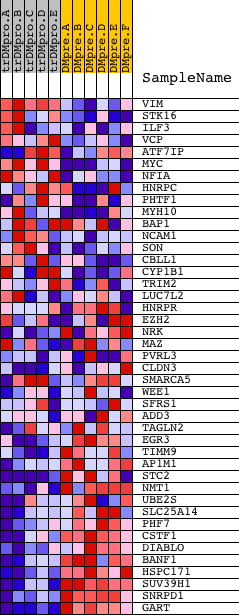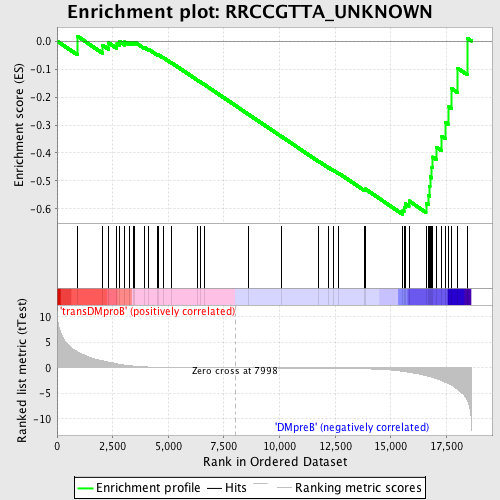
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
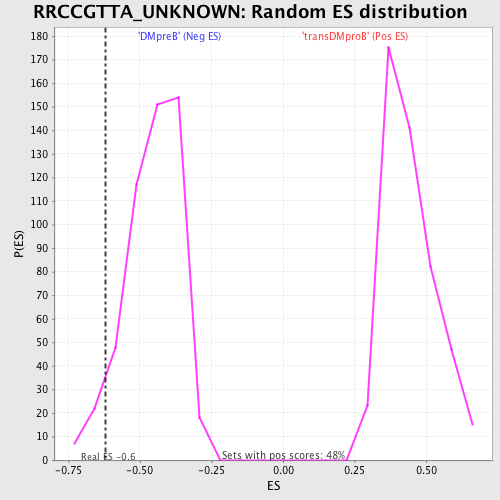
| GeneSet | RRCCGTTA_UNKNOWN |
| Enrichment Score (ES) | -0.6198013 |
| Normalized Enrichment Score (NES) | -1.3672391 |
| Nominal p-value | 0.056092843 |
| FDR q-value | 0.38120714 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VIM | 80 | 907 | 3.168 | 0.0177 | No | ||
| 2 | STK16 | 14216 | 2035 | 1.372 | -0.0142 | No | ||
| 3 | ILF3 | 3110 3030 9176 | 2324 | 1.096 | -0.0067 | No | ||
| 4 | VCP | 11254 | 2689 | 0.797 | -0.0096 | No | ||
| 5 | ATF7IP | 12028 13038 | 2811 | 0.682 | -0.0018 | No | ||
| 6 | MYC | 22465 9435 | 3045 | 0.531 | -0.0032 | No | ||
| 7 | NFIA | 16172 5170 | 3241 | 0.430 | -0.0047 | No | ||
| 8 | HNRPC | 9102 4859 | 3416 | 0.354 | -0.0066 | No | ||
| 9 | PHTF1 | 15469 | 3494 | 0.323 | -0.0040 | No | ||
| 10 | MYH10 | 8077 | 3920 | 0.200 | -0.0227 | No | ||
| 11 | BAP1 | 22057 | 4112 | 0.162 | -0.0296 | No | ||
| 12 | NCAM1 | 5149 | 4518 | 0.105 | -0.0492 | No | ||
| 13 | SON | 5473 1657 1684 | 4552 | 0.100 | -0.0488 | No | ||
| 14 | CBLL1 | 4188 | 4767 | 0.081 | -0.0587 | No | ||
| 15 | CYP1B1 | 4581 | 5159 | 0.060 | -0.0784 | No | ||
| 16 | TRIM2 | 13485 8157 | 6306 | 0.026 | -0.1396 | No | ||
| 17 | LUC7L2 | 5313 5312 | 6449 | 0.023 | -0.1467 | No | ||
| 18 | HNRPR | 13196 7909 2500 | 6617 | 0.020 | -0.1553 | No | ||
| 19 | EZH2 | 1092 17163 | 8583 | -0.008 | -0.2609 | No | ||
| 20 | NRK | 6559 | 10107 | -0.030 | -0.3423 | No | ||
| 21 | MAZ | 1327 17623 | 11744 | -0.064 | -0.4290 | No | ||
| 22 | PVRL3 | 1674 22579 1675 | 12215 | -0.080 | -0.4527 | No | ||
| 23 | CLDN3 | 16680 | 12429 | -0.090 | -0.4622 | No | ||
| 24 | SMARCA5 | 8215 | 12643 | -0.099 | -0.4716 | No | ||
| 25 | WEE1 | 18127 | 13796 | -0.184 | -0.5298 | No | ||
| 26 | SFRS1 | 8492 | 13847 | -0.190 | -0.5285 | No | ||
| 27 | ADD3 | 23812 3694 | 15544 | -0.676 | -0.6056 | Yes | ||
| 28 | TAGLN2 | 14043 | 15596 | -0.708 | -0.5935 | Yes | ||
| 29 | EGR3 | 4656 | 15650 | -0.741 | -0.5808 | Yes | ||
| 30 | TIMM9 | 19686 | 15817 | -0.873 | -0.5714 | Yes | ||
| 31 | AP1M1 | 18571 | 16582 | -1.540 | -0.5802 | Yes | ||
| 32 | STC2 | 5526 | 16684 | -1.618 | -0.5517 | Yes | ||
| 33 | NMT1 | 20638 | 16738 | -1.674 | -0.5194 | Yes | ||
| 34 | UBE2S | 17979 | 16761 | -1.697 | -0.4850 | Yes | ||
| 35 | SLC25A14 | 24344 | 16846 | -1.807 | -0.4516 | Yes | ||
| 36 | PHF7 | 21890 | 16880 | -1.861 | -0.4143 | Yes | ||
| 37 | CSTF1 | 12515 | 17048 | -2.098 | -0.3793 | Yes | ||
| 38 | DIABLO | 16375 | 17272 | -2.465 | -0.3395 | Yes | ||
| 39 | BANF1 | 6216 10706 3760 | 17446 | -2.797 | -0.2901 | Yes | ||
| 40 | HSPC171 | 18494 | 17574 | -3.032 | -0.2333 | Yes | ||
| 41 | SUV39H1 | 24194 | 17743 | -3.415 | -0.1707 | Yes | ||
| 42 | SNRPD1 | 23622 | 17992 | -4.171 | -0.0965 | Yes | ||
| 43 | GART | 22543 1754 | 18431 | -6.196 | 0.0100 | Yes |