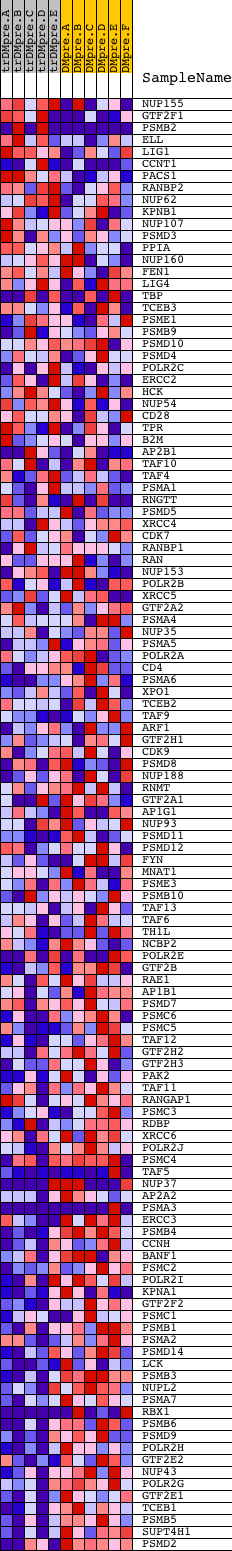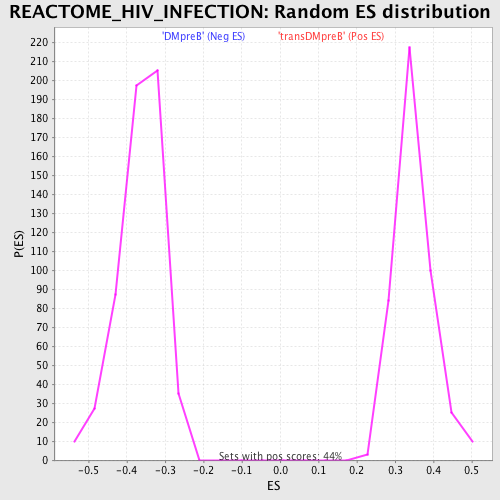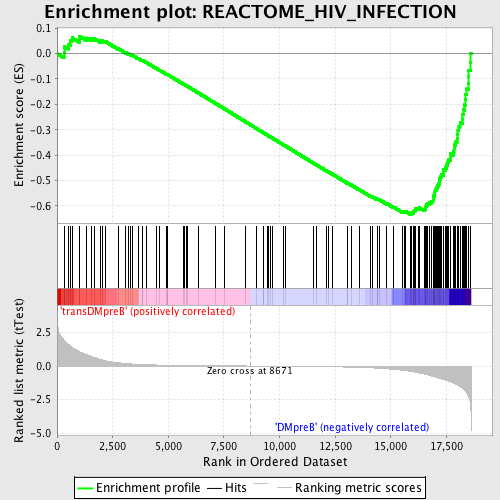
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | REACTOME_HIV_INFECTION |
| Enrichment Score (ES) | -0.6340271 |
| Normalized Enrichment Score (NES) | -1.7500781 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.007836382 |
| FWER p-Value | 0.073 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NUP155 | 2298 5027 | 311 | 1.904 | 0.0056 | No | ||
| 2 | GTF2F1 | 13599 | 318 | 1.883 | 0.0275 | No | ||
| 3 | PSMB2 | 2324 16078 | 494 | 1.634 | 0.0373 | No | ||
| 4 | ELL | 18586 | 584 | 1.520 | 0.0504 | No | ||
| 5 | LIG1 | 18388 1749 1493 | 670 | 1.402 | 0.0624 | No | ||
| 6 | CCNT1 | 22140 11607 | 1002 | 1.050 | 0.0569 | No | ||
| 7 | PACS1 | 23975 | 1008 | 1.045 | 0.0689 | No | ||
| 8 | RANBP2 | 20019 | 1330 | 0.846 | 0.0615 | No | ||
| 9 | NUP62 | 9497 | 1532 | 0.721 | 0.0592 | No | ||
| 10 | KPNB1 | 20274 | 1674 | 0.643 | 0.0591 | No | ||
| 11 | NUP107 | 8337 | 1954 | 0.481 | 0.0497 | No | ||
| 12 | PSMD3 | 5803 | 2030 | 0.446 | 0.0509 | No | ||
| 13 | PPIA | 1284 11188 | 2164 | 0.393 | 0.0484 | No | ||
| 14 | NUP160 | 14957 | 2748 | 0.241 | 0.0197 | No | ||
| 15 | FEN1 | 8961 | 3090 | 0.188 | 0.0035 | No | ||
| 16 | LIG4 | 11454 | 3210 | 0.171 | -0.0010 | No | ||
| 17 | TBP | 671 1554 | 3303 | 0.158 | -0.0041 | No | ||
| 18 | TCEB3 | 15707 | 3375 | 0.150 | -0.0061 | No | ||
| 19 | PSME1 | 5300 9637 | 3639 | 0.127 | -0.0189 | No | ||
| 20 | PSMB9 | 23021 | 3824 | 0.113 | -0.0275 | No | ||
| 21 | PSMD10 | 2578 2597 24046 | 3855 | 0.111 | -0.0278 | No | ||
| 22 | PSMD4 | 15251 | 4024 | 0.100 | -0.0357 | No | ||
| 23 | POLR2C | 9750 | 4458 | 0.078 | -0.0582 | No | ||
| 24 | ERCC2 | 1549 18363 3812 | 4596 | 0.072 | -0.0647 | No | ||
| 25 | HCK | 14787 | 4919 | 0.061 | -0.0814 | No | ||
| 26 | NUP54 | 11231 11232 6516 | 4961 | 0.060 | -0.0829 | No | ||
| 27 | CD28 | 14239 4092 | 4981 | 0.059 | -0.0833 | No | ||
| 28 | TPR | 927 4255 | 5663 | 0.042 | -0.1196 | No | ||
| 29 | B2M | 72 14882 | 5712 | 0.042 | -0.1217 | No | ||
| 30 | AP2B1 | 12958 7752 12957 1280 | 5802 | 0.040 | -0.1260 | No | ||
| 31 | TAF10 | 10785 | 5865 | 0.038 | -0.1289 | No | ||
| 32 | TAF4 | 14319 | 6357 | 0.030 | -0.1551 | No | ||
| 33 | PSMA1 | 1627 17669 | 7105 | 0.019 | -0.1953 | No | ||
| 34 | RNGTT | 10769 2354 6272 | 7529 | 0.013 | -0.2180 | No | ||
| 35 | PSMD5 | 2774 14612 | 8474 | 0.002 | -0.2690 | No | ||
| 36 | XRCC4 | 4238 8432 | 8973 | -0.003 | -0.2959 | No | ||
| 37 | CDK7 | 21365 | 9267 | -0.007 | -0.3117 | No | ||
| 38 | RANBP1 | 9692 5357 | 9289 | -0.008 | -0.3127 | No | ||
| 39 | RAN | 5356 9691 | 9465 | -0.010 | -0.3220 | No | ||
| 40 | NUP153 | 21474 | 9500 | -0.010 | -0.3238 | No | ||
| 41 | POLR2B | 16817 | 9610 | -0.012 | -0.3295 | No | ||
| 42 | XRCC5 | 14229 | 9669 | -0.013 | -0.3325 | No | ||
| 43 | GTF2A2 | 10654 | 10157 | -0.019 | -0.3586 | No | ||
| 44 | PSMA4 | 11179 | 10263 | -0.020 | -0.3640 | No | ||
| 45 | NUP35 | 12803 | 11524 | -0.039 | -0.4317 | No | ||
| 46 | PSMA5 | 6464 | 11660 | -0.042 | -0.4385 | No | ||
| 47 | POLR2A | 5394 | 12113 | -0.052 | -0.4623 | No | ||
| 48 | CD4 | 16999 | 12204 | -0.054 | -0.4666 | No | ||
| 49 | PSMA6 | 21270 | 12356 | -0.058 | -0.4741 | No | ||
| 50 | XPO1 | 4172 | 13031 | -0.079 | -0.5096 | No | ||
| 51 | TCEB2 | 12563 23102 7476 | 13069 | -0.080 | -0.5106 | No | ||
| 52 | TAF9 | 3213 8433 | 13209 | -0.086 | -0.5171 | No | ||
| 53 | ARF1 | 64 | 13575 | -0.103 | -0.5356 | No | ||
| 54 | GTF2H1 | 4069 18236 | 14080 | -0.132 | -0.5613 | No | ||
| 55 | CDK9 | 14619 | 14177 | -0.139 | -0.5649 | No | ||
| 56 | PSMD8 | 7166 | 14395 | -0.160 | -0.5747 | No | ||
| 57 | NUP188 | 15053 | 14398 | -0.160 | -0.5729 | No | ||
| 58 | RNMT | 7501 | 14503 | -0.169 | -0.5766 | No | ||
| 59 | GTF2A1 | 2064 8187 8188 13512 | 14827 | -0.206 | -0.5916 | No | ||
| 60 | AP1G1 | 4392 | 15128 | -0.245 | -0.6049 | No | ||
| 61 | NUP93 | 7762 | 15520 | -0.314 | -0.6224 | No | ||
| 62 | PSMD11 | 12772 7600 | 15592 | -0.330 | -0.6223 | No | ||
| 63 | PSMD12 | 20621 | 15664 | -0.347 | -0.6221 | No | ||
| 64 | FYN | 3375 3395 20052 | 15886 | -0.398 | -0.6293 | Yes | ||
| 65 | MNAT1 | 9396 2161 | 15949 | -0.416 | -0.6278 | Yes | ||
| 66 | PSME3 | 20657 | 16006 | -0.433 | -0.6257 | Yes | ||
| 67 | PSMB10 | 5299 18761 | 16020 | -0.436 | -0.6213 | Yes | ||
| 68 | TAF13 | 8288 | 16047 | -0.444 | -0.6174 | Yes | ||
| 69 | TAF6 | 16322 891 | 16105 | -0.458 | -0.6151 | Yes | ||
| 70 | TH1L | 14715 | 16116 | -0.462 | -0.6102 | Yes | ||
| 71 | NCBP2 | 12643 | 16223 | -0.492 | -0.6101 | Yes | ||
| 72 | POLR2E | 3325 19699 | 16276 | -0.508 | -0.6070 | Yes | ||
| 73 | GTF2B | 10489 | 16522 | -0.597 | -0.6132 | Yes | ||
| 74 | RAE1 | 12395 | 16554 | -0.613 | -0.6076 | Yes | ||
| 75 | AP1B1 | 8599 | 16576 | -0.624 | -0.6014 | Yes | ||
| 76 | PSMD7 | 18752 3825 | 16614 | -0.638 | -0.5959 | Yes | ||
| 77 | PSMC6 | 7379 | 16655 | -0.652 | -0.5904 | Yes | ||
| 78 | PSMC5 | 1348 20624 | 16737 | -0.692 | -0.5866 | Yes | ||
| 79 | TAF12 | 2537 16058 2338 | 16807 | -0.723 | -0.5818 | Yes | ||
| 80 | GTF2H2 | 6236 | 16895 | -0.768 | -0.5774 | Yes | ||
| 81 | GTF2H3 | 5551 | 16901 | -0.772 | -0.5686 | Yes | ||
| 82 | PAK2 | 10310 5875 | 16924 | -0.783 | -0.5605 | Yes | ||
| 83 | TAF11 | 12733 | 16966 | -0.807 | -0.5532 | Yes | ||
| 84 | RANGAP1 | 2180 22195 | 16983 | -0.817 | -0.5445 | Yes | ||
| 85 | PSMC3 | 9636 | 16989 | -0.820 | -0.5351 | Yes | ||
| 86 | RDBP | 6589 | 17040 | -0.853 | -0.5277 | Yes | ||
| 87 | XRCC6 | 2286 8991 | 17103 | -0.878 | -0.5207 | Yes | ||
| 88 | POLR2J | 16672 | 17132 | -0.889 | -0.5117 | Yes | ||
| 89 | PSMC4 | 2141 17915 | 17184 | -0.914 | -0.5037 | Yes | ||
| 90 | TAF5 | 23833 5934 3759 | 17191 | -0.916 | -0.4932 | Yes | ||
| 91 | NUP37 | 3294 3326 19909 | 17218 | -0.932 | -0.4836 | Yes | ||
| 92 | AP2A2 | 18009 | 17274 | -0.963 | -0.4753 | Yes | ||
| 93 | PSMA3 | 9632 5298 | 17363 | -1.000 | -0.4682 | Yes | ||
| 94 | ERCC3 | 23605 | 17381 | -1.003 | -0.4573 | Yes | ||
| 95 | PSMB4 | 15252 | 17460 | -1.042 | -0.4492 | Yes | ||
| 96 | CCNH | 7322 | 17501 | -1.064 | -0.4388 | Yes | ||
| 97 | BANF1 | 6216 10706 3760 | 17547 | -1.087 | -0.4285 | Yes | ||
| 98 | PSMC2 | 16909 | 17578 | -1.113 | -0.4170 | Yes | ||
| 99 | POLR2I | 12839 | 17691 | -1.175 | -0.4092 | Yes | ||
| 100 | KPNA1 | 22776 | 17694 | -1.176 | -0.3954 | Yes | ||
| 101 | GTF2F2 | 21750 | 17807 | -1.258 | -0.3866 | Yes | ||
| 102 | PSMC1 | 9635 | 17840 | -1.277 | -0.3733 | Yes | ||
| 103 | PSMB1 | 23118 | 17872 | -1.303 | -0.3596 | Yes | ||
| 104 | PSMA2 | 9631 | 17925 | -1.349 | -0.3465 | Yes | ||
| 105 | PSMD14 | 15005 | 17976 | -1.387 | -0.3329 | Yes | ||
| 106 | LCK | 15746 | 17991 | -1.398 | -0.3171 | Yes | ||
| 107 | PSMB3 | 11180 | 18016 | -1.424 | -0.3016 | Yes | ||
| 108 | NUPL2 | 6072 | 18061 | -1.465 | -0.2867 | Yes | ||
| 109 | PSMA7 | 14318 | 18110 | -1.516 | -0.2714 | Yes | ||
| 110 | RBX1 | 12130 7123 | 18213 | -1.640 | -0.2576 | Yes | ||
| 111 | PSMB6 | 9634 | 18218 | -1.647 | -0.2384 | Yes | ||
| 112 | PSMD9 | 3461 7393 | 18267 | -1.723 | -0.2207 | Yes | ||
| 113 | POLR2H | 10888 | 18321 | -1.818 | -0.2021 | Yes | ||
| 114 | GTF2E2 | 18635 | 18339 | -1.848 | -0.1812 | Yes | ||
| 115 | NUP43 | 20094 | 18353 | -1.873 | -0.1599 | Yes | ||
| 116 | POLR2G | 23753 | 18407 | -1.978 | -0.1394 | Yes | ||
| 117 | GTF2E1 | 22597 | 18481 | -2.218 | -0.1172 | Yes | ||
| 118 | TCEB1 | 13995 | 18503 | -2.265 | -0.0916 | Yes | ||
| 119 | PSMB5 | 9633 | 18507 | -2.296 | -0.0647 | Yes | ||
| 120 | SUPT4H1 | 9938 | 18579 | -2.818 | -0.0353 | Yes | ||
| 121 | PSMD2 | 10137 5724 | 18598 | -3.160 | 0.0010 | Yes |