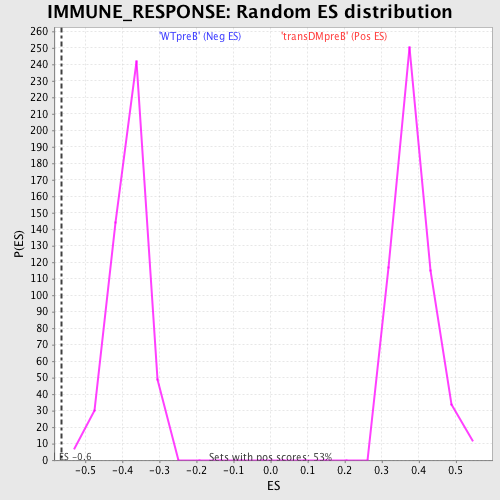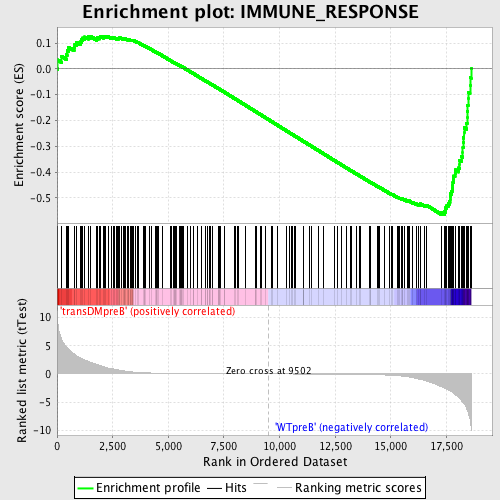
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | IMMUNE_RESPONSE |
| Enrichment Score (ES) | -0.5647526 |
| Normalized Enrichment Score (NES) | -1.4807731 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.6871146 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IKBKAP | 2540 6045 | 22 | 9.493 | 0.0348 | No | ||
| 2 | TNFAIP1 | 20333 | 196 | 6.200 | 0.0490 | No | ||
| 3 | CTSC | 18195 2261 | 425 | 4.779 | 0.0547 | No | ||
| 4 | CEBPB | 8733 | 449 | 4.666 | 0.0712 | No | ||
| 5 | BST2 | 18845 | 521 | 4.321 | 0.0837 | No | ||
| 6 | C1QBP | 20364 | 763 | 3.544 | 0.0841 | No | ||
| 7 | CD86 | 1660 22601 | 798 | 3.433 | 0.0953 | No | ||
| 8 | MAP3K7 | 16255 | 878 | 3.198 | 0.1031 | No | ||
| 9 | CD274 | 12244 | 1046 | 2.806 | 0.1047 | No | ||
| 10 | LY86 | 21665 | 1089 | 2.738 | 0.1128 | No | ||
| 11 | MALT1 | 6274 | 1143 | 2.647 | 0.1200 | No | ||
| 12 | UBE2N | 8216 19898 | 1244 | 2.470 | 0.1239 | No | ||
| 13 | APOA1 | 8614 | 1393 | 2.248 | 0.1244 | No | ||
| 14 | CHUK | 23665 | 1487 | 2.079 | 0.1272 | No | ||
| 15 | APOA2 | 8615 4044 | 1783 | 1.660 | 0.1175 | No | ||
| 16 | IL10 | 14145 1510 1553 22902 | 1820 | 1.629 | 0.1217 | No | ||
| 17 | EBI2 | 21718 | 1926 | 1.507 | 0.1218 | No | ||
| 18 | MAP4K2 | 6457 | 1945 | 1.471 | 0.1264 | No | ||
| 19 | IFITM3 | 17566 | 2071 | 1.317 | 0.1246 | No | ||
| 20 | LTF | 9289 | 2148 | 1.213 | 0.1251 | No | ||
| 21 | GPR65 | 21187 | 2196 | 1.158 | 0.1269 | No | ||
| 22 | TARBP2 | 10033 10034 5626 | 2300 | 1.037 | 0.1252 | No | ||
| 23 | YTHDF2 | 5623 5624 | 2459 | 0.931 | 0.1202 | No | ||
| 24 | TLR7 | 24004 | 2518 | 0.887 | 0.1204 | No | ||
| 25 | LTB4R | 22003 | 2578 | 0.841 | 0.1204 | No | ||
| 26 | BLNK | 23681 3691 | 2670 | 0.771 | 0.1184 | No | ||
| 27 | AQP9 | 7218 | 2720 | 0.728 | 0.1185 | No | ||
| 28 | THY1 | 19481 | 2772 | 0.683 | 0.1183 | No | ||
| 29 | HRH2 | 9120 | 2810 | 0.653 | 0.1188 | No | ||
| 30 | RSAD2 | 21094 | 2816 | 0.648 | 0.1210 | No | ||
| 31 | CCBP2 | 19262 | 2888 | 0.598 | 0.1194 | No | ||
| 32 | FCGR3A | 8960 | 2982 | 0.537 | 0.1164 | No | ||
| 33 | EBI3 | 23193 | 3029 | 0.502 | 0.1158 | No | ||
| 34 | IFITM2 | 13483 | 3040 | 0.496 | 0.1171 | No | ||
| 35 | CCL5 | 20320 | 3082 | 0.473 | 0.1167 | No | ||
| 36 | IL18BP | 17730 | 3170 | 0.424 | 0.1136 | No | ||
| 37 | IL6ST | 4920 | 3188 | 0.416 | 0.1142 | No | ||
| 38 | IL6R | 1862 4919 | 3230 | 0.397 | 0.1135 | No | ||
| 39 | TNFSF13 | 12806 | 3304 | 0.371 | 0.1109 | No | ||
| 40 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 3317 | 0.365 | 0.1117 | No | ||
| 41 | TGFB2 | 5743 | 3353 | 0.350 | 0.1111 | No | ||
| 42 | TCF7 | 1467 20466 | 3394 | 0.332 | 0.1102 | No | ||
| 43 | CCL27 | 5419 | 3395 | 0.331 | 0.1115 | No | ||
| 44 | ETS1 | 10715 6230 3135 | 3419 | 0.324 | 0.1114 | No | ||
| 45 | FOXP3 | 9804 5426 | 3523 | 0.292 | 0.1070 | No | ||
| 46 | DMBT1 | 18050 | 3609 | 0.270 | 0.1034 | No | ||
| 47 | POU2F2 | 17929 | 3654 | 0.260 | 0.1020 | No | ||
| 48 | CIITA | 22859 | 3874 | 0.215 | 0.0909 | No | ||
| 49 | IL4 | 9174 | 3912 | 0.208 | 0.0897 | No | ||
| 50 | MBL2 | 23886 | 3920 | 0.207 | 0.0901 | No | ||
| 51 | NCF4 | 22435 | 3992 | 0.194 | 0.0870 | No | ||
| 52 | IL15 | 18826 3801 | 4142 | 0.172 | 0.0795 | No | ||
| 53 | BCAR1 | 18741 | 4251 | 0.158 | 0.0743 | No | ||
| 54 | IL17A | 14289 | 4436 | 0.138 | 0.0648 | No | ||
| 55 | FYB | 10717 | 4452 | 0.137 | 0.0645 | No | ||
| 56 | CST7 | 14805 | 4484 | 0.133 | 0.0633 | No | ||
| 57 | GPR44 | 23926 | 4533 | 0.129 | 0.0612 | No | ||
| 58 | GTPBP1 | 4814 2255 | 4540 | 0.128 | 0.0614 | No | ||
| 59 | SP2 | 8130 | 4756 | 0.112 | 0.0501 | No | ||
| 60 | PTGER4 | 9649 | 5103 | 0.090 | 0.0317 | No | ||
| 61 | SFTPD | 21867 | 5147 | 0.088 | 0.0297 | No | ||
| 62 | LY75 | 9313 5026 | 5243 | 0.084 | 0.0248 | No | ||
| 63 | CCL4 | 1362 20730 | 5258 | 0.083 | 0.0244 | No | ||
| 64 | SEMA7A | 19426 9803 | 5281 | 0.082 | 0.0235 | No | ||
| 65 | IKBKG | 2570 2562 4908 | 5302 | 0.081 | 0.0227 | No | ||
| 66 | MBP | 23502 | 5324 | 0.079 | 0.0219 | No | ||
| 67 | TREM1 | 23205 | 5371 | 0.077 | 0.0197 | No | ||
| 68 | MS4A2 | 23729 | 5490 | 0.073 | 0.0135 | No | ||
| 69 | GBP2 | 9006 | 5532 | 0.071 | 0.0116 | No | ||
| 70 | BNIP3 | 17581 | 5548 | 0.070 | 0.0110 | No | ||
| 71 | CD28 | 14239 4092 | 5583 | 0.069 | 0.0095 | No | ||
| 72 | C2 | 23013 | 5605 | 0.068 | 0.0086 | No | ||
| 73 | HAMP | 17878 | 5623 | 0.067 | 0.0079 | No | ||
| 74 | CX3CL1 | 9791 | 5692 | 0.065 | 0.0045 | No | ||
| 75 | ANXA11 | 22075 | 5851 | 0.060 | -0.0039 | No | ||
| 76 | CRTAM | 19160 | 6000 | 0.056 | -0.0117 | No | ||
| 77 | IL6 | 16895 | 6126 | 0.053 | -0.0183 | No | ||
| 78 | CCL22 | 18518 | 6137 | 0.053 | -0.0186 | No | ||
| 79 | ZAP70 | 14271 4042 | 6321 | 0.048 | -0.0284 | No | ||
| 80 | CCL2 | 9788 | 6471 | 0.045 | -0.0363 | No | ||
| 81 | TLR8 | 9308 | 6497 | 0.044 | -0.0375 | No | ||
| 82 | EREG | 4679 16797 | 6648 | 0.041 | -0.0455 | No | ||
| 83 | CTSW | 23980 | 6662 | 0.041 | -0.0460 | No | ||
| 84 | SPINK5 | 23569 | 6759 | 0.039 | -0.0511 | No | ||
| 85 | IL27 | 17636 | 6764 | 0.039 | -0.0512 | No | ||
| 86 | SEMA3C | 16914 | 6840 | 0.038 | -0.0551 | No | ||
| 87 | COLEC12 | 23624 8946 | 6859 | 0.037 | -0.0559 | No | ||
| 88 | CXCL12 | 9792 1182 1031 | 6880 | 0.037 | -0.0569 | No | ||
| 89 | DEFB1 | 18663 | 6980 | 0.035 | -0.0621 | No | ||
| 90 | CD7 | 20105 | 6991 | 0.035 | -0.0625 | No | ||
| 91 | CCR2 | 19250 | 7241 | 0.030 | -0.0759 | No | ||
| 92 | CCRL1 | 10909 3077 | 7246 | 0.030 | -0.0760 | No | ||
| 93 | IFNK | 11837 | 7302 | 0.030 | -0.0789 | No | ||
| 94 | CADM1 | 7057 | 7304 | 0.030 | -0.0789 | No | ||
| 95 | NCR1 | 18409 | 7362 | 0.028 | -0.0818 | No | ||
| 96 | SECTM1 | 20104 | 7533 | 0.026 | -0.0910 | No | ||
| 97 | DEFB103A | 18665 | 7537 | 0.026 | -0.0910 | No | ||
| 98 | MADCAM1 | 19966 | 7982 | 0.020 | -0.1151 | No | ||
| 99 | IL1R2 | 14265 | 8032 | 0.019 | -0.1177 | No | ||
| 100 | CD40LG | 24330 | 8127 | 0.018 | -0.1227 | No | ||
| 101 | CCR5 | 8754 4534 | 8140 | 0.018 | -0.1233 | No | ||
| 102 | CHST4 | 18749 | 8469 | 0.013 | -0.1410 | No | ||
| 103 | IL12B | 20918 | 8911 | 0.007 | -0.1649 | No | ||
| 104 | CCL24 | 16342 | 8922 | 0.007 | -0.1655 | No | ||
| 105 | TNFRSF4 | 15954 | 8960 | 0.007 | -0.1674 | No | ||
| 106 | RGS1 | 3970 6936 | 9151 | 0.005 | -0.1777 | No | ||
| 107 | LAT | 17643 | 9183 | 0.004 | -0.1794 | No | ||
| 108 | CCR1 | 18951 | 9345 | 0.002 | -0.1881 | No | ||
| 109 | CD96 | 22580 | 9628 | -0.002 | -0.2034 | No | ||
| 110 | ODZ1 | 24168 | 9659 | -0.002 | -0.2051 | No | ||
| 111 | CCR8 | 19272 | 9904 | -0.005 | -0.2183 | No | ||
| 112 | IL17B | 12074 | 10294 | -0.010 | -0.2394 | No | ||
| 113 | TREM2 | 23201 | 10443 | -0.012 | -0.2474 | No | ||
| 114 | DEFB4 | 18662 18668 | 10537 | -0.013 | -0.2524 | No | ||
| 115 | OPRD1 | 15737 | 10577 | -0.014 | -0.2544 | No | ||
| 116 | APLN | 24164 | 10655 | -0.015 | -0.2585 | No | ||
| 117 | GZMA | 21343 | 10727 | -0.016 | -0.2623 | No | ||
| 118 | GEM | 16274 | 11061 | -0.021 | -0.2803 | No | ||
| 119 | CCR9 | 8753 | 11073 | -0.021 | -0.2809 | No | ||
| 120 | C5AR1 | 17957 | 11086 | -0.021 | -0.2814 | No | ||
| 121 | APOA4 | 4401 | 11350 | -0.025 | -0.2956 | No | ||
| 122 | CCL19 | 10773 | 11417 | -0.026 | -0.2991 | No | ||
| 123 | TRAF2 | 14657 | 11751 | -0.032 | -0.3171 | No | ||
| 124 | IL7 | 4921 | 11984 | -0.036 | -0.3295 | No | ||
| 125 | CMKLR1 | 16429 | 12486 | -0.047 | -0.3565 | No | ||
| 126 | IL2 | 15354 | 12608 | -0.050 | -0.3629 | No | ||
| 127 | CRHR1 | 1464 1468 20633 | 12768 | -0.054 | -0.3713 | No | ||
| 128 | CFHR1 | 11926 13815 | 13017 | -0.062 | -0.3846 | No | ||
| 129 | VTN | 20756 | 13186 | -0.068 | -0.3934 | No | ||
| 130 | IGSF6 | 17653 | 13239 | -0.070 | -0.3960 | No | ||
| 131 | CTLA4 | 14238 | 13463 | -0.078 | -0.4078 | No | ||
| 132 | CXCL13 | 16789 | 13595 | -0.086 | -0.4146 | No | ||
| 133 | RFX1 | 18548 | 13642 | -0.088 | -0.4167 | No | ||
| 134 | SOCS5 | 1619 23141 | 14043 | -0.114 | -0.4380 | No | ||
| 135 | VIPR1 | 3048 19264 3012 | 14084 | -0.117 | -0.4398 | No | ||
| 136 | LCP2 | 4988 9268 | 14104 | -0.118 | -0.4403 | No | ||
| 137 | IL28RA | 10823 16038 | 14393 | -0.144 | -0.4554 | No | ||
| 138 | CCR4 | 18978 | 14458 | -0.152 | -0.4583 | No | ||
| 139 | PDCD1 | 13870 | 14508 | -0.157 | -0.4604 | No | ||
| 140 | CTSG | 21815 | 14729 | -0.188 | -0.4716 | No | ||
| 141 | IL2RG | 24096 | 14951 | -0.226 | -0.4828 | No | ||
| 142 | SEMA4D | 9800 | 15030 | -0.243 | -0.4861 | No | ||
| 143 | FYN | 3375 3395 20052 | 15076 | -0.254 | -0.4876 | No | ||
| 144 | CCR6 | 1505 23390 | 15311 | -0.312 | -0.4991 | No | ||
| 145 | CCL25 | 9789 3848 | 15338 | -0.319 | -0.4993 | No | ||
| 146 | OPRK1 | 14300 | 15391 | -0.339 | -0.5008 | No | ||
| 147 | FCGRT | 17839 | 15491 | -0.377 | -0.5048 | No | ||
| 148 | PTPRC | 5327 9662 | 15502 | -0.380 | -0.5039 | No | ||
| 149 | PAX5 | 15892 | 15518 | -0.387 | -0.5032 | No | ||
| 150 | DEFB127 | 622 | 15633 | -0.435 | -0.5077 | No | ||
| 151 | IL27RA | 11949 | 15768 | -0.497 | -0.5131 | No | ||
| 152 | IK | 23594 | 15794 | -0.514 | -0.5125 | No | ||
| 153 | MR1 | 4837 | 15818 | -0.524 | -0.5118 | No | ||
| 154 | CNIH | 4536 23971 | 15994 | -0.658 | -0.5188 | No | ||
| 155 | ARHGDIB | 16950 | 16142 | -0.803 | -0.5237 | No | ||
| 156 | TAPBP | 5625 | 16232 | -0.893 | -0.5252 | No | ||
| 157 | CD22 | 17881 | 16316 | -0.978 | -0.5260 | No | ||
| 158 | DPP8 | 19408 | 16324 | -0.985 | -0.5226 | No | ||
| 159 | CTSS | 15505 | 16505 | -1.130 | -0.5281 | No | ||
| 160 | NFIL3 | 21463 | 16622 | -1.258 | -0.5296 | No | ||
| 161 | IL4R | 18085 | 17270 | -2.250 | -0.5562 | Yes | ||
| 162 | IL10RB | 9169 | 17406 | -2.493 | -0.5541 | Yes | ||
| 163 | PSMB10 | 5299 18761 | 17450 | -2.545 | -0.5468 | Yes | ||
| 164 | TNFRSF14 | 15650 | 17472 | -2.579 | -0.5381 | Yes | ||
| 165 | BST1 | 16850 | 17520 | -2.674 | -0.5305 | Yes | ||
| 166 | POU2AF1 | 19455 | 17577 | -2.801 | -0.5229 | Yes | ||
| 167 | FCGR2B | 8959 | 17622 | -2.880 | -0.5144 | Yes | ||
| 168 | CD164 | 20046 | 17660 | -2.966 | -0.5051 | Yes | ||
| 169 | FTH1 | 8986 | 17665 | -2.977 | -0.4941 | Yes | ||
| 170 | WAS | 24193 | 17688 | -3.040 | -0.4837 | Yes | ||
| 171 | LAT2 | 16351 | 17736 | -3.151 | -0.4743 | Yes | ||
| 172 | CD79B | 20185 1309 | 17759 | -3.226 | -0.4633 | Yes | ||
| 173 | CD79A | 18342 | 17781 | -3.312 | -0.4519 | Yes | ||
| 174 | TGFB1 | 18332 | 17790 | -3.337 | -0.4396 | Yes | ||
| 175 | ATP6V0A2 | 5760 | 17810 | -3.402 | -0.4277 | Yes | ||
| 176 | ZEB1 | 5647 | 17824 | -3.443 | -0.4154 | Yes | ||
| 177 | EDG6 | 19672 | 17886 | -3.639 | -0.4049 | Yes | ||
| 178 | PRKRA | 357 | 17904 | -3.689 | -0.3918 | Yes | ||
| 179 | IL2RA | 4918 | 18055 | -4.139 | -0.3843 | Yes | ||
| 180 | DPP4 | 14579 | 18084 | -4.277 | -0.3695 | Yes | ||
| 181 | IL16 | 9171 | 18088 | -4.289 | -0.3534 | Yes | ||
| 182 | BCL10 | 15397 | 18173 | -4.717 | -0.3401 | Yes | ||
| 183 | CD83 | 21653 | 18239 | -5.076 | -0.3244 | Yes | ||
| 184 | IL18 | 9172 | 18242 | -5.084 | -0.3052 | Yes | ||
| 185 | TCF12 | 10044 3125 | 18254 | -5.152 | -0.2862 | Yes | ||
| 186 | MS4A1 | 59 | 18263 | -5.193 | -0.2670 | Yes | ||
| 187 | TRAF6 | 5797 14940 | 18289 | -5.308 | -0.2482 | Yes | ||
| 188 | IL12A | 4913 | 18294 | -5.328 | -0.2282 | Yes | ||
| 189 | IRF8 | 18443 | 18397 | -5.983 | -0.2110 | Yes | ||
| 190 | BNIP3L | 21775 | 18430 | -6.311 | -0.1888 | Yes | ||
| 191 | ST6GAL1 | 22806 | 18435 | -6.343 | -0.1650 | Yes | ||
| 192 | CNR2 | 16035 | 18454 | -6.573 | -0.1410 | Yes | ||
| 193 | RAG1 | 9689 | 18474 | -6.799 | -0.1162 | Yes | ||
| 194 | XBP1 | 10318 | 18480 | -6.901 | -0.0903 | Yes | ||
| 195 | IL7R | 9175 4922 | 18559 | -8.025 | -0.0641 | Yes | ||
| 196 | CXCR4 | 13844 | 18566 | -8.219 | -0.0332 | Yes | ||
| 197 | CD97 | 18822 3874 | 18609 | -9.463 | 0.0004 | Yes |