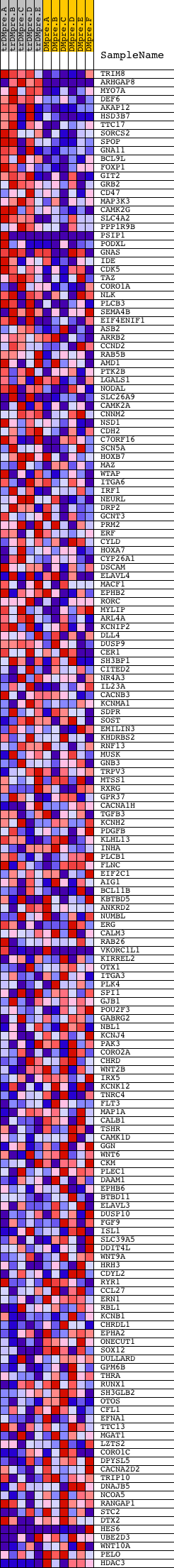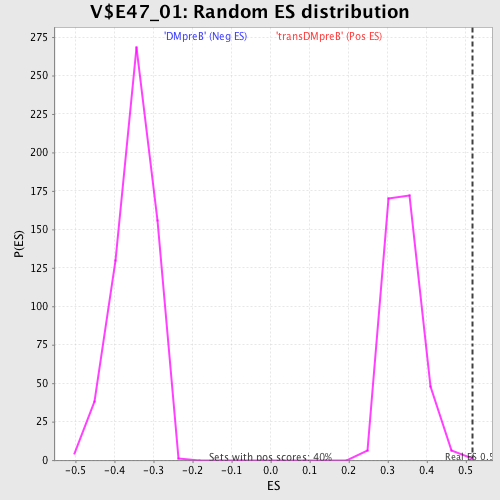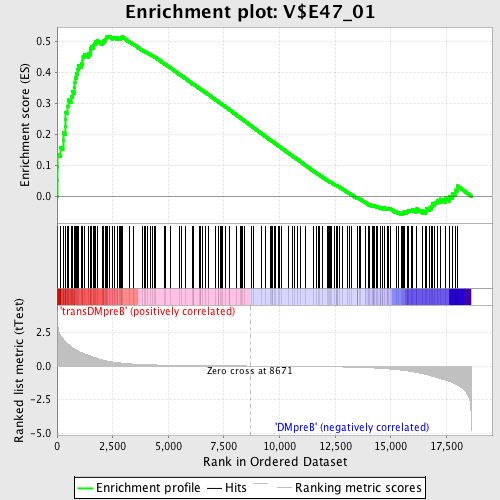
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | V$E47_01 |
| Enrichment Score (ES) | 0.5178849 |
| Normalized Enrichment Score (NES) | 1.5254688 |
| Nominal p-value | 0.0024813896 |
| FDR q-value | 0.17291573 |
| FWER p-Value | 0.719 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TRIM8 | 23824 | 5 | 3.715 | 0.0499 | Yes | ||
| 2 | ARHGAP8 | 4268 2196 | 8 | 3.396 | 0.0957 | Yes | ||
| 3 | MYO7A | 9439 | 33 | 2.928 | 0.1339 | Yes | ||
| 4 | DEF6 | 23319 | 145 | 2.255 | 0.1583 | Yes | ||
| 5 | AKAP12 | 17600 | 266 | 2.027 | 0.1792 | Yes | ||
| 6 | HSD3B7 | 4158 | 272 | 2.007 | 0.2060 | Yes | ||
| 7 | TTC17 | 13219 | 375 | 1.799 | 0.2248 | Yes | ||
| 8 | SORCS2 | 16553 | 382 | 1.794 | 0.2487 | Yes | ||
| 9 | SPOP | 5498 | 390 | 1.780 | 0.2724 | Yes | ||
| 10 | GNA11 | 3300 3345 19670 | 465 | 1.674 | 0.2909 | Yes | ||
| 11 | BCL9L | 19475 | 495 | 1.633 | 0.3114 | Yes | ||
| 12 | FOXP1 | 4242 | 632 | 1.442 | 0.3235 | Yes | ||
| 13 | GIT2 | 11174 6462 | 692 | 1.380 | 0.3390 | Yes | ||
| 14 | GRB2 | 20149 | 792 | 1.266 | 0.3507 | Yes | ||
| 15 | CD47 | 4933 | 801 | 1.260 | 0.3673 | Yes | ||
| 16 | MAP3K3 | 20626 | 808 | 1.254 | 0.3839 | Yes | ||
| 17 | CAMK2G | 21905 | 866 | 1.191 | 0.3969 | Yes | ||
| 18 | SLC4A2 | 9828 | 911 | 1.153 | 0.4101 | Yes | ||
| 19 | PPP1R9B | 20698 | 947 | 1.108 | 0.4231 | Yes | ||
| 20 | PSIP1 | 4159 2507 8319 | 1095 | 0.999 | 0.4286 | Yes | ||
| 21 | PODXL | 17190 | 1160 | 0.963 | 0.4382 | Yes | ||
| 22 | GNAS | 9025 2963 2752 | 1162 | 0.961 | 0.4511 | Yes | ||
| 23 | IDE | 4893 | 1227 | 0.912 | 0.4599 | Yes | ||
| 24 | CDK5 | 16591 | 1415 | 0.791 | 0.4605 | Yes | ||
| 25 | TAZ | 12415 | 1487 | 0.750 | 0.4668 | Yes | ||
| 26 | CORO1A | 1066 17631 1074 | 1492 | 0.747 | 0.4766 | Yes | ||
| 27 | NLK | 5179 5178 | 1529 | 0.723 | 0.4844 | Yes | ||
| 28 | PLCB3 | 23799 | 1635 | 0.661 | 0.4877 | Yes | ||
| 29 | SEMA4B | 18205 | 1660 | 0.651 | 0.4952 | Yes | ||
| 30 | EIF4ENIF1 | 20971 | 1724 | 0.615 | 0.5001 | Yes | ||
| 31 | ASB2 | 20996 | 1808 | 0.558 | 0.5031 | Yes | ||
| 32 | ARRB2 | 20806 | 2022 | 0.449 | 0.4976 | Yes | ||
| 33 | CCND2 | 16987 | 2088 | 0.425 | 0.4998 | Yes | ||
| 34 | RAB5B | 359 19591 3430 | 2096 | 0.421 | 0.5051 | Yes | ||
| 35 | AMD1 | 8583 | 2155 | 0.398 | 0.5074 | Yes | ||
| 36 | PTK2B | 21776 | 2197 | 0.385 | 0.5103 | Yes | ||
| 37 | LGALS1 | 22429 | 2203 | 0.383 | 0.5152 | Yes | ||
| 38 | NODAL | 333 20010 | 2247 | 0.369 | 0.5179 | Yes | ||
| 39 | SLC26A9 | 11560 980 4067 | 2360 | 0.333 | 0.5163 | No | ||
| 40 | CAMK2A | 2024 23541 1980 | 2511 | 0.290 | 0.5121 | No | ||
| 41 | CNNM2 | 8251 | 2573 | 0.276 | 0.5125 | No | ||
| 42 | NSD1 | 2134 5197 | 2698 | 0.250 | 0.5092 | No | ||
| 43 | CDH2 | 1963 8727 8726 4508 | 2701 | 0.250 | 0.5124 | No | ||
| 44 | C7ORF16 | 17434 | 2793 | 0.234 | 0.5107 | No | ||
| 45 | SCN5A | 5412 | 2868 | 0.222 | 0.5097 | No | ||
| 46 | HOXB7 | 666 9110 | 2877 | 0.221 | 0.5122 | No | ||
| 47 | MAZ | 1327 17623 | 2901 | 0.216 | 0.5139 | No | ||
| 48 | WTAP | 7215 | 2917 | 0.212 | 0.5159 | No | ||
| 49 | ITGA6 | 4930 | 3269 | 0.163 | 0.4991 | No | ||
| 50 | IRF1 | 1336 1258 1433 9182 | 3435 | 0.144 | 0.4921 | No | ||
| 51 | NEURL | 5164 | 3823 | 0.113 | 0.4726 | No | ||
| 52 | DRP2 | 24254 | 3932 | 0.106 | 0.4682 | No | ||
| 53 | GCNT3 | 12993 | 3969 | 0.104 | 0.4677 | No | ||
| 54 | PRM2 | 9621 | 4043 | 0.099 | 0.4650 | No | ||
| 55 | ERF | 4680 8914 | 4183 | 0.090 | 0.4587 | No | ||
| 56 | CYLD | 18532 | 4266 | 0.086 | 0.4554 | No | ||
| 57 | HOXA7 | 4862 | 4378 | 0.081 | 0.4505 | No | ||
| 58 | CYP26A1 | 23871 3695 | 4425 | 0.079 | 0.4491 | No | ||
| 59 | DSCAM | 1667 4641 22530 | 4828 | 0.064 | 0.4282 | No | ||
| 60 | ELAVL4 | 15805 4889 9137 | 4859 | 0.063 | 0.4274 | No | ||
| 61 | MACF1 | 4316 15763 2382 15764 | 5108 | 0.055 | 0.4147 | No | ||
| 62 | EPHB2 | 4675 2440 8910 | 5497 | 0.046 | 0.3943 | No | ||
| 63 | RORC | 9732 | 5591 | 0.044 | 0.3898 | No | ||
| 64 | MYLIP | 21652 3189 | 5767 | 0.040 | 0.3809 | No | ||
| 65 | ARL4A | 8627 | 6104 | 0.034 | 0.3631 | No | ||
| 66 | KCNIP2 | 23655 3725 | 6122 | 0.034 | 0.3627 | No | ||
| 67 | DLL4 | 7040 | 6145 | 0.034 | 0.3619 | No | ||
| 68 | DUSP9 | 24307 | 6388 | 0.030 | 0.3492 | No | ||
| 69 | CER1 | 15853 | 6444 | 0.029 | 0.3466 | No | ||
| 70 | SH3BP1 | 632 22431 | 6449 | 0.029 | 0.3468 | No | ||
| 71 | CITED2 | 5118 14477 | 6554 | 0.027 | 0.3415 | No | ||
| 72 | NR4A3 | 9473 16212 5183 | 6648 | 0.025 | 0.3368 | No | ||
| 73 | IL23A | 19598 | 6795 | 0.023 | 0.3292 | No | ||
| 74 | CACNB3 | 22373 8679 | 7126 | 0.018 | 0.3115 | No | ||
| 75 | KCNMA1 | 4947 | 7243 | 0.017 | 0.3055 | No | ||
| 76 | SDPR | 14252 | 7261 | 0.017 | 0.3048 | No | ||
| 77 | SOST | 20210 | 7341 | 0.016 | 0.3007 | No | ||
| 78 | EMILIN3 | 11376 | 7378 | 0.015 | 0.2990 | No | ||
| 79 | KHDRBS2 | 14284 | 7428 | 0.015 | 0.2965 | No | ||
| 80 | RNF13 | 6271 | 7551 | 0.013 | 0.2901 | No | ||
| 81 | MUSK | 5199 | 7752 | 0.010 | 0.2794 | No | ||
| 82 | GNB3 | 9027 | 7761 | 0.010 | 0.2791 | No | ||
| 83 | TRPV3 | 20788 | 8056 | 0.007 | 0.2632 | No | ||
| 84 | MTSS1 | 5585 22285 | 8257 | 0.005 | 0.2524 | No | ||
| 85 | RXRG | 14063 | 8272 | 0.005 | 0.2517 | No | ||
| 86 | GPR37 | 17202 | 8328 | 0.004 | 0.2488 | No | ||
| 87 | CACNA1H | 23077 | 8432 | 0.003 | 0.2433 | No | ||
| 88 | TGFB3 | 10161 | 8719 | -0.001 | 0.2278 | No | ||
| 89 | KCNH2 | 16592 | 8828 | -0.002 | 0.2219 | No | ||
| 90 | PDGFB | 22199 2311 | 9172 | -0.006 | 0.2034 | No | ||
| 91 | KLHL13 | 12536 | 9385 | -0.009 | 0.1920 | No | ||
| 92 | INHA | 14214 | 9587 | -0.011 | 0.1813 | No | ||
| 93 | PLCB1 | 14832 2821 | 9656 | -0.012 | 0.1778 | No | ||
| 94 | FLNC | 4302 8510 | 9670 | -0.013 | 0.1772 | No | ||
| 95 | EIF2C1 | 10672 | 9702 | -0.013 | 0.1757 | No | ||
| 96 | AIG1 | 7272 | 9766 | -0.014 | 0.1725 | No | ||
| 97 | BCL11B | 7190 | 9826 | -0.015 | 0.1695 | No | ||
| 98 | KBTBD5 | 13020 | 9928 | -0.016 | 0.1642 | No | ||
| 99 | ANKRD2 | 23853 | 10014 | -0.017 | 0.1599 | No | ||
| 100 | NUMBL | 1663 9496 | 10090 | -0.018 | 0.1560 | No | ||
| 101 | ERG | 1686 8915 | 10402 | -0.022 | 0.1395 | No | ||
| 102 | CALM3 | 8682 | 10585 | -0.025 | 0.1299 | No | ||
| 103 | RAB26 | 11609 | 10665 | -0.026 | 0.1260 | No | ||
| 104 | VKORC1L1 | 102 7625 | 10824 | -0.028 | 0.1178 | No | ||
| 105 | KIRREL2 | 17889 3922 | 10918 | -0.030 | 0.1132 | No | ||
| 106 | OTX1 | 20517 | 11153 | -0.033 | 0.1009 | No | ||
| 107 | ITGA3 | 20284 | 11505 | -0.039 | 0.0824 | No | ||
| 108 | PLK4 | 1870 1874 | 11649 | -0.042 | 0.0752 | No | ||
| 109 | SPI1 | 14949 | 11750 | -0.044 | 0.0704 | No | ||
| 110 | GJB1 | 24276 | 11774 | -0.044 | 0.0698 | No | ||
| 111 | POU2F3 | 9602 | 11946 | -0.048 | 0.0611 | No | ||
| 112 | GABRG2 | 1259 4748 8996 1331 | 12136 | -0.052 | 0.0516 | No | ||
| 113 | NBL1 | 15699 | 12212 | -0.054 | 0.0483 | No | ||
| 114 | KCNJ4 | 22205 | 12261 | -0.055 | 0.0464 | No | ||
| 115 | PAK3 | 9528 | 12282 | -0.056 | 0.0461 | No | ||
| 116 | CORO2A | 4216 8411 | 12340 | -0.057 | 0.0437 | No | ||
| 117 | CHRD | 22817 | 12453 | -0.060 | 0.0385 | No | ||
| 118 | WNT2B | 15214 1838 | 12537 | -0.063 | 0.0348 | No | ||
| 119 | IRX5 | 18527 | 12551 | -0.063 | 0.0350 | No | ||
| 120 | KCNK12 | 9978 | 12602 | -0.064 | 0.0331 | No | ||
| 121 | TNRC4 | 15514 | 12615 | -0.065 | 0.0334 | No | ||
| 122 | FLT3 | 16288 | 12713 | -0.068 | 0.0290 | No | ||
| 123 | MAP1A | 5124 | 12841 | -0.072 | 0.0231 | No | ||
| 124 | CALB1 | 4469 | 13044 | -0.080 | 0.0132 | No | ||
| 125 | TSHR | 5801 10223 | 13143 | -0.083 | 0.0090 | No | ||
| 126 | CAMK1D | 5977 2948 | 13212 | -0.086 | 0.0065 | No | ||
| 127 | GGN | 2472 1171 18312 | 13481 | -0.098 | -0.0067 | No | ||
| 128 | WNT6 | 5881 | 13500 | -0.099 | -0.0063 | No | ||
| 129 | CKM | 18362 | 13520 | -0.100 | -0.0060 | No | ||
| 130 | PLEC1 | 2273 2198 2205 2178 2230 22247 2213 2231 2295 2172 5263 2266 | 13573 | -0.103 | -0.0075 | No | ||
| 131 | DAAM1 | 5536 | 13639 | -0.106 | -0.0095 | No | ||
| 132 | EPHB6 | 17473 | 13860 | -0.119 | -0.0199 | No | ||
| 133 | BTBD11 | 13147 | 14017 | -0.128 | -0.0266 | No | ||
| 134 | ELAVL3 | 9136 | 14054 | -0.131 | -0.0268 | No | ||
| 135 | DUSP10 | 4003 14016 | 14184 | -0.140 | -0.0319 | No | ||
| 136 | FGF9 | 8966 | 14217 | -0.143 | -0.0317 | No | ||
| 137 | ISL1 | 21338 | 14221 | -0.144 | -0.0299 | No | ||
| 138 | SLC39A5 | 3364 19596 | 14272 | -0.147 | -0.0306 | No | ||
| 139 | DDIT4L | 15410 | 14368 | -0.156 | -0.0337 | No | ||
| 140 | WNT9A | 5709 | 14415 | -0.162 | -0.0340 | No | ||
| 141 | HRH3 | 14317 | 14524 | -0.172 | -0.0375 | No | ||
| 142 | CDYL2 | 18736 | 14528 | -0.172 | -0.0354 | No | ||
| 143 | RYR1 | 17903 17902 9770 | 14642 | -0.185 | -0.0390 | No | ||
| 144 | CCL27 | 5419 | 14643 | -0.185 | -0.0365 | No | ||
| 145 | ERN1 | 8137 | 14698 | -0.190 | -0.0369 | No | ||
| 146 | RBL1 | 14373 | 14829 | -0.206 | -0.0411 | No | ||
| 147 | KCNB1 | 14334 | 14865 | -0.211 | -0.0402 | No | ||
| 148 | CHRDL1 | 8180 13504 | 14911 | -0.217 | -0.0397 | No | ||
| 149 | EPHA2 | 16006 | 14984 | -0.226 | -0.0405 | No | ||
| 150 | ONECUT1 | 4858 | 15232 | -0.261 | -0.0504 | No | ||
| 151 | SOX12 | 5478 | 15342 | -0.280 | -0.0525 | No | ||
| 152 | DULLARD | 20816 | 15458 | -0.300 | -0.0547 | No | ||
| 153 | GPM6B | 9034 | 15537 | -0.316 | -0.0547 | No | ||
| 154 | THRA | 1447 10171 1406 | 15565 | -0.323 | -0.0518 | No | ||
| 155 | RUNX1 | 4481 | 15616 | -0.335 | -0.0500 | No | ||
| 156 | SH3GLB2 | 14628 | 15727 | -0.361 | -0.0511 | No | ||
| 157 | OTOS | 13877 | 15765 | -0.368 | -0.0481 | No | ||
| 158 | CFL1 | 4516 | 15803 | -0.378 | -0.0450 | No | ||
| 159 | EFNA1 | 15279 | 15909 | -0.406 | -0.0452 | No | ||
| 160 | TTC13 | 10636 | 15982 | -0.427 | -0.0433 | No | ||
| 161 | MGAT1 | 9384 | 16133 | -0.467 | -0.0452 | No | ||
| 162 | LZTS2 | 10362 | 16172 | -0.479 | -0.0408 | No | ||
| 163 | CORO1C | 16425 | 16433 | -0.562 | -0.0473 | No | ||
| 164 | DPYSL5 | 92 | 16579 | -0.624 | -0.0467 | No | ||
| 165 | CACNA2D2 | 7154 | 16605 | -0.635 | -0.0395 | No | ||
| 166 | TRIP10 | 23174 | 16748 | -0.697 | -0.0378 | No | ||
| 167 | DNAJB5 | 16236 | 16830 | -0.732 | -0.0323 | No | ||
| 168 | NCOA5 | 14341 | 16875 | -0.760 | -0.0244 | No | ||
| 169 | RANGAP1 | 2180 22195 | 16983 | -0.817 | -0.0192 | No | ||
| 170 | STC2 | 5526 | 17079 | -0.870 | -0.0126 | No | ||
| 171 | DTX2 | 16674 3488 | 17246 | -0.947 | -0.0088 | No | ||
| 172 | HES6 | 13882 4058 | 17445 | -1.035 | -0.0055 | No | ||
| 173 | UBE2D3 | 7253 | 17639 | -1.143 | -0.0006 | No | ||
| 174 | WNT10A | 14221 | 17775 | -1.232 | 0.0087 | No | ||
| 175 | PELO | 8360 | 17891 | -1.314 | 0.0203 | No | ||
| 176 | HDAC3 | 4844 | 18001 | -1.408 | 0.0333 | No |