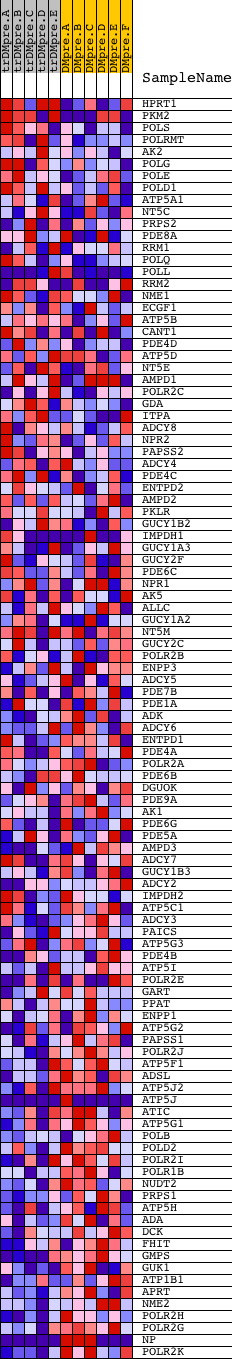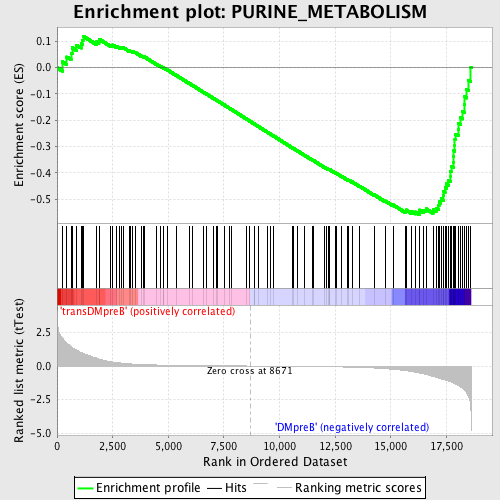
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | PURINE_METABOLISM |
| Enrichment Score (ES) | -0.55888456 |
| Normalized Enrichment Score (NES) | -1.5030318 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.25280085 |
| FWER p-Value | 0.985 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HPRT1 | 2655 24339 408 | 244 | 2.067 | 0.0220 | No | ||
| 2 | PKM2 | 3642 9573 | 439 | 1.709 | 0.0405 | No | ||
| 3 | POLS | 9963 | 629 | 1.445 | 0.0549 | No | ||
| 4 | POLRMT | 19705 | 675 | 1.394 | 0.0761 | No | ||
| 5 | AK2 | 8563 2479 16073 | 873 | 1.189 | 0.0857 | No | ||
| 6 | POLG | 17789 | 1079 | 1.005 | 0.0917 | No | ||
| 7 | POLE | 16755 | 1140 | 0.981 | 0.1052 | No | ||
| 8 | POLD1 | 17847 | 1169 | 0.959 | 0.1199 | No | ||
| 9 | ATP5A1 | 23505 | 1763 | 0.591 | 0.0980 | No | ||
| 10 | NT5C | 20151 | 1883 | 0.516 | 0.1003 | No | ||
| 11 | PRPS2 | 24003 | 1912 | 0.501 | 0.1073 | No | ||
| 12 | PDE8A | 18202 | 2417 | 0.317 | 0.0855 | No | ||
| 13 | RRM1 | 18163 | 2497 | 0.295 | 0.0862 | No | ||
| 14 | POLQ | 13407 22768 | 2673 | 0.255 | 0.0811 | No | ||
| 15 | POLL | 23658 3688 | 2816 | 0.230 | 0.0773 | No | ||
| 16 | RRM2 | 5401 5400 | 2892 | 0.217 | 0.0770 | No | ||
| 17 | NME1 | 9467 | 2961 | 0.206 | 0.0768 | No | ||
| 18 | ECGF1 | 22160 | 3245 | 0.166 | 0.0643 | No | ||
| 19 | ATP5B | 19846 | 3318 | 0.157 | 0.0631 | No | ||
| 20 | CANT1 | 13304 | 3395 | 0.148 | 0.0615 | No | ||
| 21 | PDE4D | 10722 6235 | 3504 | 0.138 | 0.0580 | No | ||
| 22 | ATP5D | 19949 | 3793 | 0.115 | 0.0444 | No | ||
| 23 | NT5E | 19360 18702 | 3904 | 0.108 | 0.0403 | No | ||
| 24 | AMPD1 | 5051 | 3916 | 0.107 | 0.0416 | No | ||
| 25 | POLR2C | 9750 | 4458 | 0.078 | 0.0137 | No | ||
| 26 | GDA | 4764 | 4626 | 0.071 | 0.0058 | No | ||
| 27 | ITPA | 9193 | 4759 | 0.066 | -0.0002 | No | ||
| 28 | ADCY8 | 22281 | 4797 | 0.065 | -0.0010 | No | ||
| 29 | NPR2 | 16230 | 4955 | 0.060 | -0.0085 | No | ||
| 30 | PAPSS2 | 6258 | 5386 | 0.049 | -0.0309 | No | ||
| 31 | ADCY4 | 2980 21818 | 5954 | 0.036 | -0.0609 | No | ||
| 32 | PDE4C | 8481 | 6098 | 0.034 | -0.0681 | No | ||
| 33 | ENTPD2 | 8713 | 6584 | 0.026 | -0.0938 | No | ||
| 34 | AMPD2 | 1931 15197 | 6692 | 0.025 | -0.0992 | No | ||
| 35 | PKLR | 1850 15545 | 7037 | 0.020 | -0.1174 | No | ||
| 36 | GUCY1B2 | 21794 | 7184 | 0.018 | -0.1250 | No | ||
| 37 | IMPDH1 | 17197 1131 | 7187 | 0.018 | -0.1248 | No | ||
| 38 | GUCY1A3 | 7216 12245 15310 | 7505 | 0.013 | -0.1417 | No | ||
| 39 | GUCY2F | 24045 | 7762 | 0.010 | -0.1554 | No | ||
| 40 | PDE6C | 8496 23868 | 7843 | 0.009 | -0.1595 | No | ||
| 41 | NPR1 | 9480 | 8528 | 0.002 | -0.1964 | No | ||
| 42 | AK5 | 6042 | 8635 | 0.000 | -0.2022 | No | ||
| 43 | ALLC | 21093 2130 | 8861 | -0.002 | -0.2143 | No | ||
| 44 | GUCY1A2 | 19580 | 8869 | -0.002 | -0.2146 | No | ||
| 45 | NT5M | 8345 4175 | 9073 | -0.005 | -0.2255 | No | ||
| 46 | GUCY2C | 1140 16954 | 9454 | -0.010 | -0.2459 | No | ||
| 47 | POLR2B | 16817 | 9610 | -0.012 | -0.2540 | No | ||
| 48 | ENPP3 | 19803 | 9728 | -0.013 | -0.2601 | No | ||
| 49 | ADCY5 | 10312 | 10596 | -0.025 | -0.3065 | No | ||
| 50 | PDE7B | 19807 | 10626 | -0.026 | -0.3077 | No | ||
| 51 | PDE1A | 9541 5234 | 10813 | -0.028 | -0.3172 | No | ||
| 52 | ADK | 3302 3454 8555 | 11121 | -0.033 | -0.3333 | No | ||
| 53 | ADCY6 | 22139 2283 8551 | 11490 | -0.039 | -0.3525 | No | ||
| 54 | ENTPD1 | 4495 | 11525 | -0.039 | -0.3537 | No | ||
| 55 | PDE4A | 3037 19542 3083 3023 | 11997 | -0.049 | -0.3783 | No | ||
| 56 | POLR2A | 5394 | 12113 | -0.052 | -0.3836 | No | ||
| 57 | PDE6B | 16760 | 12210 | -0.054 | -0.3879 | No | ||
| 58 | DGUOK | 17099 1041 | 12229 | -0.055 | -0.3879 | No | ||
| 59 | PDE9A | 23301 | 12232 | -0.055 | -0.3871 | No | ||
| 60 | AK1 | 4363 | 12500 | -0.061 | -0.4005 | No | ||
| 61 | PDE6G | 20119 | 12577 | -0.064 | -0.4035 | No | ||
| 62 | PDE5A | 15431 | 12802 | -0.071 | -0.4144 | No | ||
| 63 | AMPD3 | 4383 2110 | 13060 | -0.080 | -0.4269 | No | ||
| 64 | ADCY7 | 8552 448 | 13118 | -0.082 | -0.4286 | No | ||
| 65 | GUCY1B3 | 15311 | 13256 | -0.087 | -0.4345 | No | ||
| 66 | ADCY2 | 21412 | 13602 | -0.104 | -0.4514 | No | ||
| 67 | IMPDH2 | 10730 | 14256 | -0.146 | -0.4841 | No | ||
| 68 | ATP5C1 | 8635 | 14751 | -0.196 | -0.5075 | No | ||
| 69 | ADCY3 | 21329 894 | 15106 | -0.241 | -0.5225 | No | ||
| 70 | PAICS | 16820 | 15673 | -0.349 | -0.5472 | No | ||
| 71 | ATP5G3 | 14558 | 15683 | -0.351 | -0.5417 | No | ||
| 72 | PDE4B | 9543 | 15910 | -0.406 | -0.5470 | No | ||
| 73 | ATP5I | 8637 | 16098 | -0.457 | -0.5493 | No | ||
| 74 | POLR2E | 3325 19699 | 16276 | -0.508 | -0.5502 | Yes | ||
| 75 | GART | 22543 1754 | 16281 | -0.510 | -0.5418 | Yes | ||
| 76 | PPAT | 6081 | 16471 | -0.577 | -0.5422 | Yes | ||
| 77 | ENPP1 | 19804 | 16584 | -0.626 | -0.5376 | Yes | ||
| 78 | ATP5G2 | 12610 | 16897 | -0.770 | -0.5414 | Yes | ||
| 79 | PAPSS1 | 15419 | 17035 | -0.850 | -0.5343 | Yes | ||
| 80 | POLR2J | 16672 | 17132 | -0.889 | -0.5244 | Yes | ||
| 81 | ATP5F1 | 15212 | 17170 | -0.904 | -0.5110 | Yes | ||
| 82 | ADSL | 4358 | 17255 | -0.952 | -0.4994 | Yes | ||
| 83 | ATP5J2 | 12186 | 17355 | -0.999 | -0.4877 | Yes | ||
| 84 | ATP5J | 880 22555 | 17364 | -1.001 | -0.4712 | Yes | ||
| 85 | ATIC | 14231 3968 | 17439 | -1.030 | -0.4577 | Yes | ||
| 86 | ATP5G1 | 8636 | 17491 | -1.058 | -0.4424 | Yes | ||
| 87 | POLB | 9599 | 17589 | -1.119 | -0.4286 | Yes | ||
| 88 | POLD2 | 20537 | 17687 | -1.174 | -0.4139 | Yes | ||
| 89 | POLR2I | 12839 | 17691 | -1.175 | -0.3941 | Yes | ||
| 90 | POLR1B | 14857 | 17713 | -1.191 | -0.3750 | Yes | ||
| 91 | NUDT2 | 16239 | 17821 | -1.266 | -0.3592 | Yes | ||
| 92 | PRPS1 | 24233 | 17827 | -1.272 | -0.3379 | Yes | ||
| 93 | ATP5H | 12948 | 17834 | -1.274 | -0.3166 | Yes | ||
| 94 | ADA | 2703 14361 | 17876 | -1.307 | -0.2966 | Yes | ||
| 95 | DCK | 16808 | 17881 | -1.308 | -0.2745 | Yes | ||
| 96 | FHIT | 21920 | 17890 | -1.314 | -0.2527 | Yes | ||
| 97 | GMPS | 15578 | 18028 | -1.431 | -0.2357 | Yes | ||
| 98 | GUK1 | 20432 | 18041 | -1.442 | -0.2119 | Yes | ||
| 99 | ATP1B1 | 4420 | 18129 | -1.539 | -0.1904 | Yes | ||
| 100 | APRT | 8620 | 18225 | -1.660 | -0.1673 | Yes | ||
| 101 | NME2 | 9468 | 18309 | -1.791 | -0.1414 | Yes | ||
| 102 | POLR2H | 10888 | 18321 | -1.818 | -0.1111 | Yes | ||
| 103 | POLR2G | 23753 | 18407 | -1.978 | -0.0820 | Yes | ||
| 104 | NP | 22027 9597 5273 5274 | 18493 | -2.234 | -0.0487 | Yes | ||
| 105 | POLR2K | 9413 | 18599 | -3.250 | 0.0009 | Yes |