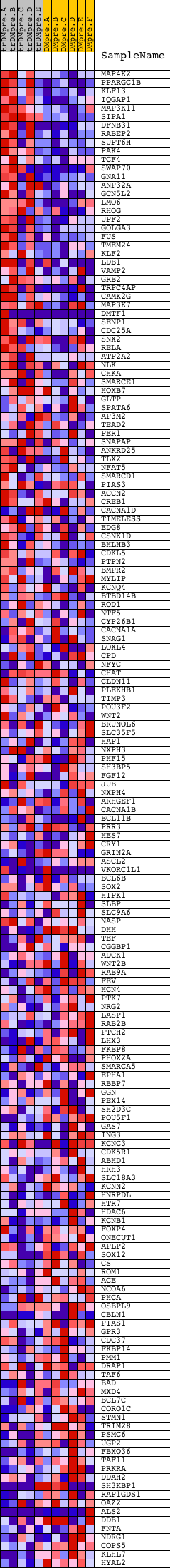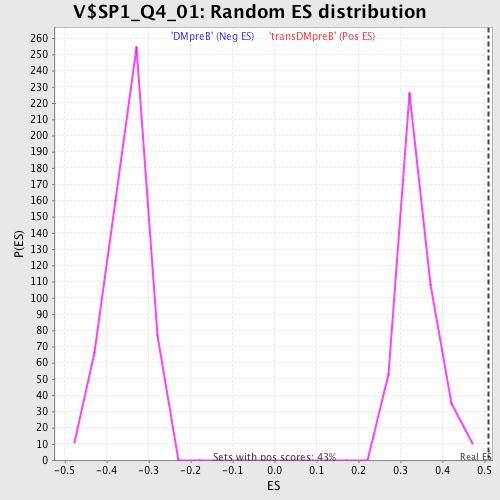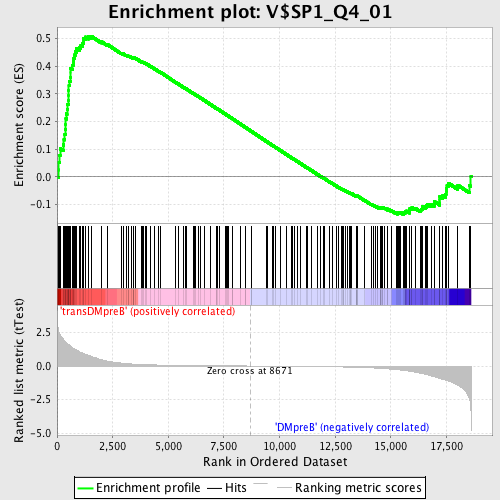
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | V$SP1_Q4_01 |
| Enrichment Score (ES) | 0.5094248 |
| Normalized Enrichment Score (NES) | 1.5052145 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.17076781 |
| FWER p-Value | 0.811 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MAP4K2 | 6457 | 72 | 2.603 | 0.0248 | Yes | ||
| 2 | PPARGC1B | 9318 | 77 | 2.580 | 0.0530 | Yes | ||
| 3 | KLF13 | 17807 | 119 | 2.372 | 0.0770 | Yes | ||
| 4 | IQGAP1 | 6619 | 136 | 2.288 | 0.1014 | Yes | ||
| 5 | MAP3K11 | 11163 | 275 | 2.001 | 0.1159 | Yes | ||
| 6 | SIPA1 | 9820 | 307 | 1.908 | 0.1353 | Yes | ||
| 7 | DFNB31 | 13139 | 347 | 1.835 | 0.1534 | Yes | ||
| 8 | RABEP2 | 18082 | 373 | 1.801 | 0.1719 | Yes | ||
| 9 | SUPT6H | 9940 | 389 | 1.782 | 0.1908 | Yes | ||
| 10 | PAK4 | 17909 | 396 | 1.777 | 0.2101 | Yes | ||
| 11 | TCF4 | 5642 | 417 | 1.735 | 0.2281 | Yes | ||
| 12 | SWAP70 | 18126 5553 9948 | 447 | 1.702 | 0.2453 | Yes | ||
| 13 | GNA11 | 3300 3345 19670 | 465 | 1.674 | 0.2629 | Yes | ||
| 14 | ANP32A | 8592 4387 19415 | 514 | 1.618 | 0.2781 | Yes | ||
| 15 | GCN5L2 | 4763 | 521 | 1.601 | 0.2954 | Yes | ||
| 16 | LMO6 | 24392 | 526 | 1.596 | 0.3128 | Yes | ||
| 17 | RHOG | 17727 | 533 | 1.576 | 0.3299 | Yes | ||
| 18 | UPF2 | 11591 | 550 | 1.558 | 0.3462 | Yes | ||
| 19 | GOLGA3 | 6535 | 590 | 1.508 | 0.3607 | Yes | ||
| 20 | FUS | 6135 | 599 | 1.491 | 0.3767 | Yes | ||
| 21 | TMEM24 | 19150 | 600 | 1.490 | 0.3932 | Yes | ||
| 22 | KLF2 | 9229 | 686 | 1.384 | 0.4038 | Yes | ||
| 23 | LDB1 | 23654 | 722 | 1.333 | 0.4166 | Yes | ||
| 24 | VAMP2 | 20826 | 750 | 1.302 | 0.4295 | Yes | ||
| 25 | GRB2 | 20149 | 792 | 1.266 | 0.4413 | Yes | ||
| 26 | TRPC4AP | 14379 12125 | 812 | 1.248 | 0.4540 | Yes | ||
| 27 | CAMK2G | 21905 | 866 | 1.191 | 0.4643 | Yes | ||
| 28 | MAP3K7 | 16255 | 1018 | 1.035 | 0.4675 | Yes | ||
| 29 | DMTF1 | 887 3662 16928 3530 | 1067 | 1.009 | 0.4760 | Yes | ||
| 30 | SENP1 | 10301 | 1136 | 0.983 | 0.4832 | Yes | ||
| 31 | CDC25A | 8721 | 1174 | 0.955 | 0.4917 | Yes | ||
| 32 | SNX2 | 1974 23554 12585 | 1183 | 0.943 | 0.5017 | Yes | ||
| 33 | RELA | 23783 | 1277 | 0.876 | 0.5063 | Yes | ||
| 34 | ATP2A2 | 4421 3481 | 1416 | 0.790 | 0.5075 | Yes | ||
| 35 | NLK | 5179 5178 | 1529 | 0.723 | 0.5094 | Yes | ||
| 36 | CHKA | 23759 | 1991 | 0.465 | 0.4896 | No | ||
| 37 | SMARCE1 | 1218 20253 | 2283 | 0.359 | 0.4777 | No | ||
| 38 | HOXB7 | 666 9110 | 2877 | 0.221 | 0.4480 | No | ||
| 39 | GLTP | 16419 | 2963 | 0.206 | 0.4457 | No | ||
| 40 | SPATA6 | 16134 | 3102 | 0.186 | 0.4402 | No | ||
| 41 | AP3M2 | 11 | 3220 | 0.169 | 0.4358 | No | ||
| 42 | TEAD2 | 18252 | 3224 | 0.168 | 0.4374 | No | ||
| 43 | PER1 | 20827 | 3332 | 0.155 | 0.4333 | No | ||
| 44 | SNAPAP | 15270 | 3411 | 0.147 | 0.4307 | No | ||
| 45 | ANKRD25 | 6161 | 3430 | 0.144 | 0.4313 | No | ||
| 46 | TLX2 | 5773 1165 | 3433 | 0.144 | 0.4328 | No | ||
| 47 | NFAT5 | 3921 7037 12036 | 3519 | 0.137 | 0.4297 | No | ||
| 48 | SMARCD1 | 2241 22360 2250 | 3772 | 0.116 | 0.4173 | No | ||
| 49 | PIAS3 | 15491 672 1906 | 3825 | 0.113 | 0.4158 | No | ||
| 50 | ACCN2 | 22363 | 3861 | 0.110 | 0.4151 | No | ||
| 51 | CREB1 | 3990 8782 4558 4093 | 3961 | 0.105 | 0.4109 | No | ||
| 52 | CACNA1D | 4464 8675 | 4027 | 0.100 | 0.4085 | No | ||
| 53 | TIMELESS | 10177 | 4208 | 0.089 | 0.3997 | No | ||
| 54 | EDG8 | 19212 | 4387 | 0.081 | 0.3909 | No | ||
| 55 | CSNK1D | 4181 8352 1387 | 4572 | 0.073 | 0.3817 | No | ||
| 56 | BHLHB3 | 16935 | 4634 | 0.071 | 0.3792 | No | ||
| 57 | CDKL5 | 24019 | 5330 | 0.050 | 0.3421 | No | ||
| 58 | PTPN2 | 23414 | 5435 | 0.047 | 0.3370 | No | ||
| 59 | BMPR2 | 14241 4081 | 5692 | 0.042 | 0.3235 | No | ||
| 60 | MYLIP | 21652 3189 | 5767 | 0.040 | 0.3200 | No | ||
| 61 | KCNQ4 | 12247 2533 | 5808 | 0.040 | 0.3182 | No | ||
| 62 | BTBD14B | 7344 7343 | 6113 | 0.034 | 0.3021 | No | ||
| 63 | ROD1 | 6047 2317 | 6181 | 0.033 | 0.2989 | No | ||
| 64 | NTF5 | 18248 | 6200 | 0.033 | 0.2982 | No | ||
| 65 | CYP26B1 | 17093 | 6349 | 0.030 | 0.2906 | No | ||
| 66 | CACNA1A | 4461 | 6455 | 0.028 | 0.2852 | No | ||
| 67 | SNAG1 | 21341 | 6644 | 0.025 | 0.2753 | No | ||
| 68 | LOXL4 | 23672 | 6916 | 0.021 | 0.2608 | No | ||
| 69 | CPD | 20344 4556 | 7162 | 0.018 | 0.2477 | No | ||
| 70 | NFYC | 9460 | 7164 | 0.018 | 0.2479 | No | ||
| 71 | CHAT | 21882 | 7211 | 0.017 | 0.2455 | No | ||
| 72 | CLDN11 | 15616 | 7276 | 0.017 | 0.2423 | No | ||
| 73 | PLEKHB1 | 21729 2755 4059 17738 | 7548 | 0.013 | 0.2277 | No | ||
| 74 | TIMP3 | 10179 5758 | 7604 | 0.012 | 0.2249 | No | ||
| 75 | POU3F2 | 15931 | 7636 | 0.012 | 0.2233 | No | ||
| 76 | WNT2 | 17212 | 7705 | 0.011 | 0.2198 | No | ||
| 77 | BRUNOL6 | 19421 | 7898 | 0.009 | 0.2094 | No | ||
| 78 | SLC35F5 | 13171 | 7900 | 0.009 | 0.2095 | No | ||
| 79 | HAP1 | 1297 20231 1231 | 8230 | 0.005 | 0.1917 | No | ||
| 80 | NXPH3 | 20281 | 8463 | 0.002 | 0.1791 | No | ||
| 81 | PHF15 | 20467 8050 | 8488 | 0.002 | 0.1779 | No | ||
| 82 | SH3BP5 | 6278 | 8721 | -0.001 | 0.1653 | No | ||
| 83 | FGF12 | 1723 22621 | 8735 | -0.001 | 0.1646 | No | ||
| 84 | JUB | 21834 3339 | 9413 | -0.009 | 0.1280 | No | ||
| 85 | NXPH4 | 19602 | 9438 | -0.010 | 0.1268 | No | ||
| 86 | ARHGEF1 | 1952 18343 3996 | 9677 | -0.013 | 0.1140 | No | ||
| 87 | CACNA1B | 8673 4462 | 9729 | -0.013 | 0.1114 | No | ||
| 88 | BCL11B | 7190 | 9826 | -0.015 | 0.1064 | No | ||
| 89 | PRR3 | 22995 | 10062 | -0.018 | 0.0938 | No | ||
| 90 | HES7 | 20828 | 10319 | -0.021 | 0.0802 | No | ||
| 91 | CRY1 | 19662 | 10517 | -0.024 | 0.0697 | No | ||
| 92 | GRIN2A | 22668 | 10572 | -0.025 | 0.0671 | No | ||
| 93 | ASCL2 | 5079 | 10681 | -0.026 | 0.0615 | No | ||
| 94 | VKORC1L1 | 102 7625 | 10824 | -0.028 | 0.0541 | No | ||
| 95 | BCL6B | 20373 | 10960 | -0.030 | 0.0472 | No | ||
| 96 | SOX2 | 9849 15612 | 11229 | -0.034 | 0.0330 | No | ||
| 97 | HIPK1 | 4851 | 11268 | -0.035 | 0.0313 | No | ||
| 98 | SLBP | 9822 | 11453 | -0.038 | 0.0218 | No | ||
| 99 | SLC9A6 | 2590 6189 | 11708 | -0.043 | 0.0085 | No | ||
| 100 | NASP | 2383 2387 6955 | 11839 | -0.046 | 0.0019 | No | ||
| 101 | DHH | 4629 | 11960 | -0.048 | -0.0041 | No | ||
| 102 | TEF | 2287 22414 2192 | 12004 | -0.049 | -0.0059 | No | ||
| 103 | CGGBP1 | 22724 | 12253 | -0.055 | -0.0187 | No | ||
| 104 | ADCK1 | 21191 | 12372 | -0.058 | -0.0245 | No | ||
| 105 | WNT2B | 15214 1838 | 12537 | -0.063 | -0.0327 | No | ||
| 106 | RAB9A | 12113 7107 | 12650 | -0.066 | -0.0380 | No | ||
| 107 | FEV | 11153 6434 | 12798 | -0.071 | -0.0452 | No | ||
| 108 | HCN4 | 9080 | 12825 | -0.072 | -0.0458 | No | ||
| 109 | PTK7 | 22959 | 12859 | -0.073 | -0.0468 | No | ||
| 110 | NRG2 | 4274 | 12885 | -0.074 | -0.0473 | No | ||
| 111 | LASP1 | 4986 4985 1248 | 12967 | -0.077 | -0.0509 | No | ||
| 112 | RAB2B | 21842 | 13034 | -0.079 | -0.0536 | No | ||
| 113 | PTCH2 | 9640 | 13134 | -0.083 | -0.0580 | No | ||
| 114 | LHX3 | 2761 4996 | 13180 | -0.085 | -0.0595 | No | ||
| 115 | FKBP8 | 18587 | 13201 | -0.085 | -0.0597 | No | ||
| 116 | PHOX2A | 18168 | 13251 | -0.087 | -0.0614 | No | ||
| 117 | SMARCA5 | 8215 | 13446 | -0.096 | -0.0708 | No | ||
| 118 | EPHA1 | 17168 8908 17167 | 13448 | -0.096 | -0.0698 | No | ||
| 119 | RBBP7 | 10887 6366 10886 | 13450 | -0.096 | -0.0688 | No | ||
| 120 | GGN | 2472 1171 18312 | 13481 | -0.098 | -0.0694 | No | ||
| 121 | PEX14 | 15668 | 13499 | -0.099 | -0.0692 | No | ||
| 122 | SH2D3C | 2864 | 13815 | -0.116 | -0.0850 | No | ||
| 123 | POU5F1 | 1547 23255 | 14116 | -0.135 | -0.0998 | No | ||
| 124 | GAS7 | 20836 1299 | 14223 | -0.144 | -0.1039 | No | ||
| 125 | ING3 | 17514 1019 | 14300 | -0.150 | -0.1064 | No | ||
| 126 | KCNC3 | 4939 9205 | 14386 | -0.159 | -0.1093 | No | ||
| 127 | CDK5R1 | 20745 | 14516 | -0.171 | -0.1144 | No | ||
| 128 | ABHD1 | 16890 3547 | 14518 | -0.171 | -0.1125 | No | ||
| 129 | HRH3 | 14317 | 14524 | -0.172 | -0.1109 | No | ||
| 130 | SLC18A3 | 21883 | 14583 | -0.178 | -0.1121 | No | ||
| 131 | KCNN2 | 8933 1949 | 14607 | -0.180 | -0.1114 | No | ||
| 132 | HNRPDL | 11947 3551 6954 | 14620 | -0.182 | -0.1100 | No | ||
| 133 | HTR7 | 23690 | 14729 | -0.194 | -0.1137 | No | ||
| 134 | HDAC6 | 24197 | 14856 | -0.210 | -0.1182 | No | ||
| 135 | KCNB1 | 14334 | 14865 | -0.211 | -0.1163 | No | ||
| 136 | FOXP4 | 22949 | 15015 | -0.230 | -0.1219 | No | ||
| 137 | ONECUT1 | 4858 | 15232 | -0.261 | -0.1307 | No | ||
| 138 | APLP2 | 19190 | 15312 | -0.276 | -0.1320 | No | ||
| 139 | SOX12 | 5478 | 15342 | -0.280 | -0.1304 | No | ||
| 140 | CS | 19839 | 15391 | -0.287 | -0.1299 | No | ||
| 141 | ROM1 | 23751 | 15429 | -0.294 | -0.1287 | No | ||
| 142 | ACE | 1419 4313 | 15573 | -0.326 | -0.1328 | No | ||
| 143 | NCOA6 | 2876 14382 | 15596 | -0.331 | -0.1304 | No | ||
| 144 | PHCA | 12301 7263 | 15640 | -0.340 | -0.1289 | No | ||
| 145 | OSBPL9 | 4141 | 15654 | -0.343 | -0.1259 | No | ||
| 146 | CBLN1 | 18800 | 15687 | -0.354 | -0.1237 | No | ||
| 147 | PIAS1 | 7126 | 15828 | -0.383 | -0.1271 | No | ||
| 148 | GPR3 | 15729 | 15836 | -0.386 | -0.1232 | No | ||
| 149 | CDC37 | 19214 | 15852 | -0.390 | -0.1197 | No | ||
| 150 | FKBP14 | 10552 | 15857 | -0.391 | -0.1156 | No | ||
| 151 | PMM1 | 22193 | 15924 | -0.409 | -0.1147 | No | ||
| 152 | DRAP1 | 23979 | 15948 | -0.416 | -0.1113 | No | ||
| 153 | TAF6 | 16322 891 | 16105 | -0.458 | -0.1147 | No | ||
| 154 | BAD | 24000 | 16322 | -0.526 | -0.1206 | No | ||
| 155 | MXD4 | 9346 | 16360 | -0.541 | -0.1167 | No | ||
| 156 | BCL7C | 4442 | 16419 | -0.559 | -0.1137 | No | ||
| 157 | CORO1C | 16425 | 16433 | -0.562 | -0.1082 | No | ||
| 158 | STMN1 | 9261 | 16546 | -0.609 | -0.1075 | No | ||
| 159 | TRIM28 | 18389 | 16604 | -0.634 | -0.1036 | No | ||
| 160 | PSMC6 | 7379 | 16655 | -0.652 | -0.0991 | No | ||
| 161 | UGP2 | 20518 | 16824 | -0.730 | -0.1002 | No | ||
| 162 | FBXO36 | 14205 | 16945 | -0.795 | -0.0979 | No | ||
| 163 | TAF11 | 12733 | 16966 | -0.807 | -0.0901 | No | ||
| 164 | PRKRA | 357 | 17196 | -0.918 | -0.0924 | No | ||
| 165 | DDAH2 | 23263 | 17200 | -0.920 | -0.0824 | No | ||
| 166 | SH3KBP1 | 2631 12206 7189 | 17203 | -0.921 | -0.0724 | No | ||
| 167 | RAP1GDS1 | 10487 | 17338 | -0.996 | -0.0686 | No | ||
| 168 | OAZ2 | 9499 | 17454 | -1.039 | -0.0634 | No | ||
| 169 | ALS2 | 7882 13151 | 17484 | -1.055 | -0.0533 | No | ||
| 170 | DDB1 | 23931 | 17493 | -1.060 | -0.0421 | No | ||
| 171 | FNTA | 18904 | 17524 | -1.076 | -0.0319 | No | ||
| 172 | NDRG1 | 22276 | 17581 | -1.116 | -0.0226 | No | ||
| 173 | COPS5 | 4117 14002 | 18012 | -1.419 | -0.0302 | No | ||
| 174 | KLHL7 | 16907 | 18534 | -2.424 | -0.0318 | No | ||
| 175 | HYAL2 | 4891 | 18601 | -3.278 | 0.0008 | No |