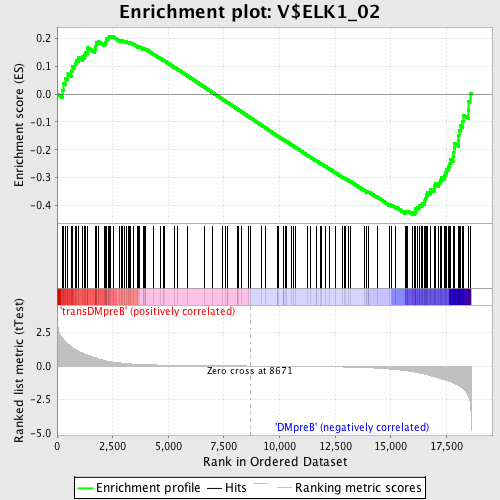
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | V$ELK1_02 |
| Enrichment Score (ES) | -0.43244487 |
| Normalized Enrichment Score (NES) | -1.2169846 |
| Nominal p-value | 0.08600337 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EIF4G1 | 22818 | 252 | 2.051 | 0.0134 | No | ||
| 2 | CHERP | 11367 | 290 | 1.945 | 0.0371 | No | ||
| 3 | GTF3C2 | 7749 | 380 | 1.797 | 0.0560 | No | ||
| 4 | GOLGA1 | 14595 | 488 | 1.641 | 0.0719 | No | ||
| 5 | UBE2N | 8216 19898 | 638 | 1.437 | 0.0828 | No | ||
| 6 | PURB | 5335 | 674 | 1.394 | 0.0993 | No | ||
| 7 | RAB11B | 9676 | 804 | 1.258 | 0.1090 | No | ||
| 8 | DHX30 | 18989 | 872 | 1.189 | 0.1210 | No | ||
| 9 | STK4 | 14743 | 965 | 1.084 | 0.1304 | No | ||
| 10 | SMS | 2573 24023 | 1142 | 0.980 | 0.1338 | No | ||
| 11 | SUPT5H | 9939 | 1232 | 0.907 | 0.1410 | No | ||
| 12 | MARK3 | 9369 | 1293 | 0.862 | 0.1491 | No | ||
| 13 | COG3 | 21751 | 1365 | 0.825 | 0.1561 | No | ||
| 14 | PRKACB | 15140 | 1381 | 0.813 | 0.1661 | No | ||
| 15 | NFYA | 5172 | 1703 | 0.626 | 0.1570 | No | ||
| 16 | MTBP | 22473 8374 | 1723 | 0.616 | 0.1641 | No | ||
| 17 | EIF5A | 11345 20379 6590 | 1726 | 0.615 | 0.1721 | No | ||
| 18 | COX8A | 8777 | 1748 | 0.599 | 0.1789 | No | ||
| 19 | ATP5A1 | 23505 | 1763 | 0.591 | 0.1859 | No | ||
| 20 | SEC24C | 22086 | 1839 | 0.542 | 0.1890 | No | ||
| 21 | HMGA1 | 23323 | 2116 | 0.415 | 0.1795 | No | ||
| 22 | MAPK8IP3 | 11402 23080 | 2160 | 0.395 | 0.1824 | No | ||
| 23 | STOML2 | 15902 | 2180 | 0.388 | 0.1865 | No | ||
| 24 | NAT5 | 18601 7496 12597 | 2199 | 0.385 | 0.1906 | No | ||
| 25 | ACP2 | 4318 | 2237 | 0.374 | 0.1936 | No | ||
| 26 | DNAJC7 | 20227 | 2240 | 0.373 | 0.1984 | No | ||
| 27 | EPN1 | 18401 | 2291 | 0.357 | 0.2004 | No | ||
| 28 | TRMT1 | 2307 12985 | 2328 | 0.347 | 0.2030 | No | ||
| 29 | FANCD2 | 17326 464 | 2356 | 0.335 | 0.2060 | No | ||
| 30 | SNRPB | 9842 5469 2736 | 2403 | 0.322 | 0.2078 | No | ||
| 31 | BMP2K | 3657 16786 | 2512 | 0.289 | 0.2057 | No | ||
| 32 | POLL | 23658 3688 | 2816 | 0.230 | 0.1924 | No | ||
| 33 | TRAF7 | 10325 | 2912 | 0.213 | 0.1900 | No | ||
| 34 | SMUG1 | 22109 | 2934 | 0.209 | 0.1916 | No | ||
| 35 | TRIM46 | 1779 15281 | 3014 | 0.198 | 0.1900 | No | ||
| 36 | TRIM44 | 8164 13487 | 3106 | 0.185 | 0.1875 | No | ||
| 37 | PHKB | 18537 3873 | 3196 | 0.172 | 0.1850 | No | ||
| 38 | NF1 | 5165 | 3253 | 0.164 | 0.1841 | No | ||
| 39 | TCF7 | 1467 20466 | 3291 | 0.161 | 0.1842 | No | ||
| 40 | DDX5 | 4606 8843 4607 | 3424 | 0.145 | 0.1790 | No | ||
| 41 | MTX2 | 14980 | 3620 | 0.129 | 0.1701 | No | ||
| 42 | GTF3C1 | 10607 17646 | 3650 | 0.126 | 0.1702 | No | ||
| 43 | DPP8 | 19408 | 3695 | 0.122 | 0.1695 | No | ||
| 44 | ZNF23 | 6149 | 3885 | 0.109 | 0.1607 | No | ||
| 45 | CALU | 1124 1149 8683 1162 4472 | 3891 | 0.108 | 0.1618 | No | ||
| 46 | MTPN | 17183 | 3913 | 0.107 | 0.1621 | No | ||
| 47 | SON | 5473 1657 1684 | 3954 | 0.105 | 0.1613 | No | ||
| 48 | DNAJA2 | 12133 18806 | 3986 | 0.103 | 0.1610 | No | ||
| 49 | BCL2L1 | 4440 2930 8652 | 4329 | 0.083 | 0.1436 | No | ||
| 50 | LYPLA2 | 15708 | 4635 | 0.071 | 0.1280 | No | ||
| 51 | OGG1 | 17331 | 4763 | 0.066 | 0.1220 | No | ||
| 52 | RPS18 | 1529 5397 1603 | 4809 | 0.065 | 0.1204 | No | ||
| 53 | SDHD | 19125 | 5278 | 0.051 | 0.0958 | No | ||
| 54 | CDC45L | 22642 1752 | 5403 | 0.048 | 0.0897 | No | ||
| 55 | RPL37 | 12502 7421 22521 | 5880 | 0.038 | 0.0644 | No | ||
| 56 | RBM22 | 23542 | 6627 | 0.026 | 0.0243 | No | ||
| 57 | NDUFS1 | 13940 | 6971 | 0.020 | 0.0060 | No | ||
| 58 | FMR1 | 24321 | 7412 | 0.015 | -0.0176 | No | ||
| 59 | CRK | 4559 1249 | 7549 | 0.013 | -0.0248 | No | ||
| 60 | LEPRE1 | 2384 12122 | 7662 | 0.012 | -0.0307 | No | ||
| 61 | FBXO3 | 2662 14934 | 7670 | 0.011 | -0.0309 | No | ||
| 62 | MYST2 | 20283 | 8091 | 0.007 | -0.0536 | No | ||
| 63 | ITGB1BP1 | 9189 | 8106 | 0.006 | -0.0542 | No | ||
| 64 | GABARAPL2 | 8211 13552 | 8166 | 0.006 | -0.0574 | No | ||
| 65 | INSIG2 | 3944 4124 4073 | 8293 | 0.004 | -0.0641 | No | ||
| 66 | HMG20A | 19436 | 8593 | 0.001 | -0.0803 | No | ||
| 67 | UBR1 | 5828 | 8690 | -0.000 | -0.0855 | No | ||
| 68 | SRP19 | 2004 12341 | 9192 | -0.006 | -0.1125 | No | ||
| 69 | CPSF3 | 7038 | 9369 | -0.009 | -0.1220 | No | ||
| 70 | ZBTB11 | 11309 | 9904 | -0.016 | -0.1507 | No | ||
| 71 | ACTR2 | 20523 | 9911 | -0.016 | -0.1508 | No | ||
| 72 | FBXO22 | 19442 | 9955 | -0.016 | -0.1529 | No | ||
| 73 | GTF2A2 | 10654 | 10157 | -0.019 | -0.1635 | No | ||
| 74 | NUDT12 | 7518 | 10158 | -0.019 | -0.1633 | No | ||
| 75 | EN1 | 387 14155 | 10178 | -0.019 | -0.1641 | No | ||
| 76 | PPIE | 15766 | 10269 | -0.020 | -0.1687 | No | ||
| 77 | IMMT | 367 8029 | 10295 | -0.021 | -0.1697 | No | ||
| 78 | TBCC | 23215 | 10538 | -0.024 | -0.1825 | No | ||
| 79 | PAPD4 | 8299 8300 4146 | 10625 | -0.026 | -0.1868 | No | ||
| 80 | STX12 | 4140 | 10696 | -0.026 | -0.1903 | No | ||
| 81 | CLDN7 | 20817 | 11238 | -0.034 | -0.2191 | No | ||
| 82 | SIRT6 | 19668 3447 | 11387 | -0.037 | -0.2266 | No | ||
| 83 | AKT1S1 | 12557 | 11638 | -0.041 | -0.2396 | No | ||
| 84 | SYT5 | 17986 | 11672 | -0.042 | -0.2409 | No | ||
| 85 | LSM5 | 7286 | 11845 | -0.046 | -0.2496 | No | ||
| 86 | FOXH1 | 22238 | 11877 | -0.046 | -0.2506 | No | ||
| 87 | TRO | 7071 | 12047 | -0.050 | -0.2591 | No | ||
| 88 | CGGBP1 | 22724 | 12253 | -0.055 | -0.2695 | No | ||
| 89 | NRAS | 5191 | 12495 | -0.061 | -0.2817 | No | ||
| 90 | IL2RG | 24096 | 12833 | -0.072 | -0.2990 | No | ||
| 91 | CRB3 | 23176 | 12916 | -0.075 | -0.3025 | No | ||
| 92 | HAT1 | 14990 | 12963 | -0.077 | -0.3040 | No | ||
| 93 | TBX6 | 1993 18075 | 13091 | -0.081 | -0.3098 | No | ||
| 94 | SHC1 | 9813 9812 5430 | 13187 | -0.085 | -0.3138 | No | ||
| 95 | VPS52 | 23287 1622 | 13812 | -0.116 | -0.3461 | No | ||
| 96 | GGTL3 | 14381 | 13924 | -0.122 | -0.3505 | No | ||
| 97 | ZNF22 | 12497 7417 | 13993 | -0.127 | -0.3525 | No | ||
| 98 | SAMD8 | 22082 | 14014 | -0.128 | -0.3519 | No | ||
| 99 | CLDN5 | 22826 | 14380 | -0.158 | -0.3695 | No | ||
| 100 | ELAVL1 | 4888 | 14931 | -0.219 | -0.3964 | No | ||
| 101 | UBE2L3 | 5818 22647 | 15040 | -0.233 | -0.3992 | No | ||
| 102 | PEX11A | 1056 5241 | 15229 | -0.260 | -0.4059 | No | ||
| 103 | PSMD12 | 20621 | 15664 | -0.347 | -0.4248 | No | ||
| 104 | NEDD1 | 19643 | 15691 | -0.355 | -0.4216 | No | ||
| 105 | NKIRAS2 | 12979 | 15770 | -0.370 | -0.4209 | No | ||
| 106 | TUFM | 10608 | 15984 | -0.427 | -0.4268 | Yes | ||
| 107 | ARCN1 | 19143 | 16063 | -0.448 | -0.4251 | Yes | ||
| 108 | NR1H2 | 17846 | 16108 | -0.459 | -0.4214 | Yes | ||
| 109 | BAT3 | 5896 | 16113 | -0.460 | -0.4156 | Yes | ||
| 110 | AP1M1 | 18571 | 16119 | -0.463 | -0.4097 | Yes | ||
| 111 | EEF1B2 | 4131 12063 | 16213 | -0.489 | -0.4083 | Yes | ||
| 112 | RAPH1 | 19110 | 16268 | -0.504 | -0.4046 | Yes | ||
| 113 | GART | 22543 1754 | 16281 | -0.510 | -0.3985 | Yes | ||
| 114 | TOMM40 | 6997 | 16391 | -0.550 | -0.3971 | Yes | ||
| 115 | PCYT1A | 4572 | 16425 | -0.560 | -0.3915 | Yes | ||
| 116 | MAPKAP1 | 15030 5993 | 16526 | -0.600 | -0.3890 | Yes | ||
| 117 | ZDHHC5 | 14547 | 16531 | -0.601 | -0.3813 | Yes | ||
| 118 | AP1B1 | 8599 | 16576 | -0.624 | -0.3754 | Yes | ||
| 119 | PRKAG2 | 24 | 16583 | -0.625 | -0.3675 | Yes | ||
| 120 | KRTCAP2 | 15541 | 16619 | -0.639 | -0.3610 | Yes | ||
| 121 | PSMC6 | 7379 | 16655 | -0.652 | -0.3543 | Yes | ||
| 122 | ITM2C | 14204 | 16765 | -0.706 | -0.3508 | Yes | ||
| 123 | RPL19 | 9736 | 16798 | -0.718 | -0.3431 | Yes | ||
| 124 | FBXO36 | 14205 | 16945 | -0.795 | -0.3405 | Yes | ||
| 125 | FBXW9 | 18540 | 16948 | -0.797 | -0.3301 | Yes | ||
| 126 | ARHGAP1 | 6001 10448 | 17007 | -0.832 | -0.3222 | Yes | ||
| 127 | PCQAP | 1661 13581 | 17164 | -0.903 | -0.3188 | Yes | ||
| 128 | TSTA3 | 22253 | 17227 | -0.936 | -0.3098 | Yes | ||
| 129 | UQCRH | 12378 | 17279 | -0.965 | -0.2998 | Yes | ||
| 130 | MRPL13 | 2249 22294 | 17396 | -1.010 | -0.2927 | Yes | ||
| 131 | UBADC1 | 2962 13608 | 17474 | -1.051 | -0.2830 | Yes | ||
| 132 | RFC4 | 1735 22627 | 17513 | -1.069 | -0.2709 | Yes | ||
| 133 | PSMC2 | 16909 | 17578 | -1.113 | -0.2597 | Yes | ||
| 134 | DDOST | 16021 | 17656 | -1.152 | -0.2487 | Yes | ||
| 135 | LMAN2 | 21454 | 17669 | -1.163 | -0.2340 | Yes | ||
| 136 | TBC1D17 | 6121 | 17797 | -1.251 | -0.2243 | Yes | ||
| 137 | UFD1L | 22825 | 17813 | -1.263 | -0.2085 | Yes | ||
| 138 | PSMC1 | 9635 | 17840 | -1.277 | -0.1930 | Yes | ||
| 139 | IPO4 | 7983 | 17857 | -1.291 | -0.1768 | Yes | ||
| 140 | HSPC171 | 18494 | 18047 | -1.450 | -0.1679 | Yes | ||
| 141 | SLC30A7 | 15186 | 18052 | -1.453 | -0.1489 | Yes | ||
| 142 | KLHDC3 | 22956 | 18067 | -1.470 | -0.1303 | Yes | ||
| 143 | RAP2C | 24156 | 18131 | -1.543 | -0.1133 | Yes | ||
| 144 | TIMM8B | 11391 | 18239 | -1.678 | -0.0969 | Yes | ||
| 145 | EXTL2 | 15440 | 18279 | -1.744 | -0.0760 | Yes | ||
| 146 | DIABLO | 16375 | 18496 | -2.246 | -0.0580 | Yes | ||
| 147 | MIR16 | 17663 1354 | 18505 | -2.285 | -0.0283 | Yes | ||
| 148 | MRPL3 | 3116 19335 | 18559 | -2.594 | 0.0031 | Yes |

