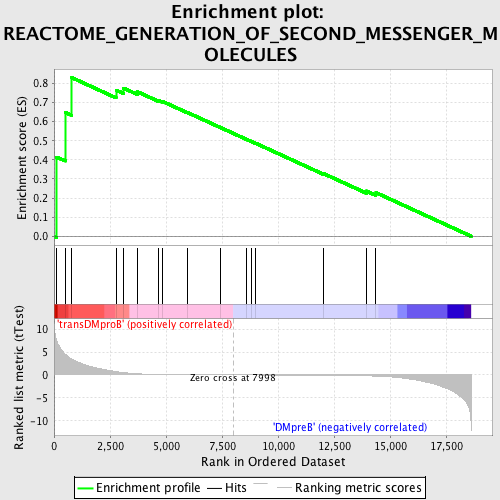
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | REACTOME_GENERATION_OF_SECOND_MESSENGER_MOLECULES |
| Enrichment Score (ES) | 0.8309101 |
| Normalized Enrichment Score (NES) | 1.5411503 |
| Nominal p-value | 0.015250545 |
| FDR q-value | 0.23449741 |
| FWER p-Value | 0.973 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LAT | 17643 | 116 | 7.434 | 0.4146 | Yes | ||
| 2 | WAS | 24193 | 499 | 4.495 | 0.6486 | Yes | ||
| 3 | LCK | 15746 | 786 | 3.491 | 0.8309 | Yes | ||
| 4 | PAK1 | 9527 | 2767 | 0.720 | 0.7653 | No | ||
| 5 | CD3D | 19473 | 3103 | 0.500 | 0.7756 | No | ||
| 6 | FYB | 10717 | 3715 | 0.252 | 0.7570 | No | ||
| 7 | ZAP70 | 14271 4042 | 4655 | 0.091 | 0.7116 | No | ||
| 8 | CD3G | 19139 | 4851 | 0.076 | 0.7055 | No | ||
| 9 | GRAP2 | 5113 9398 | 5961 | 0.034 | 0.6477 | No | ||
| 10 | PLCG1 | 14753 | 7414 | 0.008 | 0.5701 | No | ||
| 11 | ITK | 4934 | 8570 | -0.008 | 0.5084 | No | ||
| 12 | CD3E | 8714 | 8819 | -0.011 | 0.4957 | No | ||
| 13 | NCK1 | 9447 5152 | 8993 | -0.014 | 0.4872 | No | ||
| 14 | CD4 | 16999 | 12030 | -0.073 | 0.3281 | No | ||
| 15 | ENAH | 4665 8901 | 13933 | -0.198 | 0.2371 | No | ||
| 16 | LCP2 | 4988 9268 | 14367 | -0.258 | 0.2284 | No |

