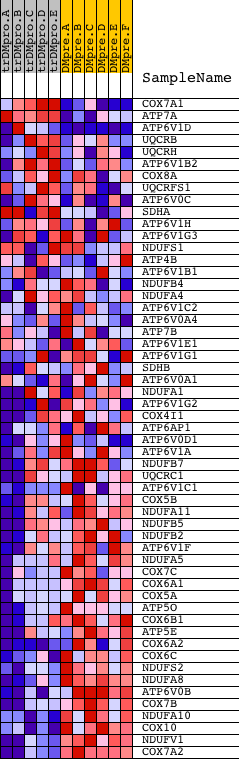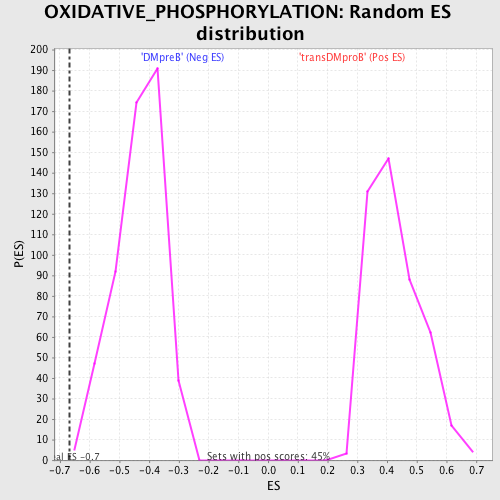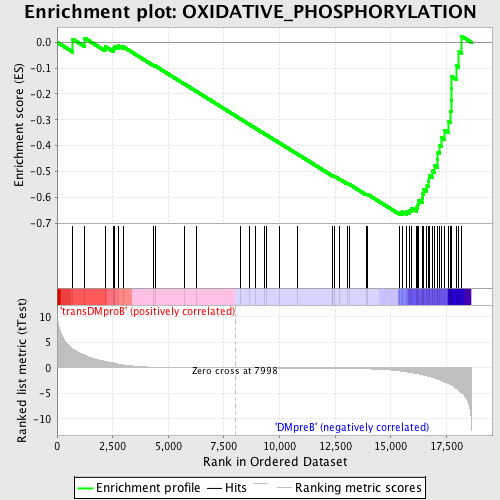
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | -0.66621065 |
| Normalized Enrichment Score (NES) | -1.5480462 |
| Nominal p-value | 0.0018248175 |
| FDR q-value | 0.100151226 |
| FWER p-Value | 0.806 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COX7A1 | 4554 | 709 | 3.674 | 0.0116 | No | ||
| 2 | ATP7A | 4425 | 1239 | 2.443 | 0.0162 | No | ||
| 3 | ATP6V1D | 7872 | 2157 | 1.261 | -0.0162 | No | ||
| 4 | UQCRB | 12545 | 2534 | 0.936 | -0.0238 | No | ||
| 5 | UQCRH | 12378 | 2594 | 0.878 | -0.0150 | No | ||
| 6 | ATP6V1B2 | 18599 | 2743 | 0.750 | -0.0129 | No | ||
| 7 | COX8A | 8777 | 2967 | 0.573 | -0.0171 | No | ||
| 8 | UQCRFS1 | 21499 | 4321 | 0.132 | -0.0882 | No | ||
| 9 | ATP6V0C | 8643 | 4403 | 0.119 | -0.0910 | No | ||
| 10 | SDHA | 12436 | 5741 | 0.039 | -0.1625 | No | ||
| 11 | ATP6V1H | 14301 | 6270 | 0.027 | -0.1906 | No | ||
| 12 | ATP6V1G3 | 14107 | 8243 | -0.003 | -0.2967 | No | ||
| 13 | NDUFS1 | 13940 | 8627 | -0.008 | -0.3173 | No | ||
| 14 | ATP4B | 18928 | 8912 | -0.012 | -0.3324 | No | ||
| 15 | ATP6V1B1 | 8499 | 9323 | -0.018 | -0.3542 | No | ||
| 16 | NDUFB4 | 22596 7541 | 9397 | -0.019 | -0.3579 | No | ||
| 17 | NDUFA4 | 9451 | 9985 | -0.028 | -0.3892 | No | ||
| 18 | ATP6V1C2 | 21098 | 10801 | -0.042 | -0.4325 | No | ||
| 19 | ATP6V0A4 | 8934 | 12390 | -0.088 | -0.5168 | No | ||
| 20 | ATP7B | 18918 4426 4427 8641 | 12393 | -0.089 | -0.5157 | No | ||
| 21 | ATP6V1E1 | 4423 | 12449 | -0.091 | -0.5175 | No | ||
| 22 | ATP6V1G1 | 12322 | 12693 | -0.101 | -0.5292 | No | ||
| 23 | SDHB | 2348 12566 14880 | 13036 | -0.121 | -0.5460 | No | ||
| 24 | ATP6V0A1 | 8640 4424 1197 | 13141 | -0.128 | -0.5499 | No | ||
| 25 | NDUFA1 | 24170 | 13888 | -0.194 | -0.5874 | No | ||
| 26 | ATP6V1G2 | 23257 1534 | 13949 | -0.200 | -0.5880 | No | ||
| 27 | COX4I1 | 18444 | 15393 | -0.584 | -0.6578 | Yes | ||
| 28 | ATP6AP1 | 24296 | 15535 | -0.670 | -0.6563 | Yes | ||
| 29 | ATP6V0D1 | 3895 8639 3920 | 15720 | -0.803 | -0.6553 | Yes | ||
| 30 | ATP6V1A | 8638 | 15844 | -0.888 | -0.6499 | Yes | ||
| 31 | NDUFB7 | 12429 | 15927 | -0.961 | -0.6413 | Yes | ||
| 32 | UQCRC1 | 10259 | 16148 | -1.088 | -0.6384 | Yes | ||
| 33 | ATP6V1C1 | 22487 12329 | 16220 | -1.155 | -0.6266 | Yes | ||
| 34 | COX5B | 4552 | 16259 | -1.193 | -0.6125 | Yes | ||
| 35 | NDUFA11 | 12832 | 16407 | -1.341 | -0.6023 | Yes | ||
| 36 | NDUFB5 | 12277 | 16443 | -1.381 | -0.5855 | Yes | ||
| 37 | NDUFB2 | 7542 | 16478 | -1.433 | -0.5679 | Yes | ||
| 38 | ATP6V1F | 12291 | 16625 | -1.572 | -0.5545 | Yes | ||
| 39 | NDUFA5 | 1076 7544 | 16708 | -1.641 | -0.5367 | Yes | ||
| 40 | COX7C | 8776 | 16715 | -1.648 | -0.5147 | Yes | ||
| 41 | COX6A1 | 8774 4553 | 16863 | -1.830 | -0.4978 | Yes | ||
| 42 | COX5A | 19431 | 16953 | -1.962 | -0.4761 | Yes | ||
| 43 | ATP5O | 22539 | 17106 | -2.180 | -0.4547 | Yes | ||
| 44 | COX6B1 | 4283 8479 | 17117 | -2.194 | -0.4255 | Yes | ||
| 45 | ATP5E | 14321 | 17201 | -2.299 | -0.3989 | Yes | ||
| 46 | COX6A2 | 17605 | 17290 | -2.499 | -0.3698 | Yes | ||
| 47 | COX6C | 8775 | 17413 | -2.723 | -0.3395 | Yes | ||
| 48 | NDUFS2 | 4089 13760 | 17608 | -3.085 | -0.3081 | Yes | ||
| 49 | NDUFA8 | 14605 | 17673 | -3.259 | -0.2675 | Yes | ||
| 50 | ATP6V0B | 15786 | 17714 | -3.351 | -0.2242 | Yes | ||
| 51 | COX7B | 7260 | 17731 | -3.397 | -0.1791 | Yes | ||
| 52 | NDUFA10 | 7420 | 17735 | -3.403 | -0.1332 | Yes | ||
| 53 | COX10 | 20408 | 17939 | -4.003 | -0.0899 | Yes | ||
| 54 | NDUFV1 | 23953 | 18063 | -4.450 | -0.0362 | Yes | ||
| 55 | COX7A2 | 19053 | 18175 | -4.873 | 0.0238 | Yes |