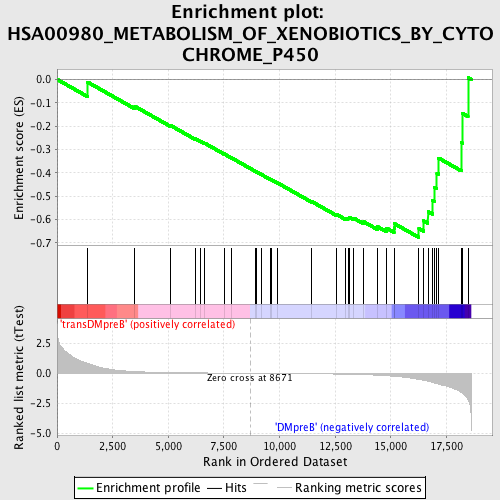
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | HSA00980_METABOLISM_OF_XENOBIOTICS_BY_CYTOCHROME_P450 |
| Enrichment Score (ES) | -0.67661166 |
| Normalized Enrichment Score (NES) | -1.550397 |
| Nominal p-value | 0.009090909 |
| FDR q-value | 0.23874205 |
| FWER p-Value | 0.898 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | UGT1A10 | 6908 | 1378 | 0.814 | -0.0119 | No | ||
| 2 | UGT2A1 | 8247 | 3495 | 0.138 | -0.1152 | No | ||
| 3 | GSTA2 | 4808 6173 | 5087 | 0.056 | -0.1965 | No | ||
| 4 | ALDH3A1 | 20854 | 6209 | 0.032 | -0.2544 | No | ||
| 5 | CYP1B1 | 4581 | 6434 | 0.029 | -0.2643 | No | ||
| 6 | ADH4 | 15405 | 6640 | 0.025 | -0.2733 | No | ||
| 7 | GSTM2 | 9049 4810 | 7504 | 0.013 | -0.3187 | No | ||
| 8 | GSTA1 | 4807 | 7853 | 0.009 | -0.3368 | No | ||
| 9 | EPHX1 | 13734 | 8898 | -0.003 | -0.3928 | No | ||
| 10 | GSTA3 | 14287 | 8943 | -0.003 | -0.3949 | No | ||
| 11 | UGT1A1 | 11851 | 9205 | -0.006 | -0.4084 | No | ||
| 12 | CYP2E1 | 18025 | 9585 | -0.011 | -0.4280 | No | ||
| 13 | ADH7 | 15408 | 9628 | -0.012 | -0.4293 | No | ||
| 14 | ALDH1A3 | 17802 | 9918 | -0.016 | -0.4437 | No | ||
| 15 | CYP1A1 | 19428 | 11450 | -0.038 | -0.5231 | No | ||
| 16 | ADH1A | 15406 | 12545 | -0.063 | -0.5772 | No | ||
| 17 | CYP1A2 | 19097 | 12978 | -0.077 | -0.5945 | No | ||
| 18 | GSTM3 | 9050 | 13076 | -0.081 | -0.5936 | No | ||
| 19 | UGT1A6 | 3969 4079 6911 13591 | 13131 | -0.083 | -0.5902 | No | ||
| 20 | GSTM1 | 4809 9048 | 13320 | -0.090 | -0.5934 | No | ||
| 21 | ADHFE1 | 4049 8008 | 13789 | -0.114 | -0.6098 | No | ||
| 22 | UGT1A9 | 11849 11850 | 14404 | -0.161 | -0.6306 | No | ||
| 23 | ALDH3B1 | 12569 23949 | 14816 | -0.204 | -0.6371 | No | ||
| 24 | CYP2S1 | 17921 | 15146 | -0.247 | -0.6359 | No | ||
| 25 | MGST2 | 15597 | 15169 | -0.251 | -0.6178 | No | ||
| 26 | ADH5 | 8554 15404 | 16262 | -0.501 | -0.6383 | Yes | ||
| 27 | GSTP1 | 9051 4812 4811 | 16478 | -0.579 | -0.6055 | Yes | ||
| 28 | GSTA4 | 19371 | 16671 | -0.658 | -0.5656 | Yes | ||
| 29 | GSTZ1 | 21193 | 16879 | -0.761 | -0.5185 | Yes | ||
| 30 | GSTM4 | 15199 | 16952 | -0.801 | -0.4612 | Yes | ||
| 31 | GSTT1 | 19730 | 17072 | -0.866 | -0.4014 | Yes | ||
| 32 | GSTM5 | 15455 | 17148 | -0.898 | -0.3368 | Yes | ||
| 33 | GSTT2 | 9052 | 18171 | -1.584 | -0.2706 | Yes | ||
| 34 | MGST1 | 17254 | 18229 | -1.667 | -0.1462 | Yes | ||
| 35 | MGST3 | 13771 12349 | 18471 | -2.184 | 0.0078 | Yes |

