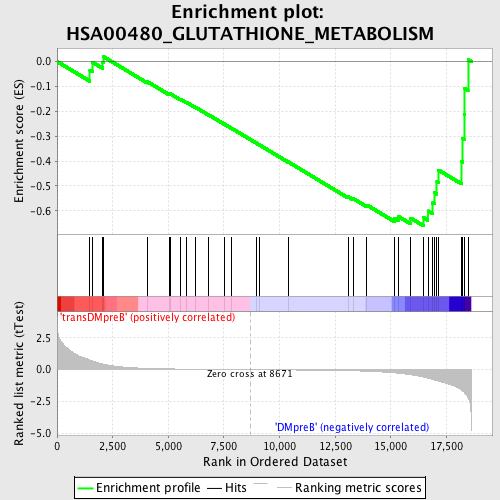
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | HSA00480_GLUTATHIONE_METABOLISM |
| Enrichment Score (ES) | -0.66100067 |
| Normalized Enrichment Score (NES) | -1.504326 |
| Nominal p-value | 0.016885553 |
| FDR q-value | 0.27398005 |
| FWER p-Value | 0.985 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IDH2 | 17781 | 1475 | 0.755 | -0.0363 | No | ||
| 2 | GPX1 | 19310 | 1580 | 0.693 | -0.0024 | No | ||
| 3 | GPX2 | 9037 | 2054 | 0.438 | -0.0028 | No | ||
| 4 | GCLC | 19374 | 2099 | 0.420 | 0.0188 | No | ||
| 5 | OPLAH | 22246 | 4053 | 0.099 | -0.0807 | No | ||
| 6 | GGT1 | 19987 | 5034 | 0.058 | -0.1301 | No | ||
| 7 | GSTA2 | 4808 6173 | 5087 | 0.056 | -0.1297 | No | ||
| 8 | GPX6 | 21698 | 5549 | 0.045 | -0.1520 | No | ||
| 9 | GPX3 | 20880 | 5793 | 0.040 | -0.1628 | No | ||
| 10 | GSS | 14380 9047 | 6204 | 0.032 | -0.1830 | No | ||
| 11 | GCLM | 9019 | 6802 | 0.023 | -0.2138 | No | ||
| 12 | GSTM2 | 9049 4810 | 7504 | 0.013 | -0.2508 | No | ||
| 13 | GSTA1 | 4807 | 7853 | 0.009 | -0.2690 | No | ||
| 14 | GSTA3 | 14287 | 8943 | -0.003 | -0.3274 | No | ||
| 15 | GPX5 | 21532 3242 | 9106 | -0.005 | -0.3358 | No | ||
| 16 | ANPEP | 17784 | 10389 | -0.022 | -0.4035 | No | ||
| 17 | GSTM3 | 9050 | 13076 | -0.081 | -0.5435 | No | ||
| 18 | GSTM1 | 4809 9048 | 13320 | -0.090 | -0.5514 | No | ||
| 19 | GGTL3 | 14381 | 13924 | -0.122 | -0.5769 | No | ||
| 20 | MGST2 | 15597 | 15169 | -0.251 | -0.6295 | No | ||
| 21 | GSR | 18634 | 15328 | -0.279 | -0.6221 | No | ||
| 22 | IDH1 | 4894 | 15895 | -0.401 | -0.6297 | Yes | ||
| 23 | GSTP1 | 9051 4812 4811 | 16478 | -0.579 | -0.6280 | Yes | ||
| 24 | GSTA4 | 19371 | 16671 | -0.658 | -0.6008 | Yes | ||
| 25 | GSTZ1 | 21193 | 16879 | -0.761 | -0.5685 | Yes | ||
| 26 | GSTM4 | 15199 | 16952 | -0.801 | -0.5267 | Yes | ||
| 27 | GSTT1 | 19730 | 17072 | -0.866 | -0.4837 | Yes | ||
| 28 | GSTM5 | 15455 | 17148 | -0.898 | -0.4365 | Yes | ||
| 29 | GSTT2 | 9052 | 18171 | -1.584 | -0.4011 | Yes | ||
| 30 | MGST1 | 17254 | 18229 | -1.667 | -0.3091 | Yes | ||
| 31 | GPX7 | 15809 | 18296 | -1.769 | -0.2118 | Yes | ||
| 32 | GPX4 | 9038 | 18324 | -1.829 | -0.1089 | Yes | ||
| 33 | MGST3 | 13771 12349 | 18471 | -2.184 | 0.0078 | Yes |

