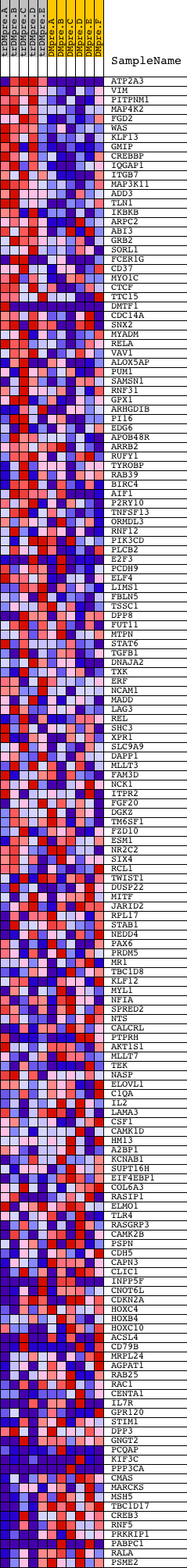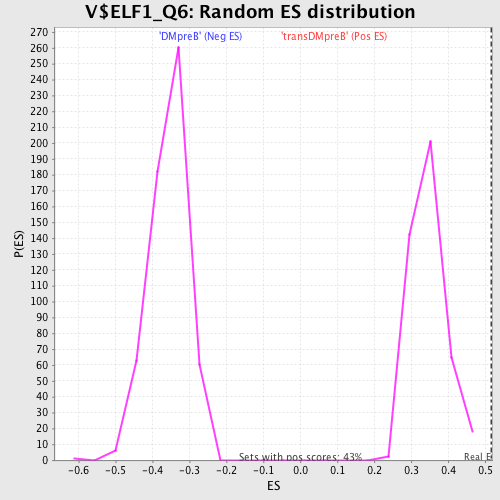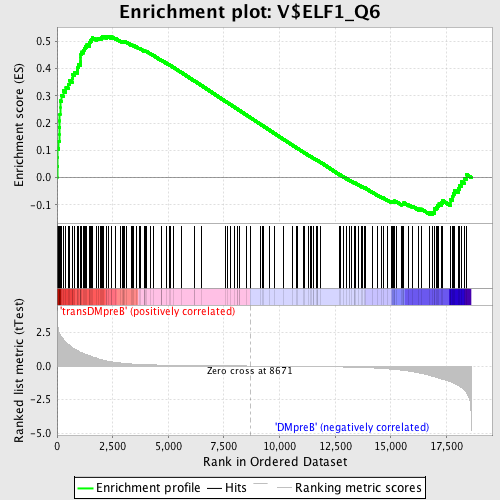
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | V$ELF1_Q6 |
| Enrichment Score (ES) | 0.51881665 |
| Normalized Enrichment Score (NES) | 1.4980701 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.16695063 |
| FWER p-Value | 0.838 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP2A3 | 20795 1314 | 3 | 3.765 | 0.0402 | Yes | ||
| 2 | VIM | 80 | 14 | 3.258 | 0.0745 | Yes | ||
| 3 | PITPNM1 | 23770 | 20 | 3.033 | 0.1067 | Yes | ||
| 4 | MAP4K2 | 6457 | 72 | 2.603 | 0.1318 | Yes | ||
| 5 | FGD2 | 23309 | 94 | 2.487 | 0.1573 | Yes | ||
| 6 | WAS | 24193 | 99 | 2.439 | 0.1832 | Yes | ||
| 7 | KLF13 | 17807 | 119 | 2.372 | 0.2076 | Yes | ||
| 8 | GMIP | 18597 | 124 | 2.341 | 0.2325 | Yes | ||
| 9 | CREBBP | 22682 8783 | 133 | 2.312 | 0.2568 | Yes | ||
| 10 | IQGAP1 | 6619 | 136 | 2.288 | 0.2812 | Yes | ||
| 11 | ITGB7 | 22112 | 191 | 2.180 | 0.3016 | Yes | ||
| 12 | MAP3K11 | 11163 | 275 | 2.001 | 0.3185 | Yes | ||
| 13 | ADD3 | 23812 3694 | 397 | 1.777 | 0.3310 | Yes | ||
| 14 | TLN1 | 15899 | 507 | 1.626 | 0.3425 | Yes | ||
| 15 | IKBKB | 4907 | 558 | 1.550 | 0.3564 | Yes | ||
| 16 | ARPC2 | 14225 | 677 | 1.393 | 0.3649 | Yes | ||
| 17 | ABI3 | 7314 | 691 | 1.380 | 0.3790 | Yes | ||
| 18 | GRB2 | 20149 | 792 | 1.266 | 0.3872 | Yes | ||
| 19 | SORL1 | 5474 | 904 | 1.159 | 0.3935 | Yes | ||
| 20 | FCER1G | 13759 | 931 | 1.131 | 0.4043 | Yes | ||
| 21 | CD37 | 17835 | 952 | 1.101 | 0.4150 | Yes | ||
| 22 | MYO1C | 20777 | 1045 | 1.018 | 0.4209 | Yes | ||
| 23 | CTCF | 18490 | 1058 | 1.013 | 0.4311 | Yes | ||
| 24 | TTC15 | 5720 | 1062 | 1.010 | 0.4417 | Yes | ||
| 25 | DMTF1 | 887 3662 16928 3530 | 1067 | 1.009 | 0.4523 | Yes | ||
| 26 | CDC14A | 15184 | 1112 | 0.993 | 0.4606 | Yes | ||
| 27 | SNX2 | 1974 23554 12585 | 1183 | 0.943 | 0.4669 | Yes | ||
| 28 | MYADM | 11946 | 1215 | 0.920 | 0.4751 | Yes | ||
| 29 | RELA | 23783 | 1277 | 0.876 | 0.4811 | Yes | ||
| 30 | VAV1 | 23173 | 1340 | 0.841 | 0.4868 | Yes | ||
| 31 | ALOX5AP | 16616 | 1445 | 0.771 | 0.4894 | Yes | ||
| 32 | PUM1 | 8160 | 1456 | 0.766 | 0.4971 | Yes | ||
| 33 | SAMSN1 | 7485 | 1490 | 0.749 | 0.5033 | Yes | ||
| 34 | RNF31 | 11206 | 1550 | 0.706 | 0.5077 | Yes | ||
| 35 | GPX1 | 19310 | 1580 | 0.693 | 0.5135 | Yes | ||
| 36 | ARHGDIB | 16950 | 1789 | 0.567 | 0.5083 | Yes | ||
| 37 | PI16 | 23310 | 1857 | 0.534 | 0.5104 | Yes | ||
| 38 | EDG6 | 19672 | 1960 | 0.476 | 0.5100 | Yes | ||
| 39 | APOB48R | 18081 | 2004 | 0.456 | 0.5125 | Yes | ||
| 40 | ARRB2 | 20806 | 2022 | 0.449 | 0.5164 | Yes | ||
| 41 | RUFY1 | 20476 | 2071 | 0.432 | 0.5185 | Yes | ||
| 42 | TYROBP | 18302 | 2209 | 0.381 | 0.5151 | Yes | ||
| 43 | RAB39 | 19113 | 2241 | 0.373 | 0.5174 | Yes | ||
| 44 | BIRC4 | 2638 8609 | 2287 | 0.358 | 0.5188 | Yes | ||
| 45 | AIF1 | 1582 23005 | 2447 | 0.306 | 0.5135 | No | ||
| 46 | P2RY10 | 2613 24269 | 2461 | 0.302 | 0.5160 | No | ||
| 47 | TNFSF13 | 12806 | 2618 | 0.265 | 0.5104 | No | ||
| 48 | ORMDL3 | 12386 | 2842 | 0.227 | 0.5007 | No | ||
| 49 | RNF12 | 5384 | 2950 | 0.207 | 0.4972 | No | ||
| 50 | PIK3CD | 9563 | 2973 | 0.204 | 0.4982 | No | ||
| 51 | PLCB2 | 5262 | 2988 | 0.201 | 0.4996 | No | ||
| 52 | E2F3 | 21500 8874 | 3044 | 0.194 | 0.4987 | No | ||
| 53 | PCDH9 | 21736 | 3111 | 0.185 | 0.4971 | No | ||
| 54 | ELF4 | 24162 | 3341 | 0.153 | 0.4863 | No | ||
| 55 | LIMS1 | 8493 4290 | 3372 | 0.150 | 0.4863 | No | ||
| 56 | FBLN5 | 21002 | 3448 | 0.142 | 0.4837 | No | ||
| 57 | TSSC1 | 2053 2063 6890 2144 | 3553 | 0.134 | 0.4795 | No | ||
| 58 | DPP8 | 19408 | 3695 | 0.122 | 0.4732 | No | ||
| 59 | FUT11 | 13084 | 3727 | 0.120 | 0.4728 | No | ||
| 60 | MTPN | 17183 | 3913 | 0.107 | 0.4639 | No | ||
| 61 | STAT6 | 19854 9909 | 3938 | 0.106 | 0.4638 | No | ||
| 62 | TGFB1 | 18332 | 3946 | 0.105 | 0.4645 | No | ||
| 63 | DNAJA2 | 12133 18806 | 3986 | 0.103 | 0.4635 | No | ||
| 64 | TXK | 16513 | 4030 | 0.100 | 0.4623 | No | ||
| 65 | ERF | 4680 8914 | 4183 | 0.090 | 0.4550 | No | ||
| 66 | NCAM1 | 5149 | 4347 | 0.082 | 0.4470 | No | ||
| 67 | MADD | 2926 14526 | 4688 | 0.069 | 0.4294 | No | ||
| 68 | LAG3 | 17000 | 4693 | 0.069 | 0.4299 | No | ||
| 69 | REL | 9716 | 4702 | 0.068 | 0.4302 | No | ||
| 70 | SHC3 | 21465 | 4896 | 0.062 | 0.4204 | No | ||
| 71 | XPR1 | 5383 | 5038 | 0.058 | 0.4134 | No | ||
| 72 | SLC9A9 | 11661 | 5080 | 0.056 | 0.4117 | No | ||
| 73 | DAPP1 | 11159 6445 6446 | 5222 | 0.053 | 0.4047 | No | ||
| 74 | MLLT3 | 2359 15847 2413 | 5589 | 0.044 | 0.3853 | No | ||
| 75 | FAM3D | 21922 | 6164 | 0.033 | 0.3546 | No | ||
| 76 | NCK1 | 9447 5152 | 6189 | 0.033 | 0.3536 | No | ||
| 77 | ITPR2 | 9194 | 6505 | 0.028 | 0.3369 | No | ||
| 78 | FGF20 | 18892 | 7588 | 0.013 | 0.2784 | No | ||
| 79 | DGKZ | 2836 14522 | 7644 | 0.012 | 0.2755 | No | ||
| 80 | TM6SF1 | 8416 | 7805 | 0.010 | 0.2670 | No | ||
| 81 | FZD10 | 13565 16693 | 7981 | 0.008 | 0.2576 | No | ||
| 82 | ESM1 | 21560 | 8098 | 0.006 | 0.2513 | No | ||
| 83 | NR2C2 | 10216 | 8122 | 0.006 | 0.2502 | No | ||
| 84 | SIX4 | 5445 | 8186 | 0.006 | 0.2468 | No | ||
| 85 | RCL1 | 3728 3743 23894 | 8219 | 0.005 | 0.2451 | No | ||
| 86 | TWIST1 | 21291 | 8501 | 0.002 | 0.2299 | No | ||
| 87 | DUSP22 | 466 21680 | 8675 | -0.000 | 0.2206 | No | ||
| 88 | MITF | 17349 | 9128 | -0.005 | 0.1961 | No | ||
| 89 | JARID2 | 9200 | 9218 | -0.007 | 0.1914 | No | ||
| 90 | RPL17 | 11429 6653 | 9298 | -0.008 | 0.1872 | No | ||
| 91 | STAB1 | 21892 | 9553 | -0.011 | 0.1735 | No | ||
| 92 | NEDD4 | 19384 3101 | 9763 | -0.014 | 0.1624 | No | ||
| 93 | PAX6 | 5223 | 10166 | -0.019 | 0.1408 | No | ||
| 94 | PRDM5 | 17424 | 10599 | -0.025 | 0.1176 | No | ||
| 95 | MR1 | 4837 | 10777 | -0.028 | 0.1084 | No | ||
| 96 | TBC1D8 | 12044 | 10815 | -0.028 | 0.1066 | No | ||
| 97 | KLF12 | 4960 2637 21732 9228 | 11071 | -0.032 | 0.0932 | No | ||
| 98 | MYL1 | 4015 | 11099 | -0.032 | 0.0921 | No | ||
| 99 | NFIA | 16172 5170 | 11303 | -0.036 | 0.0814 | No | ||
| 100 | SPRED2 | 8539 4332 | 11409 | -0.037 | 0.0762 | No | ||
| 101 | NTS | 19635 3297 | 11414 | -0.037 | 0.0763 | No | ||
| 102 | CALCRL | 12041 | 11508 | -0.039 | 0.0717 | No | ||
| 103 | PTPRH | 11861 | 11537 | -0.040 | 0.0706 | No | ||
| 104 | AKT1S1 | 12557 | 11638 | -0.041 | 0.0657 | No | ||
| 105 | MLLT7 | 24278 | 11716 | -0.043 | 0.0619 | No | ||
| 106 | TEK | 16175 | 11723 | -0.043 | 0.0621 | No | ||
| 107 | NASP | 2383 2387 6955 | 11839 | -0.046 | 0.0563 | No | ||
| 108 | ELOVL1 | 16114 2484 12026 | 12709 | -0.068 | 0.0100 | No | ||
| 109 | C1QA | 8666 | 12718 | -0.068 | 0.0103 | No | ||
| 110 | IL2 | 15354 | 12873 | -0.073 | 0.0027 | No | ||
| 111 | LAMA3 | 23620 2009 | 13027 | -0.079 | -0.0047 | No | ||
| 112 | CSF1 | 15202 | 13152 | -0.084 | -0.0105 | No | ||
| 113 | CAMK1D | 5977 2948 | 13212 | -0.086 | -0.0128 | No | ||
| 114 | HM13 | 2880 2707 14793 | 13345 | -0.091 | -0.0190 | No | ||
| 115 | A2BP1 | 11212 1652 1689 | 13356 | -0.092 | -0.0186 | No | ||
| 116 | KCNAB1 | 1791 15577 | 13396 | -0.094 | -0.0197 | No | ||
| 117 | SUPT16H | 8541 | 13534 | -0.101 | -0.0260 | No | ||
| 118 | EIF4EBP1 | 8891 4661 | 13698 | -0.110 | -0.0337 | No | ||
| 119 | COL6A3 | 4546 13885 13884 | 13739 | -0.111 | -0.0347 | No | ||
| 120 | RASIP1 | 12838 | 13828 | -0.117 | -0.0382 | No | ||
| 121 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 13839 | -0.118 | -0.0374 | No | ||
| 122 | TLR4 | 2329 10191 5770 | 14171 | -0.139 | -0.0539 | No | ||
| 123 | RASGRP3 | 10767 | 14421 | -0.162 | -0.0656 | No | ||
| 124 | CAMK2B | 20536 | 14588 | -0.178 | -0.0727 | No | ||
| 125 | PSPN | 22921 | 14648 | -0.185 | -0.0739 | No | ||
| 126 | CDH5 | 18512 | 14864 | -0.211 | -0.0833 | No | ||
| 127 | CAPN3 | 8686 2868 | 15012 | -0.230 | -0.0888 | No | ||
| 128 | CLIC1 | 23262 | 15052 | -0.234 | -0.0884 | No | ||
| 129 | INPP5F | 18055 3877 | 15134 | -0.245 | -0.0902 | No | ||
| 130 | CNOT6L | 6084 | 15143 | -0.247 | -0.0880 | No | ||
| 131 | CDKN2A | 2491 15841 | 15149 | -0.248 | -0.0856 | No | ||
| 132 | HOXC4 | 4864 | 15252 | -0.263 | -0.0883 | No | ||
| 133 | HOXB4 | 20686 | 15485 | -0.307 | -0.0976 | No | ||
| 134 | HOXC10 | 22342 | 15539 | -0.316 | -0.0971 | No | ||
| 135 | ACSL4 | 24043 11936 24042 2614 6940 | 15587 | -0.328 | -0.0961 | No | ||
| 136 | CD79B | 20185 1309 | 15588 | -0.328 | -0.0926 | No | ||
| 137 | MRPL24 | 15556 | 15805 | -0.378 | -0.1003 | No | ||
| 138 | AGPAT1 | 23275 | 15995 | -0.430 | -0.1059 | No | ||
| 139 | RAB25 | 15287 | 16264 | -0.502 | -0.1150 | No | ||
| 140 | RAC1 | 16302 | 16377 | -0.547 | -0.1153 | No | ||
| 141 | CENTA1 | 10546 16319 | 16732 | -0.690 | -0.1270 | No | ||
| 142 | IL7R | 9175 4922 | 16887 | -0.764 | -0.1272 | No | ||
| 143 | GPR120 | 23869 | 16959 | -0.803 | -0.1225 | No | ||
| 144 | STIM1 | 18164 | 16965 | -0.807 | -0.1141 | No | ||
| 145 | DPP3 | 7954 3746 | 17037 | -0.851 | -0.1088 | No | ||
| 146 | GNGT2 | 20691 | 17099 | -0.878 | -0.1027 | No | ||
| 147 | PCQAP | 1661 13581 | 17164 | -0.903 | -0.0965 | No | ||
| 148 | KIF3C | 9220 4954 | 17287 | -0.971 | -0.0927 | No | ||
| 149 | PPP3CA | 1863 5284 | 17331 | -0.993 | -0.0844 | No | ||
| 150 | CMAS | 17245 | 17689 | -1.175 | -0.0912 | No | ||
| 151 | MARCKS | 9331 | 17692 | -1.175 | -0.0787 | No | ||
| 152 | MSH5 | 23010 1599 | 17786 | -1.243 | -0.0704 | No | ||
| 153 | TBC1D17 | 6121 | 17797 | -1.251 | -0.0576 | No | ||
| 154 | CREB3 | 16231 | 17855 | -1.288 | -0.0469 | No | ||
| 155 | RNF5 | 63 | 18034 | -1.434 | -0.0412 | No | ||
| 156 | PRKRIP1 | 12407 7336 | 18075 | -1.483 | -0.0275 | No | ||
| 157 | PABPC1 | 5219 9522 9523 23572 | 18166 | -1.581 | -0.0154 | No | ||
| 158 | RALA | 21541 | 18299 | -1.773 | -0.0036 | No | ||
| 159 | PSME2 | 5301 | 18390 | -1.933 | 0.0122 | No |