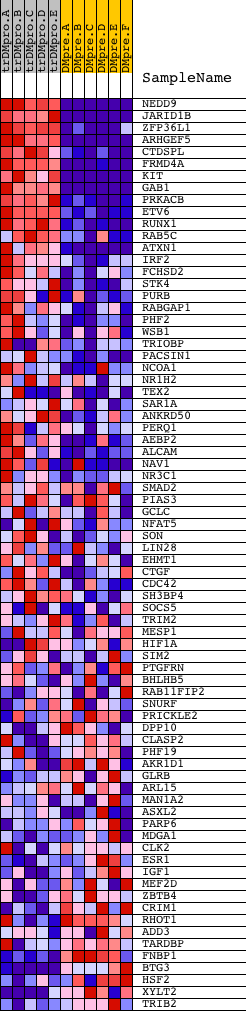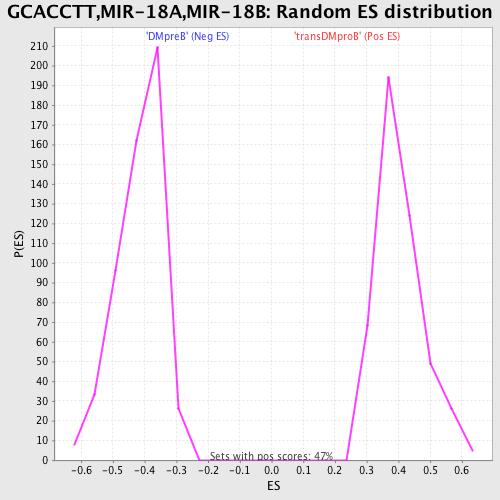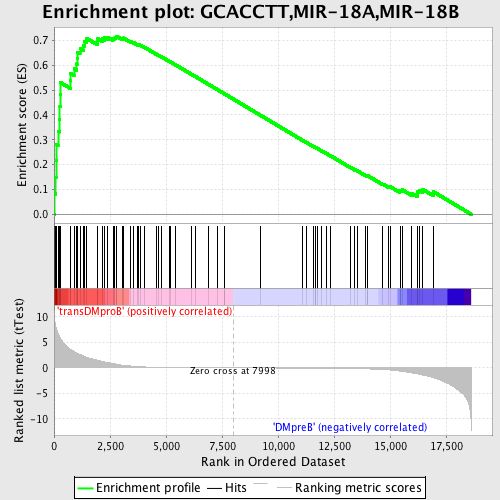
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | GCACCTT,MIR-18A,MIR-18B |
| Enrichment Score (ES) | 0.7176614 |
| Normalized Enrichment Score (NES) | 1.781936 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.011416538 |
| FWER p-Value | 0.013 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NEDD9 | 3206 21483 | 16 | 9.974 | 0.0841 | Yes | ||
| 2 | JARID1B | 14123 | 75 | 8.103 | 0.1501 | Yes | ||
| 3 | ZFP36L1 | 4453 4454 | 88 | 7.945 | 0.2171 | Yes | ||
| 4 | ARHGEF5 | 12025 | 106 | 7.710 | 0.2819 | Yes | ||
| 5 | CTDSPL | 19279 | 180 | 6.657 | 0.3347 | Yes | ||
| 6 | FRMD4A | 5557 | 244 | 6.127 | 0.3835 | Yes | ||
| 7 | KIT | 16823 | 259 | 5.993 | 0.4339 | Yes | ||
| 8 | GAB1 | 18828 | 276 | 5.806 | 0.4825 | Yes | ||
| 9 | PRKACB | 15140 | 286 | 5.707 | 0.5306 | Yes | ||
| 10 | ETV6 | 17264 | 737 | 3.610 | 0.5371 | Yes | ||
| 11 | RUNX1 | 4481 | 743 | 3.588 | 0.5674 | Yes | ||
| 12 | RAB5C | 20226 | 892 | 3.201 | 0.5867 | Yes | ||
| 13 | ATXN1 | 21477 | 1007 | 2.918 | 0.6055 | Yes | ||
| 14 | IRF2 | 18621 | 1027 | 2.872 | 0.6289 | Yes | ||
| 15 | FCHSD2 | 18172 | 1039 | 2.856 | 0.6527 | Yes | ||
| 16 | STK4 | 14743 | 1158 | 2.598 | 0.6684 | Yes | ||
| 17 | PURB | 5335 | 1325 | 2.278 | 0.6789 | Yes | ||
| 18 | RABGAP1 | 10439 | 1353 | 2.227 | 0.6964 | Yes | ||
| 19 | PHF2 | 21470 | 1448 | 2.064 | 0.7089 | Yes | ||
| 20 | WSB1 | 20330 | 1938 | 1.473 | 0.6951 | Yes | ||
| 21 | TRIOBP | 2280 670 8476 | 1953 | 1.455 | 0.7068 | Yes | ||
| 22 | PACSIN1 | 6257 23322 10744 | 2152 | 1.266 | 0.7069 | Yes | ||
| 23 | NCOA1 | 5154 | 2258 | 1.157 | 0.7111 | Yes | ||
| 24 | NR1H2 | 17846 | 2387 | 1.040 | 0.7130 | Yes | ||
| 25 | TEX2 | 5725 20181 20182 | 2629 | 0.846 | 0.7072 | Yes | ||
| 26 | SAR1A | 20008 | 2699 | 0.782 | 0.7102 | Yes | ||
| 27 | ANKRD50 | 533 | 2763 | 0.723 | 0.7130 | Yes | ||
| 28 | PERQ1 | 7168 | 2788 | 0.705 | 0.7177 | Yes | ||
| 29 | AEBP2 | 4359 | 3063 | 0.518 | 0.7073 | No | ||
| 30 | ALCAM | 4367 | 3090 | 0.505 | 0.7102 | No | ||
| 31 | NAV1 | 3978 5681 | 3402 | 0.360 | 0.6965 | No | ||
| 32 | NR3C1 | 9043 | 3543 | 0.304 | 0.6915 | No | ||
| 33 | SMAD2 | 23511 | 3741 | 0.245 | 0.6830 | No | ||
| 34 | PIAS3 | 15491 672 1906 | 3768 | 0.238 | 0.6836 | No | ||
| 35 | GCLC | 19374 | 3842 | 0.221 | 0.6816 | No | ||
| 36 | NFAT5 | 3921 7037 12036 | 4055 | 0.171 | 0.6716 | No | ||
| 37 | SON | 5473 1657 1684 | 4552 | 0.100 | 0.6457 | No | ||
| 38 | LIN28 | 15723 | 4666 | 0.090 | 0.6404 | No | ||
| 39 | EHMT1 | 13401 14670 2756 8090 | 4810 | 0.079 | 0.6333 | No | ||
| 40 | CTGF | 20068 | 5148 | 0.061 | 0.6157 | No | ||
| 41 | CDC42 | 4503 8722 4504 2465 | 5195 | 0.059 | 0.6137 | No | ||
| 42 | SH3BP4 | 8270 | 5415 | 0.049 | 0.6023 | No | ||
| 43 | SOCS5 | 1619 23141 | 6137 | 0.030 | 0.5637 | No | ||
| 44 | TRIM2 | 13485 8157 | 6306 | 0.026 | 0.5548 | No | ||
| 45 | MESP1 | 17786 | 6900 | 0.015 | 0.5230 | No | ||
| 46 | HIF1A | 4850 | 7306 | 0.009 | 0.5012 | No | ||
| 47 | SIM2 | 22693 | 7604 | 0.005 | 0.4852 | No | ||
| 48 | PTGFRN | 15224 | 7620 | 0.005 | 0.4845 | No | ||
| 49 | BHLHB5 | 15629 | 9205 | -0.017 | 0.3992 | No | ||
| 50 | RAB11FIP2 | 23635 | 11086 | -0.048 | 0.2982 | No | ||
| 51 | SNURF | 2251 | 11283 | -0.052 | 0.2881 | No | ||
| 52 | PRICKLE2 | 10837 | 11559 | -0.059 | 0.2737 | No | ||
| 53 | DPP10 | 13850 | 11676 | -0.062 | 0.2680 | No | ||
| 54 | CLASP2 | 13338 | 11689 | -0.062 | 0.2679 | No | ||
| 55 | PHF19 | 13150 | 11777 | -0.065 | 0.2638 | No | ||
| 56 | AKR1D1 | 17483 | 11954 | -0.071 | 0.2549 | No | ||
| 57 | GLRB | 15312 1846 | 12154 | -0.078 | 0.2448 | No | ||
| 58 | ARL15 | 21559 | 12343 | -0.086 | 0.2354 | No | ||
| 59 | MAN1A2 | 5076 5075 | 13233 | -0.135 | 0.1886 | No | ||
| 60 | ASXL2 | 7957 | 13430 | -0.149 | 0.1793 | No | ||
| 61 | PARP6 | 19419 2979 3156 | 13536 | -0.159 | 0.1750 | No | ||
| 62 | MDGA1 | 13231 | 13898 | -0.194 | 0.1572 | No | ||
| 63 | CLK2 | 1883 15544 | 13983 | -0.204 | 0.1544 | No | ||
| 64 | ESR1 | 20097 4685 | 14010 | -0.207 | 0.1547 | No | ||
| 65 | IGF1 | 3352 9156 3409 | 14678 | -0.326 | 0.1215 | No | ||
| 66 | MEF2D | 9379 | 14907 | -0.385 | 0.1125 | No | ||
| 67 | ZBTB4 | 13267 | 14995 | -0.412 | 0.1113 | No | ||
| 68 | CRIM1 | 403 | 15440 | -0.622 | 0.0927 | No | ||
| 69 | RHOT1 | 12227 | 15467 | -0.638 | 0.0967 | No | ||
| 70 | ADD3 | 23812 3694 | 15544 | -0.676 | 0.0984 | No | ||
| 71 | TARDBP | 10520 6066 | 15969 | -0.990 | 0.0840 | No | ||
| 72 | FNBP1 | 4732 | 16216 | -1.150 | 0.0805 | No | ||
| 73 | BTG3 | 8664 | 16219 | -1.154 | 0.0902 | No | ||
| 74 | HSF2 | 9128 | 16290 | -1.219 | 0.0968 | No | ||
| 75 | XYLT2 | 20288 | 16461 | -1.417 | 0.0997 | No | ||
| 76 | TRIB2 | 21102 | 16928 | -1.928 | 0.0910 | No |