 7759 | 2612 | 0.862 | 0.5670 | Yes | ||
| 36 | ZADH2 | 23501 | 2700 | 0.782 | 0.5663 | Yes | ||
| 37 | IL6ST | 4920 | 2717 | 0.771 | 0.5694 | Yes | ||
| 38 | PAK1 | 9527 | 2767 | 0.720 | 0.5705 | Yes | ||
| 39 | BCAS3 | 5314 | 2779 | 0.711 | 0.5736 | Yes | ||
| 40 | SH3GL3 | 18201 | 2790 | 0.702 | 0.5767 | Yes | ||
| 41 | MRVI1 | 17674 1020 2526 | 2854 | 0.647 | 0.5767 | Yes | ||
| 42 | MBNL1 | 1921 15582 | 2963 | 0.575 | 0.5738 | Yes | ||
| 43 | DNAJB12 | 7146 | 2971 | 0.571 | 0.5764 | Yes | ||
| 44 | TCF15 | 14798 | 2988 | 0.559 | 0.5785 | Yes | ||
| 45 | NAP1L3 | 24266 | 3011 | 0.549 | 0.5801 | Yes | ||
| 46 | CPNE1 | 11186 | 3061 | 0.519 | 0.5802 | Yes | ||
| 47 | RPS18 | 1529 5397 1603 | 3093 | 0.504 | 0.5811 | Yes | ||
| 48 | ATBF1 | 916 8633 | 3161 | 0.464 | 0.5799 | No | ||
| 49 | NFRKB | 10642 | 3361 | 0.376 | 0.5710 | No | ||
| 50 | MAP4K5 | 11853 21048 2101 2142 | 3458 | 0.336 | 0.5676 | No | ||
| 51 | HIPK1 | 4851 | 3522 | 0.312 | 0.5658 | No | ||
| 52 | PLCB2 | 5262 | 3787 | 0.233 | 0.5527 | No | ||
| 53 | THRA | 1447 10171 1406 | 3826 | 0.224 | 0.5518 | No | ||
| 54 | EYA1 | 4695 4061 | 3919 | 0.200 | 0.5478 | No | ||
| 55 | DHRS3 | 15997 | 3946 | 0.193 | 0.5474 | No | ||
| 56 | PHF15 | 20467 8050 | 3986 | 0.184 | 0.5463 | No | ||
| 57 | PELI2 | 8217 | 4050 | 0.172 | 0.5437 | No | ||
| 58 | VPS52 | 23287 1622 | 4069 | 0.169 | 0.5436 | No | ||
| 59 | PIK3C2A | 17667 | 4070 | 0.168 | 0.5445 | No | ||
| 60 | DUSP4 | 18632 3820 | 4129 | 0.160 | 0.5422 | No | ||
| 61 | STK3 | 7084 | 4281 | 0.137 | 0.5347 | No | ||
| 62 | FOXB1 | 12254 19069 | 4317 | 0.132 | 0.5335 | No | ||
| 63 | MMP13 | 19573 3007 | 4386 | 0.120 | 0.5305 | No | ||
| 64 | HEBP2 | 19812 | 4506 | 0.106 | 0.5246 | No | ||
| 65 | EZH1 | 20217 | 4546 | 0.101 | 0.5230 | No | ||
| 66 | TCF1 | 16416 | 4599 | 0.096 | 0.5206 | No | ||
| 67 | RORA | 3019 5388 | 4686 | 0.088 | 0.5164 | No | ||
| 68 | CALD1 | 4273 8463 | 4749 | 0.083 | 0.5135 | No | ||
| 69 | RASD1 | 20424 | 4940 | 0.071 | 0.5036 | No | ||
| 70 | STOML3 | 15594 | 4967 | 0.070 | 0.5025 | No | ||
| 71 | PITX2 | 15424 1878 | 5066 | 0.065 | 0.4975 | No | ||
| 72 | NRAP | 23642 | 5385 | 0.050 | 0.4806 | No | ||
| 73 | MITF | 17349 | 5440 | 0.048 | 0.4779 | No | ||
| 74 | NGEF | 13891 | 5533 | 0.045 | 0.4731 | No | ||
| 75 | VIT | 23155 | 5816 | 0.037 | 0.4580 | No | ||
| 76 | TRPC4 | 15593 | 5944 | 0.034 | 0.4513 | No | ||
| 77 | KCTD15 | 17864 | 6209 | 0.028 | 0.4371 | No | ||
| 78 | HOXA11 | 17146 | 6300 | 0.026 | 0.4324 | No | ||
| 79 | HAND2 | 18615 | 6345 | 0.025 | 0.4301 | No | ||
| 80 | MAFF | 22423 2243 | 6483 | 0.023 | 0.4228 | No | ||
| 81 | PCDH18 | 15348 | 6680 | 0.019 | 0.4123 | No | ||
| 82 | BRUNOL6 | 19421 | 6795 | 0.017 | 0.4062 | No | ||
| 83 | ACCN1 | 20327 1194 | 6825 | 0.017 | 0.4047 | No | ||
| 84 | MRGPRF | 17995 | 7111 | 0.012 | 0.3893 | No | ||
| 85 | SLC2A4 | 20380 | 7171 | 0.011 | 0.3862 | No | ||
| 86 | IRS4 | 9183 4926 | 7411 | 0.008 | 0.3733 | No | ||
| 87 | MLLT6 | 20678 | 7530 | 0.006 | 0.3669 | No | ||
| 88 | BMP2 | 14833 | 7661 | 0.004 | 0.3599 | No | ||
| 89 | GNPNAT1 | 12027 7031 | 7745 | 0.003 | 0.3554 | No | ||
| 90 | SYT1 | 5565 | 7856 | 0.002 | 0.3494 | No | ||
| 91 | HHIP | 4849 | 8030 | -0.000 | 0.3400 | No | ||
| 92 | ENPP2 | 9548 | 8049 | -0.001 | 0.3391 | No | ||
| 93 | GPR4 | 18364 | 8145 | -0.002 | 0.3339 | No | ||
| 94 | RBM16 | 11492 8387 | 8175 | -0.003 | 0.3324 | No | ||
| 95 | SCRN3 | 14984 | 8198 | -0.003 | 0.3312 | No | ||
| 96 | PLUNC | 9593 | 8412 | -0.006 | 0.3197 | No | ||
| 97 | ROR1 | 6472 | 8593 | -0.008 | 0.3099 | No | ||
| 98 | DHX8 | 20649 | 8662 | -0.009 | 0.3063 | No | ||
| 99 | RASGEF1B | 11515 16469 | 8707 | -0.010 | 0.3040 | No | ||
| 100 | CLDN8 | 22549 1747 | 8838 | -0.011 | 0.2970 | No | ||
| 101 | PURA | 9670 | 9307 | -0.018 | 0.2717 | No | ||
| 102 | DSTN | 14823 | 9445 | -0.020 | 0.2644 | No | ||
| 103 | RARA | 5358 | 9584 | -0.022 | 0.2570 | No | ||
| 104 | FOXP2 | 4312 17523 8523 | 9742 | -0.024 | 0.2486 | No | ||
| 105 | SYT6 | 13437 7042 12040 | 9929 | -0.027 | 0.2387 | No | ||
| 106 | FGF12 | 1723 22621 | 9994 | -0.028 | 0.2353 | No | ||
| 107 | AP1G2 | 2677 21827 | 10043 | -0.029 | 0.2329 | No | ||
| 108 | AKT2 | 4365 4366 | 10089 | -0.030 | 0.2306 | No | ||
| 109 | SLC34A3 | 14664 | 10267 | -0.033 | 0.2212 | No | ||
| 110 | EVI1 | 4689 8926 | 10413 | -0.036 | 0.2135 | No | ||
| 111 | CD2AP | 22975 | 10510 | -0.037 | 0.2085 | No | ||
| 112 | GREM1 | 14485 | 10521 | -0.037 | 0.2081 | No | ||
| 113 | PAPPA | 16190 | 10574 | -0.038 | 0.2055 | No | ||
| 114 | CITED2 | 5118 14477 | 10616 | -0.039 | 0.2035 | No | ||
| 115 | HOXD9 | 14982 | 10707 | -0.040 | 0.1988 | No | ||
| 116 | SLC38A3 | 13319 | 10929 | -0.045 | 0.1871 | No | ||
| 117 | HR | 3622 9119 | 11074 | -0.048 | 0.1795 | No | ||
| 118 | PCSK1 | 21600 | 11208 | -0.051 | 0.1726 | No | ||
| 119 | EN2 | 16898 | 11277 | -0.052 | 0.1691 | No | ||
| 120 | PLAG1 | 7148 15949 12161 | 11529 | -0.058 | 0.1558 | No | ||
| 121 | HOXC12 | 22343 | 11534 | -0.058 | 0.1559 | No | ||
| 122 | CXXC5 | 12523 | 11557 | -0.059 | 0.1550 | No | ||
| 123 | FOSL2 | 4733 8978 16878 | 11587 | -0.060 | 0.1538 | No | ||
| 124 | SLITRK5 | 21939 | 11609 | -0.060 | 0.1529 | No | ||
| 125 | MID1 | 5097 5098 | 11671 | -0.062 | 0.1499 | No | ||
| 126 | BDNF | 14926 2797 | 11691 | -0.062 | 0.1492 | No | ||
| 127 | PRDM1 | 19775 3337 | 11694 | -0.062 | 0.1495 | No | ||
| 128 | POU3F3 | 14261 | 11726 | -0.063 | 0.1481 | No | ||
| 129 | BMP5 | 19376 | 11749 | -0.064 | 0.1472 | No | ||
| 130 | FGF20 | 18892 | 11835 | -0.067 | 0.1430 | No | ||
| 131 | FBXW7 | 1805 11928 | 11871 | -0.068 | 0.1414 | No | ||
| 132 | NRG1 | 3885 | 12073 | -0.075 | 0.1309 | No | ||
| 133 | EPHB3 | 22816 | 12211 | -0.080 | 0.1239 | No | ||
| 134 | EN1 | 387 14155 | 12477 | -0.092 | 0.1100 | No | ||
| 135 | FGF9 | 8966 | 12531 | -0.094 | 0.1076 | No | ||
| 136 | LRP5 | 23948 9285 | 12876 | -0.111 | 0.0896 | No | ||
| 137 | CGA | 16246 | 12911 | -0.114 | 0.0883 | No | ||
| 138 | DSG4 | 9262 23616 | 13159 | -0.129 | 0.0756 | No | ||
| 139 | TGFB3 | 10161 | 13186 | -0.131 | 0.0748 | No | ||
| 140 | CD109 | 6174 10656 19369 | 13243 | -0.135 | 0.0725 | No | ||
| 141 | ERG | 1686 8915 | 13277 | -0.137 | 0.0714 | No | ||
| 142 | KLF12 | 4960 2637 21732 9228 | 13321 | -0.141 | 0.0698 | No | ||
| 143 | ADAM12 | 2207 8545 | 13702 | -0.174 | 0.0501 | No | ||
| 144 | WEE1 | 18127 | 13796 | -0.184 | 0.0460 | No | ||
| 145 | EXT1 | 22297 | 13817 | -0.187 | 0.0459 | No | ||
| 146 | GPRC5C | 12864 | 13855 | -0.191 | 0.0449 | No | ||
| 147 | OFCC1 | 21486 | 14073 | -0.215 | 0.0343 | No | ||
| 148 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 14344 | -0.253 | 0.0209 | No | ||
| 149 | TBX4 | 5633 | 14471 | -0.277 | 0.0155 | No | ||
| 150 | HABP4 | 12145 | 14531 | -0.289 | 0.0138 | No | ||
| 151 | MCC | 552 | 14634 | -0.315 | 0.0100 | No | ||
| 152 | DTNA | 8870 4647 23611 1991 | 14636 | -0.315 | 0.0115 | No | ||
| 153 | IGF1 | 3352 9156 3409 | 14678 | -0.326 | 0.0110 | No | ||
| 154 | ARID4A | 2080 6215 | 14683 | -0.328 | 0.0125 | No | ||
| 155 | SIN3A | 5442 | 14813 | -0.359 | 0.0074 | No | ||
| 156 | SMAD1 | 18831 922 | 14865 | -0.374 | 0.0065 | No | ||
| 157 | BCL11B | 7190 | 14886 | -0.380 | 0.0074 | No | ||
| 158 | RAD21 | 22298 | 14963 | -0.404 | 0.0054 | No | ||
| 159 | NFIB | 15855 | 15064 | -0.439 | 0.0023 | No | ||
| 160 | NNT | 3273 5181 9471 3238 | 15173 | -0.484 | -0.0011 | No | ||
| 161 | CDX2 | 16289 | 15282 | -0.527 | -0.0042 | No | ||
| 162 | POU3F4 | 347 | 15458 | -0.633 | -0.0104 | No | ||
| 163 | HOXC8 | 4865 | 15522 | -0.664 | -0.0104 | No | ||
| 164 | AP4M1 | 3499 16656 | 15528 | -0.668 | -0.0072 | No | ||
| 165 | HOXC4 | 4864 | 15695 | -0.780 | -0.0121 | No | ||
| 166 | E2F3 | 21500 8874 | 16104 | -1.060 | -0.0287 | No | ||
| 167 | PROX1 | 9623 | 16126 | -1.071 | -0.0243 | No | ||
| 168 | STOML2 | 15902 | 16273 | -1.202 | -0.0260 | No | ||
| 169 | CHCHD7 | 16280 | 16366 | -1.289 | -0.0242 | No | ||
| 170 | CLPX | 19403 | 16400 | -1.335 | -0.0191 | No | ||
| 171 | HBP1 | 2048 2162 21086 | 16634 | -1.578 | -0.0235 | No | ||
| 172 | DOCK3 | 9920 5537 10751 | 16654 | -1.587 | -0.0163 | No | ||
| 173 | TPP2 | 14257 | 16832 | -1.791 | -0.0166 | No | ||
| 174 | PDE7A | 9544 1788 | 16836 | -1.794 | -0.0074 | No | ||
| 175 | TOB1 | 20703 | 17150 | -2.224 | -0.0128 | No | ||
| 176 | MPP6 | 17448 | 17954 | -4.052 | -0.0353 | No | ||
| 177 | CPN1 | 23667 | 17997 | -4.189 | -0.0158 | No | ||
| 178 | LMO4 | 15151 | 18076 | -4.510 | 0.0034 | No | ||
| 179 | MCM7 | 9372 3568 | 18211 | -4.973 | 0.0220 | No |