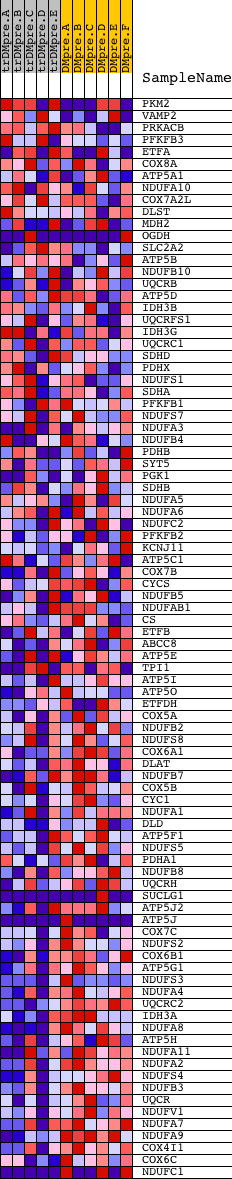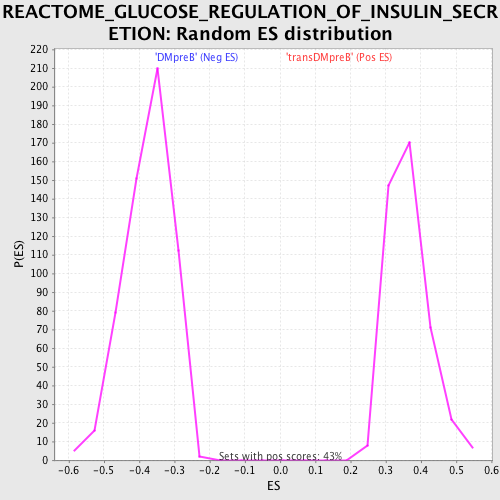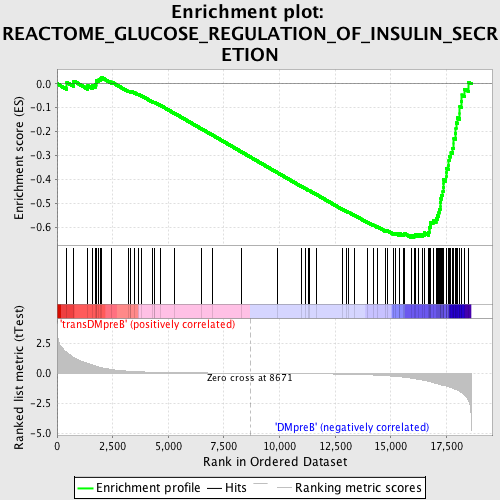
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | REACTOME_GLUCOSE_REGULATION_OF_INSULIN_SECRETION |
| Enrichment Score (ES) | -0.64275044 |
| Normalized Enrichment Score (NES) | -1.7193832 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0069261054 |
| FWER p-Value | 0.125 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PKM2 | 3642 9573 | 439 | 1.709 | 0.0057 | No | ||
| 2 | VAMP2 | 20826 | 750 | 1.302 | 0.0113 | No | ||
| 3 | PRKACB | 15140 | 1381 | 0.813 | -0.0087 | No | ||
| 4 | PFKFB3 | 2748 | 1585 | 0.690 | -0.0078 | No | ||
| 5 | ETFA | 4292 8495 3155 | 1711 | 0.621 | -0.0038 | No | ||
| 6 | COX8A | 8777 | 1748 | 0.599 | 0.0045 | No | ||
| 7 | ATP5A1 | 23505 | 1763 | 0.591 | 0.0139 | No | ||
| 8 | NDUFA10 | 7420 | 1880 | 0.518 | 0.0166 | No | ||
| 9 | COX7A2L | 9819 | 1965 | 0.474 | 0.0202 | No | ||
| 10 | DLST | 8131 | 2008 | 0.455 | 0.0257 | No | ||
| 11 | MDH2 | 9399 | 2446 | 0.306 | 0.0074 | No | ||
| 12 | OGDH | 9502 1412 | 3215 | 0.170 | -0.0311 | No | ||
| 13 | SLC2A2 | 15623 1880 1855 | 3307 | 0.158 | -0.0333 | No | ||
| 14 | ATP5B | 19846 | 3318 | 0.157 | -0.0312 | No | ||
| 15 | NDUFB10 | 23084 | 3487 | 0.139 | -0.0378 | No | ||
| 16 | UQCRB | 12545 | 3654 | 0.125 | -0.0446 | No | ||
| 17 | ATP5D | 19949 | 3793 | 0.115 | -0.0501 | No | ||
| 18 | IDH3B | 9299 | 4288 | 0.085 | -0.0753 | No | ||
| 19 | UQCRFS1 | 21499 | 4365 | 0.081 | -0.0780 | No | ||
| 20 | IDH3G | 24132 | 4380 | 0.081 | -0.0774 | No | ||
| 21 | UQCRC1 | 10259 | 4631 | 0.071 | -0.0896 | No | ||
| 22 | SDHD | 19125 | 5278 | 0.051 | -0.1236 | No | ||
| 23 | PDHX | 14510 | 6490 | 0.028 | -0.1885 | No | ||
| 24 | NDUFS1 | 13940 | 6971 | 0.020 | -0.2141 | No | ||
| 25 | SDHA | 12436 | 8267 | 0.005 | -0.2839 | No | ||
| 26 | PFKFB1 | 9552 | 9909 | -0.016 | -0.3722 | No | ||
| 27 | NDUFS7 | 19946 | 10999 | -0.031 | -0.4304 | No | ||
| 28 | NDUFA3 | 18410 | 11178 | -0.034 | -0.4395 | No | ||
| 29 | NDUFB4 | 22596 7541 | 11298 | -0.035 | -0.4453 | No | ||
| 30 | PDHB | 12670 7548 | 11347 | -0.036 | -0.4473 | No | ||
| 31 | SYT5 | 17986 | 11672 | -0.042 | -0.4640 | No | ||
| 32 | PGK1 | 9557 5244 | 12838 | -0.072 | -0.5257 | No | ||
| 33 | SDHB | 2348 12566 14880 | 12985 | -0.078 | -0.5322 | No | ||
| 34 | NDUFA5 | 1076 7544 | 13114 | -0.082 | -0.5377 | No | ||
| 35 | NDUFA6 | 12473 | 13383 | -0.093 | -0.5506 | No | ||
| 36 | NDUFC2 | 18183 | 13965 | -0.125 | -0.5798 | No | ||
| 37 | PFKFB2 | 4116 5242 | 14211 | -0.142 | -0.5906 | No | ||
| 38 | KCNJ11 | 17822 | 14392 | -0.160 | -0.5975 | No | ||
| 39 | ATP5C1 | 8635 | 14751 | -0.196 | -0.6135 | No | ||
| 40 | COX7B | 7260 | 14837 | -0.207 | -0.6145 | No | ||
| 41 | CYCS | 8821 | 15141 | -0.247 | -0.6266 | No | ||
| 42 | NDUFB5 | 12277 | 15191 | -0.254 | -0.6249 | No | ||
| 43 | NDUFAB1 | 7667 | 15387 | -0.286 | -0.6305 | No | ||
| 44 | CS | 19839 | 15391 | -0.287 | -0.6257 | No | ||
| 45 | ETFB | 18279 | 15583 | -0.327 | -0.6304 | No | ||
| 46 | ABCC8 | 9941 11649 | 15623 | -0.336 | -0.6267 | No | ||
| 47 | ATP5E | 14321 | 15921 | -0.408 | -0.6357 | Yes | ||
| 48 | TPI1 | 5795 10212 | 16042 | -0.443 | -0.6346 | Yes | ||
| 49 | ATP5I | 8637 | 16098 | -0.457 | -0.6297 | Yes | ||
| 50 | ATP5O | 22539 | 16259 | -0.501 | -0.6297 | Yes | ||
| 51 | ETFDH | 15313 | 16430 | -0.561 | -0.6293 | Yes | ||
| 52 | COX5A | 19431 | 16505 | -0.589 | -0.6231 | Yes | ||
| 53 | NDUFB2 | 7542 | 16689 | -0.666 | -0.6215 | Yes | ||
| 54 | NDUFS8 | 23950 | 16721 | -0.685 | -0.6114 | Yes | ||
| 55 | COX6A1 | 8774 4553 | 16723 | -0.686 | -0.5997 | Yes | ||
| 56 | DLAT | 19123 | 16784 | -0.713 | -0.5907 | Yes | ||
| 57 | NDUFB7 | 12429 | 16791 | -0.715 | -0.5787 | Yes | ||
| 58 | COX5B | 4552 | 16899 | -0.772 | -0.5712 | Yes | ||
| 59 | CYC1 | 12348 | 17034 | -0.848 | -0.5639 | Yes | ||
| 60 | NDUFA1 | 24170 | 17089 | -0.873 | -0.5518 | Yes | ||
| 61 | DLD | 2097 21090 | 17160 | -0.902 | -0.5400 | Yes | ||
| 62 | ATP5F1 | 15212 | 17170 | -0.904 | -0.5250 | Yes | ||
| 63 | NDUFS5 | 9296 | 17217 | -0.932 | -0.5114 | Yes | ||
| 64 | PDHA1 | 24020 | 17220 | -0.933 | -0.4955 | Yes | ||
| 65 | NDUFB8 | 7418 | 17249 | -0.948 | -0.4807 | Yes | ||
| 66 | UQCRH | 12378 | 17279 | -0.965 | -0.4657 | Yes | ||
| 67 | SUCLG1 | 17409 1059 | 17342 | -0.997 | -0.4519 | Yes | ||
| 68 | ATP5J2 | 12186 | 17355 | -0.999 | -0.4353 | Yes | ||
| 69 | ATP5J | 880 22555 | 17364 | -1.001 | -0.4186 | Yes | ||
| 70 | COX7C | 8776 | 17367 | -1.001 | -0.4015 | Yes | ||
| 71 | NDUFS2 | 4089 13760 | 17480 | -1.054 | -0.3894 | Yes | ||
| 72 | COX6B1 | 4283 8479 | 17485 | -1.056 | -0.3714 | Yes | ||
| 73 | ATP5G1 | 8636 | 17491 | -1.058 | -0.3535 | Yes | ||
| 74 | NDUFS3 | 7553 12683 | 17602 | -1.123 | -0.3401 | Yes | ||
| 75 | NDUFA4 | 9451 | 17610 | -1.130 | -0.3211 | Yes | ||
| 76 | UQCRC2 | 18102 | 17627 | -1.137 | -0.3024 | Yes | ||
| 77 | IDH3A | 19447 | 17703 | -1.181 | -0.2861 | Yes | ||
| 78 | NDUFA8 | 14605 | 17783 | -1.240 | -0.2691 | Yes | ||
| 79 | ATP5H | 12948 | 17834 | -1.274 | -0.2499 | Yes | ||
| 80 | NDUFA11 | 12832 | 17835 | -1.274 | -0.2280 | Yes | ||
| 81 | NDUFA2 | 9450 | 17911 | -1.329 | -0.2092 | Yes | ||
| 82 | NDUFS4 | 9452 5157 3173 | 17926 | -1.349 | -0.1867 | Yes | ||
| 83 | NDUFB3 | 12362 | 17932 | -1.353 | -0.1637 | Yes | ||
| 84 | UQCR | 12382 | 17992 | -1.399 | -0.1428 | Yes | ||
| 85 | NDUFV1 | 23953 | 18102 | -1.507 | -0.1228 | Yes | ||
| 86 | NDUFA7 | 12347 | 18104 | -1.509 | -0.0969 | Yes | ||
| 87 | NDUFA9 | 12288 | 18165 | -1.581 | -0.0730 | Yes | ||
| 88 | COX4I1 | 18444 | 18198 | -1.613 | -0.0470 | Yes | ||
| 89 | COX6C | 8775 | 18317 | -1.804 | -0.0223 | Yes | ||
| 90 | NDUFC1 | 7287 12339 | 18491 | -2.233 | 0.0067 | Yes |