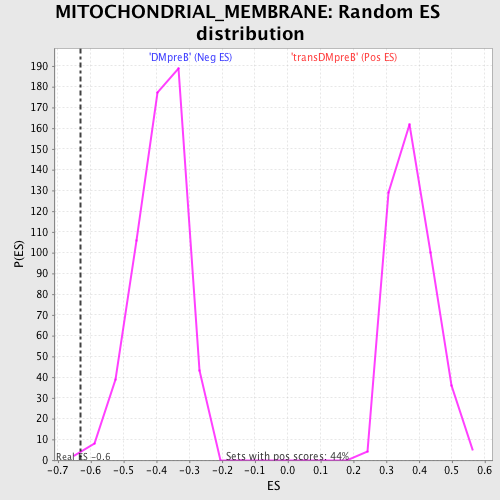
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | MITOCHONDRIAL_MEMBRANE |
| Enrichment Score (ES) | -0.6306198 |
| Normalized Enrichment Score (NES) | -1.6263514 |
| Nominal p-value | 0.0035460992 |
| FDR q-value | 0.1008613 |
| FWER p-Value | 0.729 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC25A22 | 12671 | 177 | 2.205 | 0.0333 | No | ||
| 2 | MCL1 | 15502 | 449 | 1.697 | 0.0516 | No | ||
| 3 | RAF1 | 17035 | 1186 | 0.942 | 0.0302 | No | ||
| 4 | SLC25A3 | 19644 | 1696 | 0.631 | 0.0150 | No | ||
| 5 | ATP5A1 | 23505 | 1763 | 0.591 | 0.0229 | No | ||
| 6 | TIMM10 | 11390 2158 | 1828 | 0.550 | 0.0301 | No | ||
| 7 | HSD3B2 | 4873 4874 15229 4875 9126 | 2059 | 0.437 | 0.0262 | No | ||
| 8 | RHOT2 | 1533 10073 | 2159 | 0.395 | 0.0285 | No | ||
| 9 | CASP7 | 23826 | 2325 | 0.347 | 0.0263 | No | ||
| 10 | TOMM34 | 14359 | 2482 | 0.298 | 0.0237 | No | ||
| 11 | COX6B2 | 17983 4110 | 2694 | 0.251 | 0.0172 | No | ||
| 12 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 2820 | 0.230 | 0.0149 | No | ||
| 13 | OGDH | 9502 1412 | 3215 | 0.170 | -0.0030 | No | ||
| 14 | ATP5B | 19846 | 3318 | 0.157 | -0.0055 | No | ||
| 15 | MTX2 | 14980 | 3620 | 0.129 | -0.0192 | No | ||
| 16 | UQCRB | 12545 | 3654 | 0.125 | -0.0186 | No | ||
| 17 | ATP5D | 19949 | 3793 | 0.115 | -0.0238 | No | ||
| 18 | ALAS2 | 24229 | 4176 | 0.091 | -0.0426 | No | ||
| 19 | OPA1 | 13169 22800 7894 | 4512 | 0.075 | -0.0592 | No | ||
| 20 | UQCRC1 | 10259 | 4631 | 0.071 | -0.0642 | No | ||
| 21 | AIFM2 | 3320 3407 20006 3317 | 5010 | 0.058 | -0.0835 | No | ||
| 22 | SDHD | 19125 | 5278 | 0.051 | -0.0969 | No | ||
| 23 | HSD3B1 | 15227 | 5474 | 0.047 | -0.1065 | No | ||
| 24 | MFN2 | 15678 2417 | 5481 | 0.046 | -0.1059 | No | ||
| 25 | HCCS | 9079 4842 | 5617 | 0.043 | -0.1124 | No | ||
| 26 | TIMM23 | 7004 5768 | 6105 | 0.034 | -0.1380 | No | ||
| 27 | PHB | 9558 | 6674 | 0.025 | -0.1681 | No | ||
| 28 | NDUFS1 | 13940 | 6971 | 0.020 | -0.1837 | No | ||
| 29 | UCP3 | 3972 18175 | 8050 | 0.007 | -0.2417 | No | ||
| 30 | BNIP3 | 17581 | 8252 | 0.005 | -0.2524 | No | ||
| 31 | SDHA | 12436 | 8267 | 0.005 | -0.2531 | No | ||
| 32 | IMMT | 367 8029 | 10295 | -0.021 | -0.3620 | No | ||
| 33 | NDUFS7 | 19946 | 10999 | -0.031 | -0.3994 | No | ||
| 34 | MRPL32 | 7961 | 11213 | -0.034 | -0.4102 | No | ||
| 35 | SLC25A11 | 20366 1368 | 13183 | -0.085 | -0.5147 | No | ||
| 36 | NDUFA6 | 12473 | 13383 | -0.093 | -0.5237 | No | ||
| 37 | RAB11FIP5 | 17089 | 13735 | -0.111 | -0.5405 | No | ||
| 38 | NDUFA13 | 7400 | 13927 | -0.122 | -0.5484 | No | ||
| 39 | TIMM9 | 19686 | 14011 | -0.128 | -0.5504 | No | ||
| 40 | TIMM17B | 24390 | 14649 | -0.185 | -0.5811 | No | ||
| 41 | ATP5C1 | 8635 | 14751 | -0.196 | -0.5828 | No | ||
| 42 | PPOX | 13761 4104 | 14996 | -0.228 | -0.5915 | No | ||
| 43 | NDUFAB1 | 7667 | 15387 | -0.286 | -0.6070 | No | ||
| 44 | ATP5G3 | 14558 | 15683 | -0.351 | -0.6161 | No | ||
| 45 | VDAC1 | 10282 | 15778 | -0.373 | -0.6139 | No | ||
| 46 | NNT | 3273 5181 9471 3238 | 15898 | -0.402 | -0.6125 | No | ||
| 47 | ATP5E | 14321 | 15921 | -0.408 | -0.6058 | No | ||
| 48 | RHOT1 | 12227 | 16014 | -0.434 | -0.6023 | No | ||
| 49 | ATP5O | 22539 | 16259 | -0.501 | -0.6058 | No | ||
| 50 | NDUFS8 | 23950 | 16721 | -0.685 | -0.6173 | Yes | ||
| 51 | ATP5G2 | 12610 | 16897 | -0.770 | -0.6118 | Yes | ||
| 52 | TOMM22 | 5863 | 17038 | -0.852 | -0.6028 | Yes | ||
| 53 | NDUFA1 | 24170 | 17089 | -0.873 | -0.5886 | Yes | ||
| 54 | PMPCA | 12421 7350 | 17156 | -0.900 | -0.5747 | Yes | ||
| 55 | SLC25A1 | 22645 | 17207 | -0.925 | -0.5594 | Yes | ||
| 56 | PHB2 | 17286 | 17213 | -0.930 | -0.5416 | Yes | ||
| 57 | UQCRH | 12378 | 17279 | -0.965 | -0.5264 | Yes | ||
| 58 | ATP5J | 880 22555 | 17364 | -1.001 | -0.5115 | Yes | ||
| 59 | NDUFS2 | 4089 13760 | 17480 | -1.054 | -0.4972 | Yes | ||
| 60 | ATP5G1 | 8636 | 17491 | -1.058 | -0.4772 | Yes | ||
| 61 | VDAC3 | 10284 | 17504 | -1.066 | -0.4572 | Yes | ||
| 62 | SURF1 | 14645 | 17597 | -1.122 | -0.4404 | Yes | ||
| 63 | NDUFS3 | 7553 12683 | 17602 | -1.123 | -0.4188 | Yes | ||
| 64 | TIMM17A | 13822 | 17641 | -1.144 | -0.3986 | Yes | ||
| 65 | VDAC2 | 22081 | 17737 | -1.213 | -0.3802 | Yes | ||
| 66 | COX15 | 5931 | 17880 | -1.308 | -0.3625 | Yes | ||
| 67 | NDUFA2 | 9450 | 17911 | -1.329 | -0.3383 | Yes | ||
| 68 | NDUFS4 | 9452 5157 3173 | 17926 | -1.349 | -0.3129 | Yes | ||
| 69 | BCS1L | 14223 | 17987 | -1.397 | -0.2890 | Yes | ||
| 70 | NDUFV1 | 23953 | 18102 | -1.507 | -0.2659 | Yes | ||
| 71 | NDUFA9 | 12288 | 18165 | -1.581 | -0.2385 | Yes | ||
| 72 | TIMM50 | 17911 | 18187 | -1.603 | -0.2085 | Yes | ||
| 73 | TIMM8B | 11391 | 18239 | -1.678 | -0.1787 | Yes | ||
| 74 | FIS1 | 16669 3579 | 18259 | -1.713 | -0.1465 | Yes | ||
| 75 | ABCB7 | 24085 | 18323 | -1.823 | -0.1145 | Yes | ||
| 76 | TIMM13 | 14974 | 18359 | -1.891 | -0.0797 | Yes | ||
| 77 | MPV17 | 9407 3505 3543 | 18511 | -2.319 | -0.0428 | Yes | ||
| 78 | SLC25A15 | 356 3819 | 18548 | -2.494 | 0.0037 | Yes |