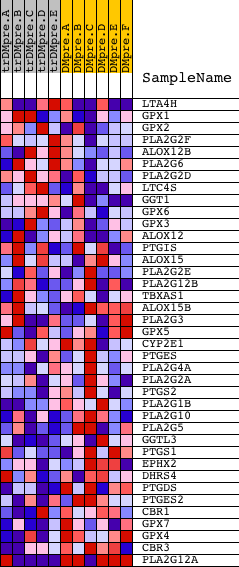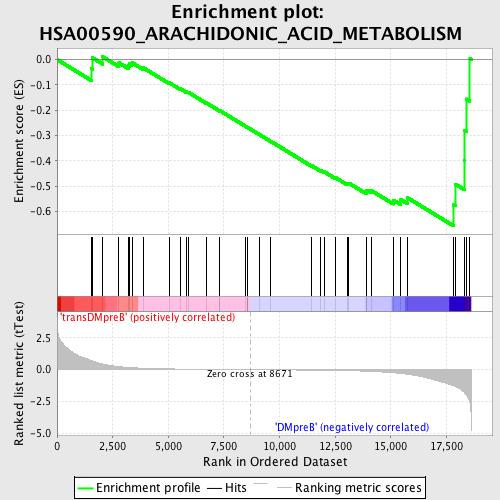
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | HSA00590_ARACHIDONIC_ACID_METABOLISM |
| Enrichment Score (ES) | -0.658011 |
| Normalized Enrichment Score (NES) | -1.5203166 |
| Nominal p-value | 0.016014235 |
| FDR q-value | 0.27644366 |
| FWER p-Value | 0.971 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LTA4H | 19906 3374 | 1534 | 0.719 | -0.0355 | No | ||
| 2 | GPX1 | 19310 | 1580 | 0.693 | 0.0075 | No | ||
| 3 | GPX2 | 9037 | 2054 | 0.438 | 0.0107 | No | ||
| 4 | PLA2G2F | 15700 | 2780 | 0.236 | -0.0128 | No | ||
| 5 | ALOX12B | 20825 | 3209 | 0.171 | -0.0247 | No | ||
| 6 | PLA2G6 | 22209 | 3248 | 0.165 | -0.0160 | No | ||
| 7 | PLA2G2D | 16018 | 3366 | 0.151 | -0.0124 | No | ||
| 8 | LTC4S | 20478 | 3882 | 0.109 | -0.0330 | No | ||
| 9 | GGT1 | 19987 | 5034 | 0.058 | -0.0912 | No | ||
| 10 | GPX6 | 21698 | 5549 | 0.045 | -0.1159 | No | ||
| 11 | GPX3 | 20880 | 5793 | 0.040 | -0.1264 | No | ||
| 12 | ALOX12 | 20372 | 5889 | 0.038 | -0.1290 | No | ||
| 13 | PTGIS | 9653 | 6708 | 0.024 | -0.1714 | No | ||
| 14 | ALOX15 | 20370 | 7315 | 0.016 | -0.2030 | No | ||
| 15 | PLA2G2E | 16016 | 7318 | 0.016 | -0.2021 | No | ||
| 16 | PLA2G12B | 20016 | 8454 | 0.003 | -0.2630 | No | ||
| 17 | TBXAS1 | 17480 1105 | 8565 | 0.001 | -0.2688 | No | ||
| 18 | ALOX15B | 20397 | 8571 | 0.001 | -0.2690 | No | ||
| 19 | PLA2G3 | 20968 | 8577 | 0.001 | -0.2692 | No | ||
| 20 | GPX5 | 21532 3242 | 9106 | -0.005 | -0.2973 | No | ||
| 21 | CYP2E1 | 18025 | 9585 | -0.011 | -0.3223 | No | ||
| 22 | PTGES | 7224 | 11445 | -0.038 | -0.4199 | No | ||
| 23 | PLA2G4A | 13809 | 11835 | -0.046 | -0.4378 | No | ||
| 24 | PLA2G2A | 16017 | 12015 | -0.049 | -0.4443 | No | ||
| 25 | PTGS2 | 5317 9655 | 12518 | -0.062 | -0.4672 | No | ||
| 26 | PLA2G1B | 9581 | 13039 | -0.079 | -0.4900 | No | ||
| 27 | PLA2G10 | 22658 | 13105 | -0.082 | -0.4882 | No | ||
| 28 | PLA2G5 | 9583 | 13892 | -0.120 | -0.5226 | No | ||
| 29 | GGTL3 | 14381 | 13924 | -0.122 | -0.5163 | No | ||
| 30 | PTGS1 | 2726 5316 15028 | 14120 | -0.135 | -0.5179 | No | ||
| 31 | EPHX2 | 21778 | 15111 | -0.242 | -0.5554 | No | ||
| 32 | DHRS4 | 6609 | 15452 | -0.299 | -0.5541 | No | ||
| 33 | PTGDS | 14658 | 15733 | -0.362 | -0.5455 | No | ||
| 34 | PTGES2 | 15038 | 17825 | -1.270 | -0.5748 | Yes | ||
| 35 | CBR1 | 22697 | 17902 | -1.322 | -0.4923 | Yes | ||
| 36 | GPX7 | 15809 | 18296 | -1.769 | -0.3977 | Yes | ||
| 37 | GPX4 | 9038 | 18324 | -1.829 | -0.2794 | Yes | ||
| 38 | CBR3 | 22696 | 18393 | -1.941 | -0.1559 | Yes | ||
| 39 | PLA2G12A | 1782 7281 1839 | 18556 | -2.564 | 0.0032 | Yes |