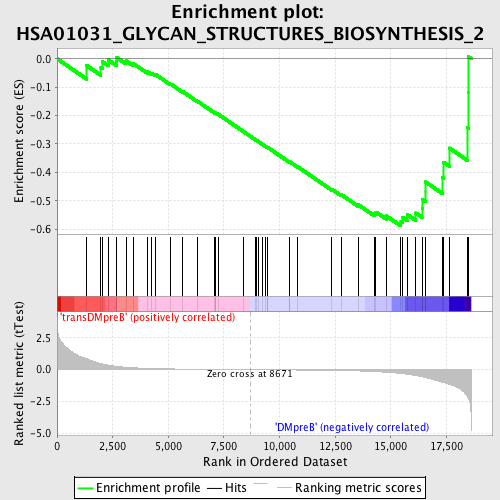
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | HSA01031_GLYCAN_STRUCTURES_BIOSYNTHESIS_2 |
| Enrichment Score (ES) | -0.58869773 |
| Normalized Enrichment Score (NES) | -1.4425651 |
| Nominal p-value | 0.02661597 |
| FDR q-value | 0.33128983 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PIGQ | 4798 4797 | 1328 | 0.846 | -0.0237 | No | ||
| 2 | B3GALT4 | 23024 | 1970 | 0.471 | -0.0316 | No | ||
| 3 | B4GALT6 | 7109 12115 | 2031 | 0.446 | -0.0096 | No | ||
| 4 | GBGT1 | 15067 | 2296 | 0.356 | -0.0037 | No | ||
| 5 | ST6GALNAC6 | 11950 2856 | 2664 | 0.257 | -0.0089 | No | ||
| 6 | PIGB | 3128 12067 | 2685 | 0.253 | 0.0043 | No | ||
| 7 | PIGA | 2592 24210 | 3096 | 0.187 | -0.0072 | No | ||
| 8 | PIGM | 14046 | 3414 | 0.146 | -0.0161 | No | ||
| 9 | PIGH | 8483 | 4070 | 0.097 | -0.0458 | No | ||
| 10 | GCNT2 | 9168 3166 3243 21661 3270 | 4241 | 0.087 | -0.0501 | No | ||
| 11 | ST3GAL1 | 5435 2227 | 4406 | 0.080 | -0.0544 | No | ||
| 12 | PIGC | 14081 | 5109 | 0.055 | -0.0890 | No | ||
| 13 | B4GALT4 | 22756 | 5648 | 0.043 | -0.1156 | No | ||
| 14 | B3GALT1 | 6497 | 6311 | 0.031 | -0.1495 | No | ||
| 15 | ABO | 14647 | 7089 | 0.019 | -0.1903 | No | ||
| 16 | B3GALT2 | 6498 | 7134 | 0.018 | -0.1916 | No | ||
| 17 | FUT4 | 19240 | 7135 | 0.018 | -0.1906 | No | ||
| 18 | ST3GAL4 | 9816 | 7267 | 0.017 | -0.1967 | No | ||
| 19 | FUT1 | 8987 | 8397 | 0.003 | -0.2573 | No | ||
| 20 | B3GALNT1 | 15319 18224 | 8919 | -0.003 | -0.2852 | No | ||
| 21 | ST3GAL3 | 15782 | 8947 | -0.003 | -0.2865 | No | ||
| 22 | FUT7 | 15084 | 9035 | -0.004 | -0.2910 | No | ||
| 23 | ST6GALNAC5 | 15136 | 9230 | -0.007 | -0.3010 | No | ||
| 24 | B3GNT1 | 11997 | 9372 | -0.009 | -0.3082 | No | ||
| 25 | ST8SIA5 | 1950 1944 23508 | 9441 | -0.010 | -0.3113 | No | ||
| 26 | ST8SIA1 | 5437 | 9444 | -0.010 | -0.3108 | No | ||
| 27 | FUT9 | 16257 | 10443 | -0.023 | -0.3633 | No | ||
| 28 | B4GALT2 | 11990 | 10456 | -0.023 | -0.3626 | No | ||
| 29 | B3GNT4 | 10542 | 10820 | -0.028 | -0.3806 | No | ||
| 30 | B3GALT5 | 22690 | 12352 | -0.057 | -0.4598 | No | ||
| 31 | ST6GALNAC4 | 15036 2702 2848 2937 | 12770 | -0.070 | -0.4783 | No | ||
| 32 | PIGN | 13865 | 13530 | -0.100 | -0.5135 | No | ||
| 33 | PIGS | 11347 | 14265 | -0.147 | -0.5447 | No | ||
| 34 | ST6GALNAC3 | 2 | 14330 | -0.152 | -0.5396 | No | ||
| 35 | FUT2 | 17829 | 14796 | -0.201 | -0.5533 | No | ||
| 36 | ST3GAL5 | 17415 | 15455 | -0.300 | -0.5718 | Yes | ||
| 37 | PIGK | 1815 6834 11629 | 15541 | -0.317 | -0.5584 | Yes | ||
| 38 | PIGO | 15903 2488 2344 | 15732 | -0.362 | -0.5482 | Yes | ||
| 39 | UGCGL1 | 6721 | 16121 | -0.463 | -0.5429 | Yes | ||
| 40 | B3GNT3 | 18842 | 16404 | -0.555 | -0.5268 | Yes | ||
| 41 | B4GALT3 | 14057 | 16437 | -0.563 | -0.4967 | Yes | ||
| 42 | UGCG | 16197 | 16559 | -0.616 | -0.4684 | Yes | ||
| 43 | PIGL | 11595 | 16560 | -0.616 | -0.4335 | Yes | ||
| 44 | B3GNT5 | 22824 | 17330 | -0.993 | -0.4189 | Yes | ||
| 45 | PIGT | 8134 1 | 17360 | -1.000 | -0.3639 | Yes | ||
| 46 | ST3GAL6 | 7046 | 17631 | -1.139 | -0.3141 | Yes | ||
| 47 | PIGF | 22879 | 18432 | -2.041 | -0.2418 | Yes | ||
| 48 | B4GALT1 | 15918 | 18484 | -2.224 | -0.1188 | Yes | ||
| 49 | PIGP | 12082 1665 | 18488 | -2.227 | 0.0069 | Yes |

