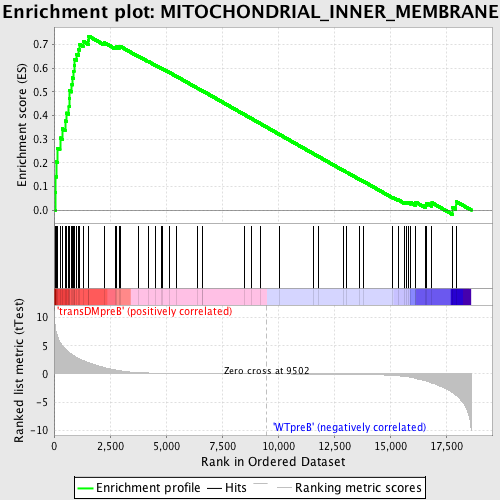
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | transDMpreB |
| GeneSet | MITOCHONDRIAL_INNER_MEMBRANE |
| Enrichment Score (ES) | 0.7359177 |
| Normalized Enrichment Score (NES) | 1.6662059 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.01676155 |
| FWER p-Value | 0.199 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TIMM13 | 14974 | 46 | 8.588 | 0.0740 | Yes | ||
| 2 | TIMM10 | 11390 2158 | 80 | 7.684 | 0.1407 | Yes | ||
| 3 | ATP5G1 | 8636 | 101 | 7.285 | 0.2045 | Yes | ||
| 4 | TIMM17A | 13822 | 154 | 6.602 | 0.2605 | Yes | ||
| 5 | SLC25A11 | 20366 1368 | 263 | 5.654 | 0.3050 | Yes | ||
| 6 | PHB2 | 17286 | 367 | 5.011 | 0.3441 | Yes | ||
| 7 | BCS1L | 14223 | 494 | 4.470 | 0.3771 | Yes | ||
| 8 | TIMM50 | 17911 | 562 | 4.171 | 0.4106 | Yes | ||
| 9 | NDUFS3 | 7553 12683 | 663 | 3.847 | 0.4395 | Yes | ||
| 10 | ATP5G2 | 12610 | 681 | 3.781 | 0.4723 | Yes | ||
| 11 | UQCRB | 12545 | 697 | 3.715 | 0.5045 | Yes | ||
| 12 | SLC25A22 | 12671 | 792 | 3.455 | 0.5302 | Yes | ||
| 13 | ATP5A1 | 23505 | 815 | 3.395 | 0.5593 | Yes | ||
| 14 | SLC25A15 | 356 3819 | 863 | 3.241 | 0.5856 | Yes | ||
| 15 | TIMM8B | 11391 | 890 | 3.180 | 0.6125 | Yes | ||
| 16 | ATP5G3 | 14558 | 918 | 3.114 | 0.6388 | Yes | ||
| 17 | NDUFA9 | 12288 | 1004 | 2.896 | 0.6600 | Yes | ||
| 18 | NDUFS7 | 19946 | 1095 | 2.728 | 0.6795 | Yes | ||
| 19 | ATP5O | 22539 | 1145 | 2.647 | 0.7004 | Yes | ||
| 20 | SLC25A3 | 19644 | 1296 | 2.390 | 0.7136 | Yes | ||
| 21 | NDUFV1 | 23953 | 1528 | 2.008 | 0.7191 | Yes | ||
| 22 | SDHD | 19125 | 1545 | 1.990 | 0.7359 | Yes | ||
| 23 | SLC25A1 | 22645 | 2243 | 1.101 | 0.7082 | No | ||
| 24 | NDUFAB1 | 7667 | 2732 | 0.718 | 0.6883 | No | ||
| 25 | SURF1 | 14645 | 2799 | 0.660 | 0.6906 | No | ||
| 26 | ATP5D | 19949 | 2903 | 0.583 | 0.6902 | No | ||
| 27 | TIMM9 | 19686 | 2960 | 0.553 | 0.6921 | No | ||
| 28 | ATP5E | 14321 | 3762 | 0.235 | 0.6511 | No | ||
| 29 | MPV17 | 9407 3505 3543 | 4216 | 0.162 | 0.6281 | No | ||
| 30 | ATP5J | 880 22555 | 4543 | 0.127 | 0.6117 | No | ||
| 31 | NDUFA2 | 9450 | 4786 | 0.110 | 0.5996 | No | ||
| 32 | HSD3B2 | 4873 4874 15229 4875 9126 | 4849 | 0.106 | 0.5972 | No | ||
| 33 | TIMM23 | 7004 5768 | 5150 | 0.088 | 0.5818 | No | ||
| 34 | COX15 | 5931 | 5462 | 0.074 | 0.5657 | No | ||
| 35 | NDUFS1 | 13940 | 6414 | 0.046 | 0.5149 | No | ||
| 36 | HCCS | 9079 4842 | 6628 | 0.042 | 0.5038 | No | ||
| 37 | UCP3 | 3972 18175 | 6633 | 0.042 | 0.5039 | No | ||
| 38 | UQCRC1 | 10259 | 8509 | 0.013 | 0.4030 | No | ||
| 39 | IMMT | 367 8029 | 8809 | 0.009 | 0.3870 | No | ||
| 40 | SDHA | 12436 | 9232 | 0.004 | 0.3643 | No | ||
| 41 | OPA1 | 13169 22800 7894 | 10064 | -0.007 | 0.3196 | No | ||
| 42 | PPOX | 13761 4104 | 11559 | -0.028 | 0.2393 | No | ||
| 43 | PHB | 9558 | 11782 | -0.032 | 0.2276 | No | ||
| 44 | HSD3B1 | 15227 | 12908 | -0.058 | 0.1675 | No | ||
| 45 | NDUFA1 | 24170 | 13060 | -0.063 | 0.1599 | No | ||
| 46 | ATP5B | 19846 | 13631 | -0.088 | 0.1300 | No | ||
| 47 | COX6B2 | 17983 4110 | 13818 | -0.099 | 0.1209 | No | ||
| 48 | NDUFS2 | 4089 13760 | 15119 | -0.261 | 0.0531 | No | ||
| 49 | NNT | 3273 5181 9471 3238 | 15351 | -0.322 | 0.0436 | No | ||
| 50 | MRPL32 | 7961 | 15632 | -0.435 | 0.0323 | No | ||
| 51 | NDUFA13 | 7400 | 15718 | -0.474 | 0.0320 | No | ||
| 52 | NDUFS4 | 9452 5157 3173 | 15820 | -0.525 | 0.0312 | No | ||
| 53 | ALAS2 | 24229 | 15930 | -0.601 | 0.0307 | No | ||
| 54 | ATP5C1 | 8635 | 16131 | -0.780 | 0.0269 | No | ||
| 55 | PMPCA | 12421 7350 | 16138 | -0.790 | 0.0336 | No | ||
| 56 | UQCRH | 12378 | 16588 | -1.209 | 0.0201 | No | ||
| 57 | NDUFS8 | 23950 | 16626 | -1.261 | 0.0294 | No | ||
| 58 | NDUFA6 | 12473 | 16866 | -1.614 | 0.0309 | No | ||
| 59 | ABCB7 | 24085 | 17789 | -3.335 | 0.0109 | No | ||
| 60 | TIMM17B | 24390 | 17940 | -3.775 | 0.0364 | No |

