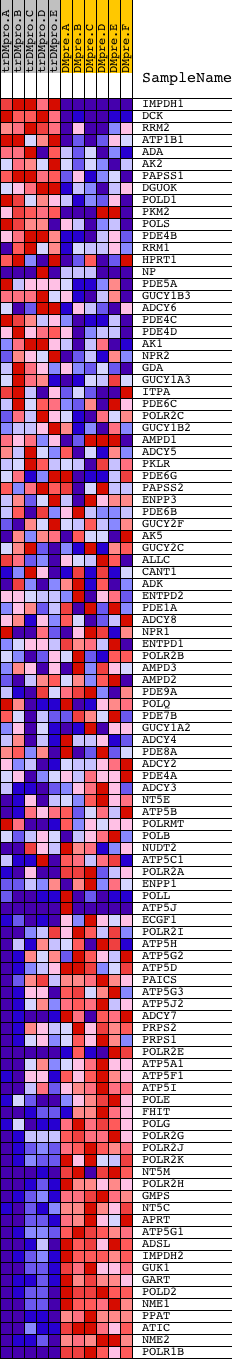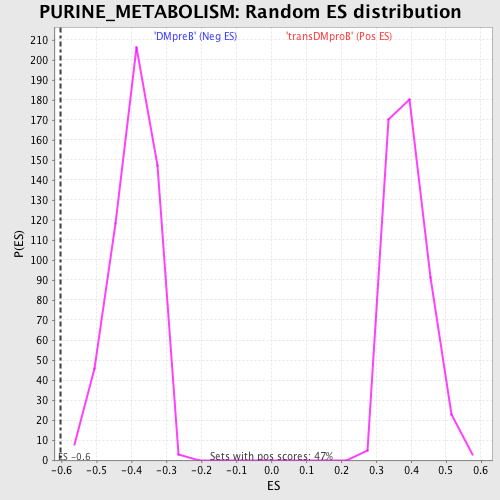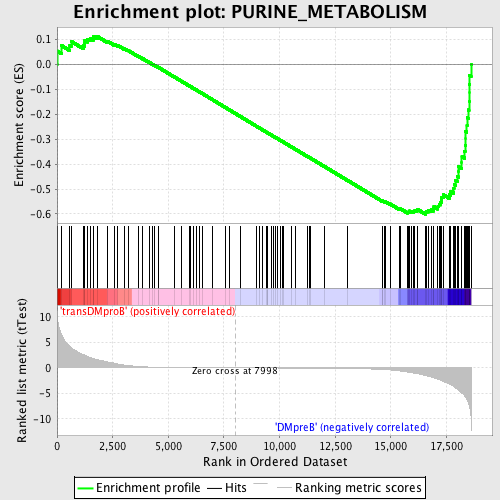
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | PURINE_METABOLISM |
| Enrichment Score (ES) | -0.6020656 |
| Normalized Enrichment Score (NES) | -1.5292982 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.12532222 |
| FWER p-Value | 0.885 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IMPDH1 | 17197 1131 | 3 | 10.857 | 0.0529 | No | ||
| 2 | DCK | 16808 | 192 | 6.569 | 0.0748 | No | ||
| 3 | RRM2 | 5401 5400 | 571 | 4.235 | 0.0751 | No | ||
| 4 | ATP1B1 | 4420 | 631 | 3.969 | 0.0913 | No | ||
| 5 | ADA | 2703 14361 | 1193 | 2.543 | 0.0734 | No | ||
| 6 | AK2 | 8563 2479 16073 | 1221 | 2.493 | 0.0841 | No | ||
| 7 | PAPSS1 | 15419 | 1245 | 2.431 | 0.0948 | No | ||
| 8 | DGUOK | 17099 1041 | 1361 | 2.209 | 0.0994 | No | ||
| 9 | POLD1 | 17847 | 1481 | 2.020 | 0.1028 | No | ||
| 10 | PKM2 | 3642 9573 | 1619 | 1.831 | 0.1044 | No | ||
| 11 | POLS | 9963 | 1634 | 1.812 | 0.1125 | No | ||
| 12 | PDE4B | 9543 | 1822 | 1.611 | 0.1102 | No | ||
| 13 | RRM1 | 18163 | 2252 | 1.164 | 0.0927 | No | ||
| 14 | HPRT1 | 2655 24339 408 | 2601 | 0.873 | 0.0782 | No | ||
| 15 | NP | 22027 9597 5273 5274 | 2722 | 0.766 | 0.0755 | No | ||
| 16 | PDE5A | 15431 | 3034 | 0.537 | 0.0613 | No | ||
| 17 | GUCY1B3 | 15311 | 3192 | 0.453 | 0.0550 | No | ||
| 18 | ADCY6 | 22139 2283 8551 | 3656 | 0.270 | 0.0313 | No | ||
| 19 | PDE4C | 8481 | 3857 | 0.217 | 0.0216 | No | ||
| 20 | PDE4D | 10722 6235 | 4135 | 0.158 | 0.0074 | No | ||
| 21 | AK1 | 4363 | 4308 | 0.133 | -0.0013 | No | ||
| 22 | NPR2 | 16230 | 4395 | 0.119 | -0.0053 | No | ||
| 23 | GDA | 4764 | 4568 | 0.098 | -0.0141 | No | ||
| 24 | GUCY1A3 | 7216 12245 15310 | 4572 | 0.098 | -0.0138 | No | ||
| 25 | ITPA | 9193 | 5267 | 0.056 | -0.0510 | No | ||
| 26 | PDE6C | 8496 23868 | 5587 | 0.044 | -0.0680 | No | ||
| 27 | POLR2C | 9750 | 5936 | 0.034 | -0.0867 | No | ||
| 28 | GUCY1B2 | 21794 | 6015 | 0.032 | -0.0907 | No | ||
| 29 | AMPD1 | 5051 | 6145 | 0.030 | -0.0976 | No | ||
| 30 | ADCY5 | 10312 | 6281 | 0.027 | -0.1047 | No | ||
| 31 | PKLR | 1850 15545 | 6419 | 0.024 | -0.1120 | No | ||
| 32 | PDE6G | 20119 | 6539 | 0.022 | -0.1183 | No | ||
| 33 | PAPSS2 | 6258 | 6978 | 0.014 | -0.1419 | No | ||
| 34 | ENPP3 | 19803 | 7572 | 0.005 | -0.1739 | No | ||
| 35 | PDE6B | 16760 | 7730 | 0.003 | -0.1824 | No | ||
| 36 | GUCY2F | 24045 | 8251 | -0.003 | -0.2105 | No | ||
| 37 | AK5 | 6042 | 8951 | -0.013 | -0.2482 | No | ||
| 38 | GUCY2C | 1140 16954 | 9096 | -0.015 | -0.2559 | No | ||
| 39 | ALLC | 21093 2130 | 9250 | -0.017 | -0.2640 | No | ||
| 40 | CANT1 | 13304 | 9398 | -0.019 | -0.2719 | No | ||
| 41 | ADK | 3302 3454 8555 | 9441 | -0.020 | -0.2741 | No | ||
| 42 | ENTPD2 | 8713 | 9449 | -0.020 | -0.2743 | No | ||
| 43 | PDE1A | 9541 5234 | 9653 | -0.023 | -0.2852 | No | ||
| 44 | ADCY8 | 22281 | 9741 | -0.024 | -0.2898 | No | ||
| 45 | NPR1 | 9480 | 9834 | -0.026 | -0.2946 | No | ||
| 46 | ENTPD1 | 4495 | 9917 | -0.027 | -0.2989 | No | ||
| 47 | POLR2B | 16817 | 10054 | -0.030 | -0.3061 | No | ||
| 48 | AMPD3 | 4383 2110 | 10111 | -0.030 | -0.3090 | No | ||
| 49 | AMPD2 | 1931 15197 | 10189 | -0.032 | -0.3130 | No | ||
| 50 | PDE9A | 23301 | 10515 | -0.037 | -0.3304 | No | ||
| 51 | POLQ | 13407 22768 | 10704 | -0.040 | -0.3403 | No | ||
| 52 | PDE7B | 19807 | 11236 | -0.051 | -0.3688 | No | ||
| 53 | GUCY1A2 | 19580 | 11338 | -0.054 | -0.3740 | No | ||
| 54 | ADCY4 | 2980 21818 | 11398 | -0.055 | -0.3769 | No | ||
| 55 | PDE8A | 18202 | 12014 | -0.073 | -0.4097 | No | ||
| 56 | ADCY2 | 21412 | 13035 | -0.121 | -0.4643 | No | ||
| 57 | PDE4A | 3037 19542 3083 3023 | 14605 | -0.307 | -0.5475 | No | ||
| 58 | ADCY3 | 21329 894 | 14643 | -0.316 | -0.5480 | No | ||
| 59 | NT5E | 19360 18702 | 14728 | -0.340 | -0.5508 | No | ||
| 60 | ATP5B | 19846 | 14751 | -0.346 | -0.5503 | No | ||
| 61 | POLRMT | 19705 | 14966 | -0.404 | -0.5599 | No | ||
| 62 | POLB | 9599 | 15405 | -0.596 | -0.5807 | No | ||
| 63 | NUDT2 | 16239 | 15441 | -0.622 | -0.5795 | No | ||
| 64 | ATP5C1 | 8635 | 15756 | -0.827 | -0.5924 | No | ||
| 65 | POLR2A | 5394 | 15814 | -0.870 | -0.5913 | No | ||
| 66 | ENPP1 | 19804 | 15834 | -0.885 | -0.5880 | No | ||
| 67 | POLL | 23658 3688 | 15940 | -0.971 | -0.5889 | No | ||
| 68 | ATP5J | 880 22555 | 16024 | -1.008 | -0.5885 | No | ||
| 69 | ECGF1 | 22160 | 16080 | -1.039 | -0.5863 | No | ||
| 70 | POLR2I | 12839 | 16178 | -1.117 | -0.5861 | No | ||
| 71 | ATP5H | 12948 | 16211 | -1.149 | -0.5822 | No | ||
| 72 | ATP5G2 | 12610 | 16579 | -1.540 | -0.5945 | Yes | ||
| 73 | ATP5D | 19949 | 16608 | -1.558 | -0.5884 | Yes | ||
| 74 | PAICS | 16820 | 16678 | -1.613 | -0.5843 | Yes | ||
| 75 | ATP5G3 | 14558 | 16819 | -1.777 | -0.5832 | Yes | ||
| 76 | ATP5J2 | 12186 | 16915 | -1.907 | -0.5790 | Yes | ||
| 77 | ADCY7 | 8552 448 | 16937 | -1.943 | -0.5706 | Yes | ||
| 78 | PRPS2 | 24003 | 17118 | -2.200 | -0.5696 | Yes | ||
| 79 | PRPS1 | 24233 | 17165 | -2.238 | -0.5611 | Yes | ||
| 80 | POLR2E | 3325 19699 | 17221 | -2.334 | -0.5527 | Yes | ||
| 81 | ATP5A1 | 23505 | 17266 | -2.447 | -0.5431 | Yes | ||
| 82 | ATP5F1 | 15212 | 17289 | -2.498 | -0.5321 | Yes | ||
| 83 | ATP5I | 8637 | 17360 | -2.639 | -0.5230 | Yes | ||
| 84 | POLE | 16755 | 17631 | -3.130 | -0.5223 | Yes | ||
| 85 | FHIT | 21920 | 17694 | -3.306 | -0.5095 | Yes | ||
| 86 | POLG | 17789 | 17827 | -3.636 | -0.4988 | Yes | ||
| 87 | POLR2G | 23753 | 17842 | -3.679 | -0.4816 | Yes | ||
| 88 | POLR2J | 16672 | 17905 | -3.892 | -0.4660 | Yes | ||
| 89 | POLR2K | 9413 | 17988 | -4.160 | -0.4501 | Yes | ||
| 90 | NT5M | 8345 4175 | 18037 | -4.318 | -0.4316 | Yes | ||
| 91 | POLR2H | 10888 | 18043 | -4.355 | -0.4105 | Yes | ||
| 92 | GMPS | 15578 | 18194 | -4.945 | -0.3945 | Yes | ||
| 93 | NT5C | 20151 | 18198 | -4.950 | -0.3705 | Yes | ||
| 94 | APRT | 8620 | 18310 | -5.406 | -0.3500 | Yes | ||
| 95 | ATP5G1 | 8636 | 18339 | -5.580 | -0.3243 | Yes | ||
| 96 | ADSL | 4358 | 18353 | -5.627 | -0.2975 | Yes | ||
| 97 | IMPDH2 | 10730 | 18355 | -5.639 | -0.2700 | Yes | ||
| 98 | GUK1 | 20432 | 18423 | -6.134 | -0.2436 | Yes | ||
| 99 | GART | 22543 1754 | 18431 | -6.196 | -0.2138 | Yes | ||
| 100 | POLD2 | 20537 | 18484 | -6.826 | -0.1832 | Yes | ||
| 101 | NME1 | 9467 | 18515 | -7.106 | -0.1501 | Yes | ||
| 102 | PPAT | 6081 | 18522 | -7.211 | -0.1152 | Yes | ||
| 103 | ATIC | 14231 3968 | 18524 | -7.214 | -0.0800 | Yes | ||
| 104 | NME2 | 9468 | 18538 | -7.449 | -0.0443 | Yes | ||
| 105 | POLR1B | 14857 | 18608 | -9.919 | 0.0004 | Yes |