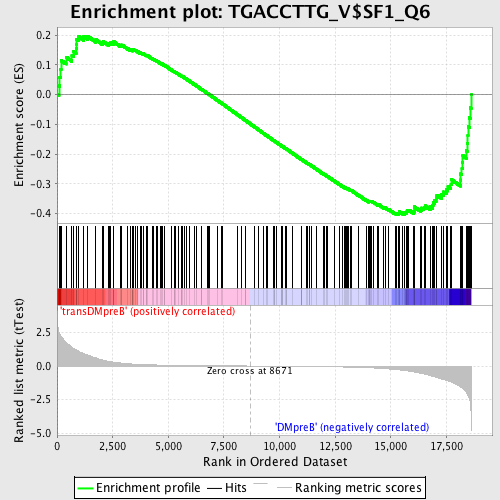
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | TGACCTTG_V$SF1_Q6 |
| Enrichment Score (ES) | -0.40338537 |
| Normalized Enrichment Score (NES) | -1.1602908 |
| Nominal p-value | 0.1362862 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CHD2 | 10847 | 85 | 2.534 | 0.0293 | No | ||
| 2 | PCTK1 | 24369 | 122 | 2.344 | 0.0587 | No | ||
| 3 | RANBP10 | 18764 | 167 | 2.217 | 0.0859 | No | ||
| 4 | STAC2 | 20265 | 178 | 2.204 | 0.1149 | No | ||
| 5 | MEF2C | 3204 9378 | 403 | 1.759 | 0.1263 | No | ||
| 6 | RBMS1 | 14580 | 667 | 1.405 | 0.1308 | No | ||
| 7 | VAMP2 | 20826 | 750 | 1.302 | 0.1437 | No | ||
| 8 | KIN | 15120 | 854 | 1.200 | 0.1542 | No | ||
| 9 | CAMK2G | 21905 | 866 | 1.191 | 0.1695 | No | ||
| 10 | AK2 | 8563 2479 16073 | 873 | 1.189 | 0.1851 | No | ||
| 11 | ESRRA | 23802 | 951 | 1.103 | 0.1957 | No | ||
| 12 | ZRANB1 | 6884 11702 | 1201 | 0.927 | 0.1946 | No | ||
| 13 | CCNE1 | 17857 | 1369 | 0.822 | 0.1965 | No | ||
| 14 | ZDHHC21 | 15854 | 1742 | 0.604 | 0.1845 | No | ||
| 15 | CRSP7 | 18838 | 2061 | 0.436 | 0.1731 | No | ||
| 16 | CKB | 4523 | 2083 | 0.426 | 0.1776 | No | ||
| 17 | APP | 4402 | 2320 | 0.349 | 0.1695 | No | ||
| 18 | FANCD2 | 17326 464 | 2356 | 0.335 | 0.1721 | No | ||
| 19 | UBL3 | 16284 | 2395 | 0.324 | 0.1743 | No | ||
| 20 | PANK1 | 1687 23692 | 2523 | 0.287 | 0.1713 | No | ||
| 21 | COMTD1 | 21903 | 2533 | 0.285 | 0.1746 | No | ||
| 22 | SV2A | 15498 12250 | 2551 | 0.280 | 0.1774 | No | ||
| 23 | SCN5A | 5412 | 2868 | 0.222 | 0.1633 | No | ||
| 24 | GABRA3 | 24139 | 2905 | 0.214 | 0.1642 | No | ||
| 25 | TRAP1 | 22683 | 2911 | 0.213 | 0.1668 | No | ||
| 26 | MAL | 2760 14437 | 3174 | 0.175 | 0.1549 | No | ||
| 27 | ATP5B | 19846 | 3318 | 0.157 | 0.1492 | No | ||
| 28 | RAB3A | 9681 | 3371 | 0.150 | 0.1484 | No | ||
| 29 | PPP1R16B | 6014 | 3381 | 0.149 | 0.1499 | No | ||
| 30 | ALDOA | 8572 | 3405 | 0.147 | 0.1507 | No | ||
| 31 | CEL | 8734 | 3415 | 0.146 | 0.1521 | No | ||
| 32 | TNNI1 | 14116 | 3515 | 0.137 | 0.1486 | No | ||
| 33 | MTX2 | 14980 | 3620 | 0.129 | 0.1447 | No | ||
| 34 | CATSPER2 | 10014 | 3733 | 0.119 | 0.1402 | No | ||
| 35 | ATP6AP2 | 2567 24380 2579 2656 | 3787 | 0.115 | 0.1389 | No | ||
| 36 | KLK15 | 11404 | 3868 | 0.110 | 0.1360 | No | ||
| 37 | TFRC | 22785 | 3875 | 0.109 | 0.1371 | No | ||
| 38 | RAPGEF5 | 5739 10155 21123 | 4014 | 0.101 | 0.1310 | No | ||
| 39 | CHGA | 21178 2151 | 4015 | 0.101 | 0.1324 | No | ||
| 40 | ARF3 | 22137 | 4049 | 0.099 | 0.1319 | No | ||
| 41 | LRFN5 | 6213 | 4305 | 0.084 | 0.1192 | No | ||
| 42 | PHF5A | 22194 | 4338 | 0.082 | 0.1186 | No | ||
| 43 | HOXC13 | 22344 | 4449 | 0.078 | 0.1137 | No | ||
| 44 | SLC2A4 | 20380 | 4496 | 0.076 | 0.1122 | No | ||
| 45 | UQCRC1 | 10259 | 4631 | 0.071 | 0.1059 | No | ||
| 46 | MADD | 2926 14526 | 4688 | 0.069 | 0.1038 | No | ||
| 47 | SLC37A2 | 19178 | 4715 | 0.068 | 0.1033 | No | ||
| 48 | NTF3 | 16991 | 4816 | 0.064 | 0.0987 | No | ||
| 49 | TBC1D15 | 19625 | 4829 | 0.064 | 0.0989 | No | ||
| 50 | CX3CL1 | 9791 | 5126 | 0.055 | 0.0836 | No | ||
| 51 | POLR3D | 21760 12456 | 5271 | 0.052 | 0.0765 | No | ||
| 52 | SDHD | 19125 | 5278 | 0.051 | 0.0768 | No | ||
| 53 | CDKL5 | 24019 | 5330 | 0.050 | 0.0747 | No | ||
| 54 | ZNF289 | 8060 13377 2750 8059 | 5477 | 0.046 | 0.0675 | No | ||
| 55 | HMGB3 | 9096 | 5610 | 0.043 | 0.0609 | No | ||
| 56 | ABHD3 | 23491 | 5612 | 0.043 | 0.0614 | No | ||
| 57 | RIN2 | 14817 | 5626 | 0.043 | 0.0613 | No | ||
| 58 | EPS8L2 | 18012 | 5719 | 0.041 | 0.0568 | No | ||
| 59 | SEMA7A | 19426 9803 | 5804 | 0.040 | 0.0528 | No | ||
| 60 | PCDH7 | 7029 3460 | 5963 | 0.036 | 0.0447 | No | ||
| 61 | CDH16 | 4507 | 6159 | 0.033 | 0.0346 | No | ||
| 62 | HTF9C | 4886 1678 | 6252 | 0.032 | 0.0301 | No | ||
| 63 | PDHX | 14510 | 6490 | 0.028 | 0.0176 | No | ||
| 64 | ATP6V1B1 | 8499 | 6500 | 0.028 | 0.0175 | No | ||
| 65 | PNOC | 21783 | 6748 | 0.024 | 0.0044 | No | ||
| 66 | MEF2D | 9379 | 6789 | 0.023 | 0.0025 | No | ||
| 67 | SEMA3F | 18999 3097 | 6837 | 0.023 | 0.0003 | No | ||
| 68 | CHAT | 21882 | 7211 | 0.017 | -0.0197 | No | ||
| 69 | NDFIP1 | 23573 | 7410 | 0.015 | -0.0302 | No | ||
| 70 | STK32A | 6509 | 7419 | 0.015 | -0.0305 | No | ||
| 71 | HCN1 | 21557 | 8086 | 0.007 | -0.0665 | No | ||
| 72 | SLC30A3 | 5995 | 8291 | 0.004 | -0.0775 | No | ||
| 73 | CACNB2 | 4466 8678 | 8470 | 0.002 | -0.0871 | No | ||
| 74 | PHF15 | 20467 8050 | 8488 | 0.002 | -0.0880 | No | ||
| 75 | CNTN2 | 5627 | 8878 | -0.002 | -0.1091 | No | ||
| 76 | UBE1 | 24370 2551 | 9039 | -0.004 | -0.1177 | No | ||
| 77 | RANBP1 | 9692 5357 | 9289 | -0.008 | -0.1311 | No | ||
| 78 | AUH | 21464 | 9412 | -0.009 | -0.1376 | No | ||
| 79 | NXPH4 | 19602 | 9438 | -0.010 | -0.1388 | No | ||
| 80 | TCF2 | 1395 20727 | 9722 | -0.013 | -0.1540 | No | ||
| 81 | NPTX2 | 16630 | 9769 | -0.014 | -0.1563 | No | ||
| 82 | CASQ2 | 15477 | 9785 | -0.014 | -0.1569 | No | ||
| 83 | RGS7 | 13743 | 9852 | -0.015 | -0.1603 | No | ||
| 84 | PHACTR3 | 7900 2770 | 10093 | -0.018 | -0.1730 | No | ||
| 85 | YWHAH | 5937 10368 | 10101 | -0.018 | -0.1732 | No | ||
| 86 | MASP1 | 1632 22626 | 10115 | -0.018 | -0.1736 | No | ||
| 87 | MYL3 | 19291 | 10129 | -0.019 | -0.1741 | No | ||
| 88 | ATP6V0C | 8643 | 10268 | -0.020 | -0.1813 | No | ||
| 89 | IMMT | 367 8029 | 10295 | -0.021 | -0.1824 | No | ||
| 90 | MNT | 20784 | 10331 | -0.021 | -0.1840 | No | ||
| 91 | XYLT1 | 18113 | 10576 | -0.025 | -0.1969 | No | ||
| 92 | RPH3A | 13628 | 10973 | -0.030 | -0.2180 | No | ||
| 93 | SH3GL2 | 9811 5429 | 10979 | -0.031 | -0.2179 | No | ||
| 94 | NNAT | 2764 5180 | 10997 | -0.031 | -0.2184 | No | ||
| 95 | KCNG4 | 18728 3845 | 11195 | -0.034 | -0.2286 | No | ||
| 96 | SDK1 | 16640 | 11266 | -0.035 | -0.2319 | No | ||
| 97 | FZD9 | 16345 | 11359 | -0.036 | -0.2364 | No | ||
| 98 | KCNMB2 | 1939 15615 | 11364 | -0.037 | -0.2361 | No | ||
| 99 | DSCR1L1 | 23231 | 11421 | -0.038 | -0.2387 | No | ||
| 100 | KIRREL3 | 12571 | 11668 | -0.042 | -0.2515 | No | ||
| 101 | ASTN2 | 15860 2524 | 11992 | -0.049 | -0.2683 | No | ||
| 102 | RHCG | 17788 | 12005 | -0.049 | -0.2683 | No | ||
| 103 | WDFY3 | 7789 3607 16462 | 12114 | -0.052 | -0.2735 | No | ||
| 104 | SLC6A8 | 24306 | 12156 | -0.053 | -0.2750 | No | ||
| 105 | SNTG1 | 7717 | 12445 | -0.060 | -0.2898 | No | ||
| 106 | AMY2A | 8587 | 12692 | -0.068 | -0.3022 | No | ||
| 107 | HCN4 | 9080 | 12825 | -0.072 | -0.3084 | No | ||
| 108 | SLC38A3 | 13319 | 12937 | -0.076 | -0.3134 | No | ||
| 109 | DNER | 13901 | 12953 | -0.076 | -0.3132 | No | ||
| 110 | VSNL1 | 6527 | 13009 | -0.078 | -0.3152 | No | ||
| 111 | AMPD3 | 4383 2110 | 13060 | -0.080 | -0.3168 | No | ||
| 112 | TBX6 | 1993 18075 | 13091 | -0.081 | -0.3173 | No | ||
| 113 | RNF139 | 7992 13291 | 13192 | -0.085 | -0.3216 | No | ||
| 114 | DIRAS1 | 9915 | 13228 | -0.086 | -0.3224 | No | ||
| 115 | GABPA | 8995 883 4744 | 13539 | -0.101 | -0.3378 | No | ||
| 116 | EEF1A2 | 8880 14309 | 13886 | -0.120 | -0.3550 | No | ||
| 117 | TIMM9 | 19686 | 14011 | -0.128 | -0.3600 | No | ||
| 118 | INSR | 18950 | 14023 | -0.129 | -0.3589 | No | ||
| 119 | ELAVL3 | 9136 | 14054 | -0.131 | -0.3587 | No | ||
| 120 | CNTF | 3741 8761 3680 3736 | 14079 | -0.132 | -0.3583 | No | ||
| 121 | NTNG2 | 2731 9330 | 14123 | -0.135 | -0.3588 | No | ||
| 122 | FGF9 | 8966 | 14217 | -0.143 | -0.3619 | No | ||
| 123 | KIF5A | 9222 | 14413 | -0.161 | -0.3703 | No | ||
| 124 | SLC9A7 | 24177 | 14464 | -0.166 | -0.3708 | No | ||
| 125 | SYNGR1 | 22420 2290 | 14672 | -0.188 | -0.3795 | No | ||
| 126 | ATP5C1 | 8635 | 14751 | -0.196 | -0.3811 | No | ||
| 127 | NCDN | 15756 2487 6471 | 14904 | -0.217 | -0.3865 | No | ||
| 128 | NDUFB5 | 12277 | 15191 | -0.254 | -0.3986 | No | ||
| 129 | RNF4 | 5385 9729 | 15271 | -0.268 | -0.3993 | No | ||
| 130 | DEXI | 7193 12212 | 15348 | -0.281 | -0.3996 | Yes | ||
| 131 | TUBB4 | 22917 | 15383 | -0.286 | -0.3977 | Yes | ||
| 132 | CS | 19839 | 15391 | -0.287 | -0.3942 | Yes | ||
| 133 | HOXC10 | 22342 | 15539 | -0.316 | -0.3979 | Yes | ||
| 134 | ACO2 | 8527 | 15620 | -0.335 | -0.3978 | Yes | ||
| 135 | ATP5G3 | 14558 | 15683 | -0.351 | -0.3964 | Yes | ||
| 136 | HSPE1 | 9133 | 15724 | -0.361 | -0.3938 | Yes | ||
| 137 | DNAJC11 | 2486 6068 | 15742 | -0.364 | -0.3898 | Yes | ||
| 138 | LAMB2 | 9265 | 15812 | -0.379 | -0.3885 | Yes | ||
| 139 | RHOT1 | 12227 | 16014 | -0.434 | -0.3936 | Yes | ||
| 140 | FSTL3 | 19961 | 16048 | -0.444 | -0.3895 | Yes | ||
| 141 | VASP | 5847 | 16051 | -0.445 | -0.3836 | Yes | ||
| 142 | GBAS | 16690 | 16053 | -0.445 | -0.3777 | Yes | ||
| 143 | CA2 | 15631 1825 | 16321 | -0.526 | -0.3851 | Yes | ||
| 144 | TOMM40 | 6997 | 16391 | -0.550 | -0.3815 | Yes | ||
| 145 | SNPH | 14395 | 16491 | -0.584 | -0.3791 | Yes | ||
| 146 | MAEA | 16877 | 16557 | -0.614 | -0.3744 | Yes | ||
| 147 | EPN3 | 20291 | 16782 | -0.712 | -0.3770 | Yes | ||
| 148 | CTSD | 22 | 16878 | -0.760 | -0.3720 | Yes | ||
| 149 | COX5B | 4552 | 16899 | -0.772 | -0.3628 | Yes | ||
| 150 | PPP3CC | 21763 | 16969 | -0.809 | -0.3557 | Yes | ||
| 151 | CYC1 | 12348 | 17034 | -0.848 | -0.3478 | Yes | ||
| 152 | MSI2 | 8030 1268 | 17070 | -0.865 | -0.3381 | Yes | ||
| 153 | AFG3L2 | 23416 | 17286 | -0.971 | -0.3368 | Yes | ||
| 154 | ATP5J | 880 22555 | 17364 | -1.001 | -0.3276 | Yes | ||
| 155 | ATP5G1 | 8636 | 17491 | -1.058 | -0.3203 | Yes | ||
| 156 | BLCAP | 14371 | 17569 | -1.106 | -0.3097 | Yes | ||
| 157 | IDH3A | 19447 | 17703 | -1.181 | -0.3011 | Yes | ||
| 158 | VDAC2 | 22081 | 17737 | -1.213 | -0.2866 | Yes | ||
| 159 | ATP1B1 | 4420 | 18129 | -1.539 | -0.2872 | Yes | ||
| 160 | HSPD1 | 4078 9129 | 18140 | -1.557 | -0.2670 | Yes | ||
| 161 | COX4I1 | 18444 | 18198 | -1.613 | -0.2485 | Yes | ||
| 162 | PRPSAP1 | 20141 | 18224 | -1.659 | -0.2277 | Yes | ||
| 163 | TIMM8B | 11391 | 18239 | -1.678 | -0.2060 | Yes | ||
| 164 | GABARAPL1 | 17268 | 18387 | -1.929 | -0.1881 | Yes | ||
| 165 | KCNK1 | 18415 | 18430 | -2.035 | -0.1632 | Yes | ||
| 166 | MTCH2 | 14954 7120 | 18465 | -2.159 | -0.1362 | Yes | ||
| 167 | MGST3 | 13771 12349 | 18471 | -2.184 | -0.1072 | Yes | ||
| 168 | UCHL1 | 16834 | 18526 | -2.404 | -0.0780 | Yes | ||
| 169 | FBXO18 | 14686 2796 | 18566 | -2.650 | -0.0447 | Yes | ||
| 170 | RBMX | 5367 9708 | 18608 | -3.540 | 0.0004 | Yes |

