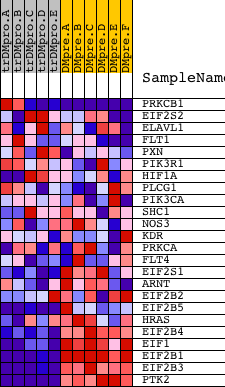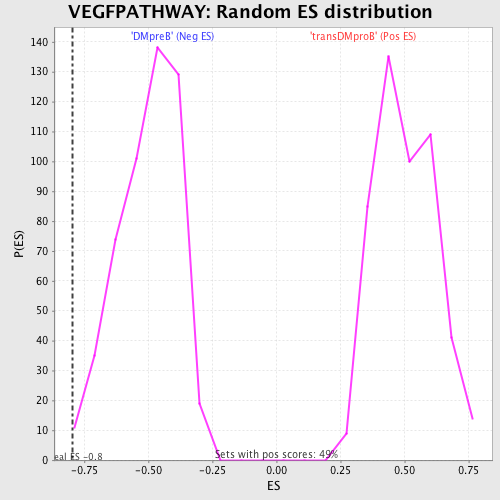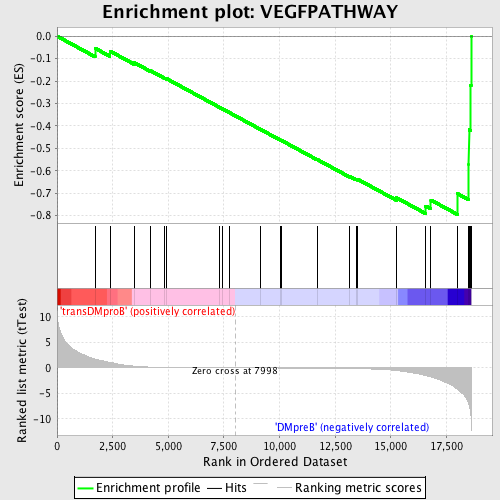
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | VEGFPATHWAY |
| Enrichment Score (ES) | -0.7945105 |
| Normalized Enrichment Score (NES) | -1.5832909 |
| Nominal p-value | 0.00591716 |
| FDR q-value | 0.074213825 |
| FWER p-Value | 0.595 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PRKCB1 | 1693 9574 | 1709 | 1.712 | -0.0540 | No | ||
| 2 | EIF2S2 | 7406 14383 | 2377 | 1.051 | -0.0665 | No | ||
| 3 | ELAVL1 | 4888 | 3484 | 0.327 | -0.1188 | No | ||
| 4 | FLT1 | 3483 16287 | 4189 | 0.149 | -0.1533 | No | ||
| 5 | PXN | 5339 3573 | 4837 | 0.077 | -0.1864 | No | ||
| 6 | PIK3R1 | 3170 | 4927 | 0.072 | -0.1896 | No | ||
| 7 | HIF1A | 4850 | 7306 | 0.009 | -0.3173 | No | ||
| 8 | PLCG1 | 14753 | 7414 | 0.008 | -0.3229 | No | ||
| 9 | PIK3CA | 9562 | 7760 | 0.003 | -0.3414 | No | ||
| 10 | SHC1 | 9813 9812 5430 | 9137 | -0.016 | -0.4150 | No | ||
| 11 | NOS3 | 16906 885 | 10060 | -0.030 | -0.4640 | No | ||
| 12 | KDR | 16509 | 10088 | -0.030 | -0.4648 | No | ||
| 13 | PRKCA | 20174 | 11681 | -0.062 | -0.5490 | No | ||
| 14 | FLT4 | 20904 1461 | 13146 | -0.128 | -0.6249 | No | ||
| 15 | EIF2S1 | 4658 | 13442 | -0.150 | -0.6375 | No | ||
| 16 | ARNT | 4413 1857 | 13512 | -0.157 | -0.6377 | No | ||
| 17 | EIF2B2 | 21204 | 15254 | -0.516 | -0.7199 | No | ||
| 18 | EIF2B5 | 1719 22822 | 16569 | -1.528 | -0.7567 | Yes | ||
| 19 | HRAS | 4868 | 16802 | -1.753 | -0.7303 | Yes | ||
| 20 | EIF2B4 | 16574 | 17996 | -4.187 | -0.7017 | Yes | ||
| 21 | EIF1 | 23563 | 18509 | -7.038 | -0.5733 | Yes | ||
| 22 | EIF2B1 | 16368 3458 | 18513 | -7.086 | -0.4164 | Yes | ||
| 23 | EIF2B3 | 16118 | 18593 | -9.059 | -0.2199 | Yes | ||
| 24 | PTK2 | 22271 | 18609 | -9.978 | 0.0004 | Yes |