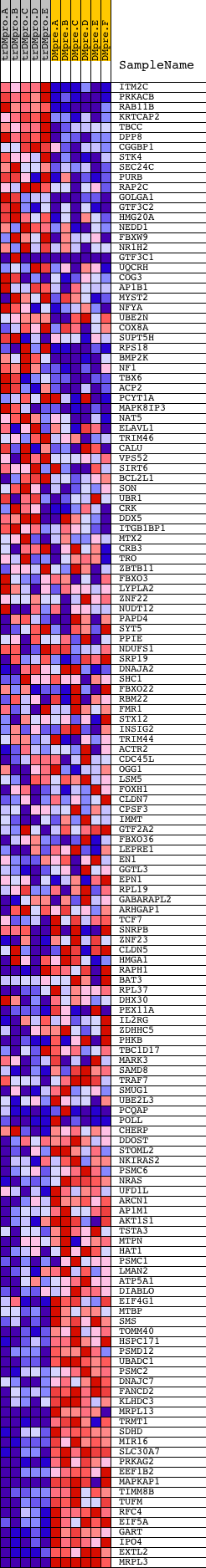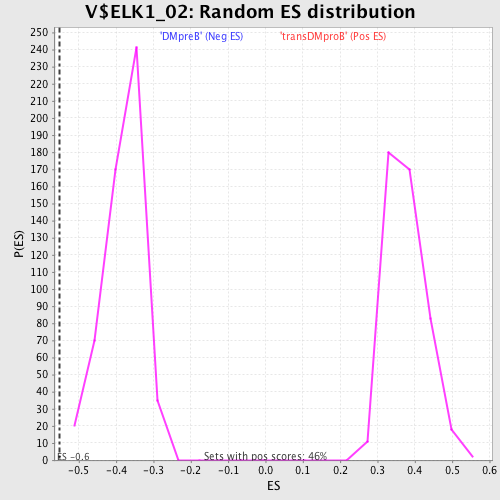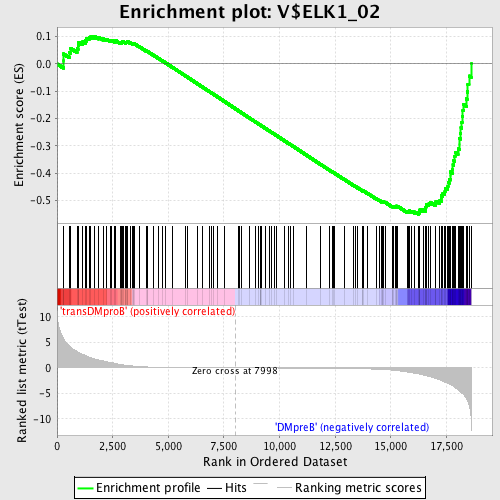
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | V$ELK1_02 |
| Enrichment Score (ES) | -0.55190957 |
| Normalized Enrichment Score (NES) | -1.4633961 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.28903794 |
| FWER p-Value | 0.94 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITM2C | 14204 | 282 | 5.751 | 0.0109 | No | ||
| 2 | PRKACB | 15140 | 286 | 5.707 | 0.0368 | No | ||
| 3 | RAB11B | 9676 | 558 | 4.276 | 0.0416 | No | ||
| 4 | KRTCAP2 | 15541 | 614 | 4.026 | 0.0569 | No | ||
| 5 | TBCC | 23215 | 897 | 3.189 | 0.0562 | No | ||
| 6 | DPP8 | 19408 | 943 | 3.063 | 0.0677 | No | ||
| 7 | CGGBP1 | 22724 | 963 | 3.008 | 0.0804 | No | ||
| 8 | STK4 | 14743 | 1158 | 2.598 | 0.0817 | No | ||
| 9 | SEC24C | 22086 | 1269 | 2.375 | 0.0866 | No | ||
| 10 | PURB | 5335 | 1325 | 2.278 | 0.0940 | No | ||
| 11 | RAP2C | 24156 | 1439 | 2.082 | 0.0973 | No | ||
| 12 | GOLGA1 | 14595 | 1519 | 1.968 | 0.1020 | No | ||
| 13 | GTF3C2 | 7749 | 1695 | 1.732 | 0.1004 | No | ||
| 14 | HMG20A | 19436 | 1864 | 1.552 | 0.0984 | No | ||
| 15 | NEDD1 | 19643 | 2081 | 1.330 | 0.0928 | No | ||
| 16 | FBXW9 | 18540 | 2239 | 1.172 | 0.0896 | No | ||
| 17 | NR1H2 | 17846 | 2387 | 1.040 | 0.0864 | No | ||
| 18 | GTF3C1 | 10607 17646 | 2465 | 0.997 | 0.0868 | No | ||
| 19 | UQCRH | 12378 | 2594 | 0.878 | 0.0838 | No | ||
| 20 | COG3 | 21751 | 2639 | 0.835 | 0.0853 | No | ||
| 21 | AP1B1 | 8599 | 2838 | 0.661 | 0.0775 | No | ||
| 22 | MYST2 | 20283 | 2892 | 0.620 | 0.0775 | No | ||
| 23 | NFYA | 5172 | 2902 | 0.613 | 0.0798 | No | ||
| 24 | UBE2N | 8216 19898 | 2919 | 0.603 | 0.0817 | No | ||
| 25 | COX8A | 8777 | 2967 | 0.573 | 0.0818 | No | ||
| 26 | SUPT5H | 9939 | 3083 | 0.511 | 0.0779 | No | ||
| 27 | RPS18 | 1529 5397 1603 | 3093 | 0.504 | 0.0797 | No | ||
| 28 | BMP2K | 3657 16786 | 3139 | 0.478 | 0.0794 | No | ||
| 29 | NF1 | 5165 | 3151 | 0.469 | 0.0810 | No | ||
| 30 | TBX6 | 1993 18075 | 3155 | 0.467 | 0.0829 | No | ||
| 31 | ACP2 | 4318 | 3295 | 0.405 | 0.0772 | No | ||
| 32 | PCYT1A | 4572 | 3387 | 0.364 | 0.0740 | No | ||
| 33 | MAPK8IP3 | 11402 23080 | 3412 | 0.354 | 0.0743 | No | ||
| 34 | NAT5 | 18601 7496 12597 | 3418 | 0.353 | 0.0756 | No | ||
| 35 | ELAVL1 | 4888 | 3484 | 0.327 | 0.0736 | No | ||
| 36 | TRIM46 | 1779 15281 | 3685 | 0.260 | 0.0640 | No | ||
| 37 | CALU | 1124 1149 8683 1162 4472 | 3997 | 0.181 | 0.0479 | No | ||
| 38 | VPS52 | 23287 1622 | 4069 | 0.169 | 0.0449 | No | ||
| 39 | SIRT6 | 19668 3447 | 4080 | 0.166 | 0.0451 | No | ||
| 40 | BCL2L1 | 4440 2930 8652 | 4316 | 0.132 | 0.0330 | No | ||
| 41 | SON | 5473 1657 1684 | 4552 | 0.100 | 0.0207 | No | ||
| 42 | UBR1 | 5828 | 4732 | 0.084 | 0.0114 | No | ||
| 43 | CRK | 4559 1249 | 4878 | 0.075 | 0.0039 | No | ||
| 44 | DDX5 | 4606 8843 4607 | 5187 | 0.059 | -0.0125 | No | ||
| 45 | ITGB1BP1 | 9189 | 5789 | 0.038 | -0.0449 | No | ||
| 46 | MTX2 | 14980 | 5878 | 0.035 | -0.0495 | No | ||
| 47 | CRB3 | 23176 | 6325 | 0.026 | -0.0735 | No | ||
| 48 | TRO | 7071 | 6527 | 0.022 | -0.0843 | No | ||
| 49 | ZBTB11 | 11309 | 6853 | 0.016 | -0.1018 | No | ||
| 50 | FBXO3 | 2662 14934 | 6937 | 0.015 | -0.1063 | No | ||
| 51 | LYPLA2 | 15708 | 7038 | 0.013 | -0.1116 | No | ||
| 52 | ZNF22 | 12497 7417 | 7214 | 0.011 | -0.1211 | No | ||
| 53 | NUDT12 | 7518 | 7517 | 0.006 | -0.1374 | No | ||
| 54 | PAPD4 | 8299 8300 4146 | 8173 | -0.003 | -0.1728 | No | ||
| 55 | SYT5 | 17986 | 8182 | -0.003 | -0.1733 | No | ||
| 56 | PPIE | 15766 | 8285 | -0.004 | -0.1788 | No | ||
| 57 | NDUFS1 | 13940 | 8627 | -0.008 | -0.1972 | No | ||
| 58 | SRP19 | 2004 12341 | 8903 | -0.012 | -0.2120 | No | ||
| 59 | DNAJA2 | 12133 18806 | 9035 | -0.014 | -0.2190 | No | ||
| 60 | SHC1 | 9813 9812 5430 | 9137 | -0.016 | -0.2244 | No | ||
| 61 | FBXO22 | 19442 | 9141 | -0.016 | -0.2245 | No | ||
| 62 | RBM22 | 23542 | 9160 | -0.016 | -0.2254 | No | ||
| 63 | FMR1 | 24321 | 9187 | -0.016 | -0.2268 | No | ||
| 64 | STX12 | 4140 | 9383 | -0.019 | -0.2372 | No | ||
| 65 | INSIG2 | 3944 4124 4073 | 9534 | -0.021 | -0.2453 | No | ||
| 66 | TRIM44 | 8164 13487 | 9617 | -0.023 | -0.2496 | No | ||
| 67 | ACTR2 | 20523 | 9775 | -0.025 | -0.2580 | No | ||
| 68 | CDC45L | 22642 1752 | 9842 | -0.026 | -0.2614 | No | ||
| 69 | OGG1 | 17331 | 10204 | -0.032 | -0.2808 | No | ||
| 70 | LSM5 | 7286 | 10421 | -0.036 | -0.2924 | No | ||
| 71 | FOXH1 | 22238 | 10491 | -0.037 | -0.2959 | No | ||
| 72 | CLDN7 | 20817 | 10609 | -0.039 | -0.3021 | No | ||
| 73 | CPSF3 | 7038 | 11212 | -0.051 | -0.3345 | No | ||
| 74 | IMMT | 367 8029 | 11818 | -0.066 | -0.3669 | No | ||
| 75 | GTF2A2 | 10654 | 12265 | -0.082 | -0.3907 | No | ||
| 76 | FBXO36 | 14205 | 12355 | -0.087 | -0.3951 | No | ||
| 77 | LEPRE1 | 2384 12122 | 12401 | -0.089 | -0.3972 | No | ||
| 78 | EN1 | 387 14155 | 12477 | -0.092 | -0.4008 | No | ||
| 79 | GGTL3 | 14381 | 12908 | -0.113 | -0.4236 | No | ||
| 80 | EPN1 | 18401 | 13314 | -0.140 | -0.4448 | No | ||
| 81 | RPL19 | 9736 | 13396 | -0.146 | -0.4486 | No | ||
| 82 | GABARAPL2 | 8211 13552 | 13523 | -0.158 | -0.4547 | No | ||
| 83 | ARHGAP1 | 6001 10448 | 13740 | -0.179 | -0.4656 | No | ||
| 84 | TCF7 | 1467 20466 | 13747 | -0.180 | -0.4651 | No | ||
| 85 | SNRPB | 9842 5469 2736 | 13755 | -0.181 | -0.4646 | No | ||
| 86 | ZNF23 | 6149 | 13936 | -0.199 | -0.4735 | No | ||
| 87 | CLDN5 | 22826 | 14338 | -0.253 | -0.4940 | No | ||
| 88 | HMGA1 | 23323 | 14469 | -0.276 | -0.4998 | No | ||
| 89 | RAPH1 | 19110 | 14598 | -0.306 | -0.5053 | No | ||
| 90 | BAT3 | 5896 | 14606 | -0.307 | -0.5043 | No | ||
| 91 | RPL37 | 12502 7421 22521 | 14614 | -0.310 | -0.5033 | No | ||
| 92 | DHX30 | 18989 | 14660 | -0.320 | -0.5043 | No | ||
| 93 | PEX11A | 1056 5241 | 14758 | -0.347 | -0.5079 | No | ||
| 94 | IL2RG | 24096 | 15069 | -0.442 | -0.5227 | No | ||
| 95 | ZDHHC5 | 14547 | 15079 | -0.445 | -0.5212 | No | ||
| 96 | PHKB | 18537 3873 | 15115 | -0.460 | -0.5210 | No | ||
| 97 | TBC1D17 | 6121 | 15220 | -0.501 | -0.5243 | No | ||
| 98 | MARK3 | 9369 | 15234 | -0.507 | -0.5227 | No | ||
| 99 | SAMD8 | 22082 | 15241 | -0.510 | -0.5207 | No | ||
| 100 | TRAF7 | 10325 | 15315 | -0.542 | -0.5222 | No | ||
| 101 | SMUG1 | 22109 | 15743 | -0.818 | -0.5416 | No | ||
| 102 | UBE2L3 | 5818 22647 | 15809 | -0.865 | -0.5411 | No | ||
| 103 | PCQAP | 1661 13581 | 15829 | -0.880 | -0.5382 | No | ||
| 104 | POLL | 23658 3688 | 15940 | -0.971 | -0.5397 | No | ||
| 105 | CHERP | 11367 | 16081 | -1.040 | -0.5425 | No | ||
| 106 | DDOST | 16021 | 16255 | -1.186 | -0.5465 | Yes | ||
| 107 | STOML2 | 15902 | 16273 | -1.202 | -0.5420 | Yes | ||
| 108 | NKIRAS2 | 12979 | 16291 | -1.219 | -0.5373 | Yes | ||
| 109 | PSMC6 | 7379 | 16310 | -1.237 | -0.5327 | Yes | ||
| 110 | NRAS | 5191 | 16449 | -1.395 | -0.5338 | Yes | ||
| 111 | UFD1L | 22825 | 16559 | -1.518 | -0.5328 | Yes | ||
| 112 | ARCN1 | 19143 | 16565 | -1.525 | -0.5261 | Yes | ||
| 113 | AP1M1 | 18571 | 16582 | -1.540 | -0.5199 | Yes | ||
| 114 | AKT1S1 | 12557 | 16605 | -1.556 | -0.5140 | Yes | ||
| 115 | TSTA3 | 22253 | 16714 | -1.648 | -0.5124 | Yes | ||
| 116 | MTPN | 17183 | 16779 | -1.716 | -0.5080 | Yes | ||
| 117 | HAT1 | 14990 | 16998 | -2.028 | -0.5106 | Yes | ||
| 118 | PSMC1 | 9635 | 17026 | -2.067 | -0.5026 | Yes | ||
| 119 | LMAN2 | 21454 | 17204 | -2.304 | -0.5017 | Yes | ||
| 120 | ATP5A1 | 23505 | 17266 | -2.447 | -0.4939 | Yes | ||
| 121 | DIABLO | 16375 | 17272 | -2.465 | -0.4829 | Yes | ||
| 122 | EIF4G1 | 22818 | 17339 | -2.599 | -0.4747 | Yes | ||
| 123 | MTBP | 22473 8374 | 17432 | -2.763 | -0.4671 | Yes | ||
| 124 | SMS | 2573 24023 | 17445 | -2.797 | -0.4550 | Yes | ||
| 125 | TOMM40 | 6997 | 17562 | -3.010 | -0.4475 | Yes | ||
| 126 | HSPC171 | 18494 | 17574 | -3.032 | -0.4343 | Yes | ||
| 127 | PSMD12 | 20621 | 17621 | -3.111 | -0.4226 | Yes | ||
| 128 | UBADC1 | 2962 13608 | 17676 | -3.265 | -0.4107 | Yes | ||
| 129 | PSMC2 | 16909 | 17677 | -3.270 | -0.3958 | Yes | ||
| 130 | DNAJC7 | 20227 | 17784 | -3.518 | -0.3855 | Yes | ||
| 131 | FANCD2 | 17326 464 | 17793 | -3.537 | -0.3698 | Yes | ||
| 132 | KLHDC3 | 22956 | 17831 | -3.644 | -0.3552 | Yes | ||
| 133 | MRPL13 | 2249 22294 | 17872 | -3.763 | -0.3403 | Yes | ||
| 134 | TRMT1 | 2307 12985 | 17888 | -3.823 | -0.3237 | Yes | ||
| 135 | SDHD | 19125 | 18022 | -4.265 | -0.3114 | Yes | ||
| 136 | MIR16 | 17663 1354 | 18084 | -4.529 | -0.2941 | Yes | ||
| 137 | SLC30A7 | 15186 | 18085 | -4.530 | -0.2735 | Yes | ||
| 138 | PRKAG2 | 24 | 18124 | -4.685 | -0.2542 | Yes | ||
| 139 | EEF1B2 | 4131 12063 | 18152 | -4.785 | -0.2338 | Yes | ||
| 140 | MAPKAP1 | 15030 5993 | 18188 | -4.914 | -0.2134 | Yes | ||
| 141 | TIMM8B | 11391 | 18205 | -4.967 | -0.1916 | Yes | ||
| 142 | TUFM | 10608 | 18239 | -5.070 | -0.1703 | Yes | ||
| 143 | RFC4 | 1735 22627 | 18260 | -5.128 | -0.1480 | Yes | ||
| 144 | EIF5A | 11345 20379 6590 | 18392 | -5.951 | -0.1280 | Yes | ||
| 145 | GART | 22543 1754 | 18431 | -6.196 | -0.1018 | Yes | ||
| 146 | IPO4 | 7983 | 18462 | -6.540 | -0.0737 | Yes | ||
| 147 | EXTL2 | 15440 | 18514 | -7.089 | -0.0441 | Yes | ||
| 148 | MRPL3 | 3116 19335 | 18611 | -10.888 | 0.0003 | Yes |