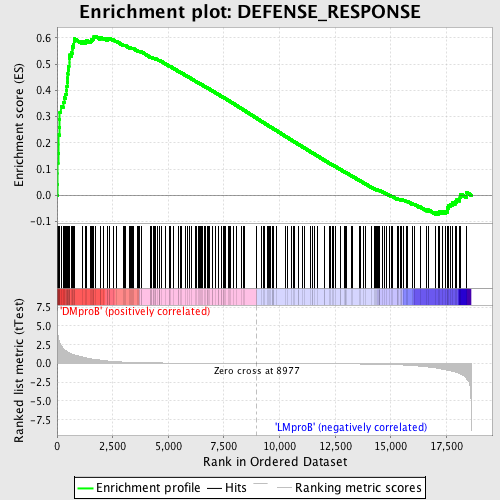
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | DEFENSE_RESPONSE |
| Enrichment Score (ES) | 0.606561 |
| Normalized Enrichment Score (NES) | 1.6661733 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.055812214 |
| FWER p-Value | 0.443 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CAMP | 18990 | 26 | 4.097 | 0.0411 | Yes | ||
| 2 | S100A9 | 9774 | 33 | 3.934 | 0.0816 | Yes | ||
| 3 | LTB4R | 22003 | 35 | 3.854 | 0.1215 | Yes | ||
| 4 | PGLYRP1 | 18369 | 41 | 3.707 | 0.1597 | Yes | ||
| 5 | CFP | 24174 | 58 | 3.500 | 0.1952 | Yes | ||
| 6 | S100A8 | 15527 | 59 | 3.473 | 0.2312 | Yes | ||
| 7 | LSP1 | 18003 | 99 | 2.973 | 0.2600 | Yes | ||
| 8 | CEBPB | 8733 | 115 | 2.822 | 0.2884 | Yes | ||
| 9 | TYROBP | 18302 | 124 | 2.734 | 0.3164 | Yes | ||
| 10 | RSAD2 | 21094 | 174 | 2.434 | 0.3389 | Yes | ||
| 11 | ALOX5AP | 16616 | 272 | 2.000 | 0.3544 | Yes | ||
| 12 | WAS | 24193 | 318 | 1.895 | 0.3717 | Yes | ||
| 13 | CCR7 | 20254 | 387 | 1.689 | 0.3855 | Yes | ||
| 14 | CD84 | 14049 | 428 | 1.624 | 0.4002 | Yes | ||
| 15 | HDAC7A | 22147 | 438 | 1.607 | 0.4164 | Yes | ||
| 16 | TNIP1 | 12192 20450 1320 | 453 | 1.579 | 0.4320 | Yes | ||
| 17 | BCL10 | 15397 | 471 | 1.553 | 0.4472 | Yes | ||
| 18 | ITGB1 | 3872 18411 | 482 | 1.537 | 0.4626 | Yes | ||
| 19 | IL12A | 4913 | 492 | 1.522 | 0.4779 | Yes | ||
| 20 | LBP | 14760 | 510 | 1.486 | 0.4924 | Yes | ||
| 21 | CD48 | 14051 | 536 | 1.448 | 0.5060 | Yes | ||
| 22 | CD19 | 17640 | 537 | 1.442 | 0.5210 | Yes | ||
| 23 | TPST1 | 10214 | 555 | 1.407 | 0.5347 | Yes | ||
| 24 | ORM1 | 9516 | 640 | 1.273 | 0.5433 | Yes | ||
| 25 | CLEC5A | 6225 | 678 | 1.218 | 0.5540 | Yes | ||
| 26 | NOD1 | 17141 | 681 | 1.214 | 0.5664 | Yes | ||
| 27 | ZNF148 | 5949 | 749 | 1.147 | 0.5747 | Yes | ||
| 28 | KCNN4 | 18353 | 759 | 1.137 | 0.5860 | Yes | ||
| 29 | NFE2L1 | 9457 | 787 | 1.113 | 0.5961 | Yes | ||
| 30 | ANXA1 | 9282 | 1145 | 0.854 | 0.5855 | Yes | ||
| 31 | CX3CR1 | 18965 | 1274 | 0.758 | 0.5865 | Yes | ||
| 32 | CD81 | 8719 | 1328 | 0.725 | 0.5911 | Yes | ||
| 33 | NFRKB | 10642 | 1510 | 0.622 | 0.5877 | Yes | ||
| 34 | PRDX5 | 7053 3732 | 1536 | 0.609 | 0.5927 | Yes | ||
| 35 | VEZF1 | 20709 | 1605 | 0.578 | 0.5950 | Yes | ||
| 36 | NFKB1 | 15160 | 1638 | 0.564 | 0.5991 | Yes | ||
| 37 | BECN1 | 7077 | 1655 | 0.558 | 0.6040 | Yes | ||
| 38 | NLRC4 | 11225 | 1711 | 0.532 | 0.6066 | Yes | ||
| 39 | NFX1 | 2475 | 1933 | 0.444 | 0.5992 | No | ||
| 40 | CBARA1 | 20015 | 1949 | 0.436 | 0.6029 | No | ||
| 41 | MEFV | 22686 | 2091 | 0.384 | 0.5992 | No | ||
| 42 | AIF1 | 1582 23005 | 2271 | 0.334 | 0.5929 | No | ||
| 43 | NOD2 | 6384 | 2275 | 0.332 | 0.5962 | No | ||
| 44 | PLA2G7 | 23233 | 2351 | 0.312 | 0.5954 | No | ||
| 45 | CXCR4 | 13844 | 2363 | 0.308 | 0.5980 | No | ||
| 46 | CYSLTR1 | 24082 | 2522 | 0.271 | 0.5922 | No | ||
| 47 | MLF2 | 17283 | 2666 | 0.239 | 0.5869 | No | ||
| 48 | AOAH | 11296 | 2967 | 0.191 | 0.5726 | No | ||
| 49 | AGER | 23276 1576 | 3017 | 0.185 | 0.5718 | No | ||
| 50 | KNG1 | 9244 22809 | 3052 | 0.180 | 0.5719 | No | ||
| 51 | CD40 | 10207 2961 2968 5783 | 3261 | 0.155 | 0.5622 | No | ||
| 52 | MST1R | 19318 3106 | 3298 | 0.151 | 0.5618 | No | ||
| 53 | ADORA2B | 20850 | 3354 | 0.145 | 0.5603 | No | ||
| 54 | CXCL9 | 5099 | 3370 | 0.144 | 0.5610 | No | ||
| 55 | TCIRG1 | 3700 3752 23951 | 3421 | 0.139 | 0.5597 | No | ||
| 56 | PRF1 | 20011 | 3634 | 0.120 | 0.5494 | No | ||
| 57 | IRAK2 | 17324 | 3658 | 0.118 | 0.5494 | No | ||
| 58 | LYST | 9325 5040 | 3701 | 0.115 | 0.5483 | No | ||
| 59 | PGLYRP2 | 7174 | 3721 | 0.113 | 0.5485 | No | ||
| 60 | XCR1 | 18953 | 3788 | 0.110 | 0.5460 | No | ||
| 61 | CCR1 | 18951 | 3798 | 0.109 | 0.5466 | No | ||
| 62 | C2 | 23013 | 4198 | 0.086 | 0.5259 | No | ||
| 63 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 4210 | 0.085 | 0.5262 | No | ||
| 64 | PLA2G2E | 16016 | 4229 | 0.084 | 0.5260 | No | ||
| 65 | STAB2 | 19656 | 4245 | 0.084 | 0.5261 | No | ||
| 66 | SFTPD | 21867 | 4349 | 0.079 | 0.5213 | No | ||
| 67 | AOX1 | 14246 4090 | 4352 | 0.079 | 0.5220 | No | ||
| 68 | TLR6 | 215 16528 | 4381 | 0.078 | 0.5213 | No | ||
| 69 | TLR8 | 9308 | 4434 | 0.076 | 0.5193 | No | ||
| 70 | TNFRSF1A | 1181 10206 | 4442 | 0.076 | 0.5197 | No | ||
| 71 | CRP | 8793 | 4489 | 0.074 | 0.5180 | No | ||
| 72 | TLR3 | 18884 | 4590 | 0.070 | 0.5133 | No | ||
| 73 | CHST2 | 7032 | 4598 | 0.070 | 0.5136 | No | ||
| 74 | IL18RAP | 14262 | 4611 | 0.070 | 0.5137 | No | ||
| 75 | GHSR | 1810 15625 | 4710 | 0.067 | 0.5091 | No | ||
| 76 | NLRP3 | 20862 1262 | 4868 | 0.062 | 0.5012 | No | ||
| 77 | IFNK | 11837 | 5039 | 0.058 | 0.4925 | No | ||
| 78 | TIAL1 | 996 5753 | 5066 | 0.057 | 0.4917 | No | ||
| 79 | CXCL2 | 9790 | 5083 | 0.057 | 0.4914 | No | ||
| 80 | CD40LG | 24330 | 5247 | 0.052 | 0.4831 | No | ||
| 81 | NCF1 | 16354 | 5443 | 0.048 | 0.4730 | No | ||
| 82 | NMI | 14590 | 5445 | 0.048 | 0.4735 | No | ||
| 83 | CCL24 | 16342 | 5538 | 0.046 | 0.4689 | No | ||
| 84 | MX1 | 9432 22529 | 5588 | 0.045 | 0.4667 | No | ||
| 85 | NFATC3 | 5169 | 5784 | 0.041 | 0.4566 | No | ||
| 86 | EREG | 4679 16797 | 5840 | 0.040 | 0.4540 | No | ||
| 87 | CCL22 | 18518 | 5947 | 0.038 | 0.4486 | No | ||
| 88 | CLEC1B | 17270 1080 | 6025 | 0.037 | 0.4448 | No | ||
| 89 | CD83 | 21653 | 6219 | 0.034 | 0.4347 | No | ||
| 90 | CD97 | 18822 3874 | 6247 | 0.033 | 0.4336 | No | ||
| 91 | DEFB103A | 18665 | 6335 | 0.031 | 0.4292 | No | ||
| 92 | CXCL10 | 16477 | 6387 | 0.031 | 0.4267 | No | ||
| 93 | COLEC12 | 23624 8946 | 6426 | 0.030 | 0.4250 | No | ||
| 94 | C5AR1 | 17957 | 6490 | 0.029 | 0.4218 | No | ||
| 95 | NFATC4 | 22002 | 6517 | 0.028 | 0.4207 | No | ||
| 96 | IL9 | 21444 | 6628 | 0.027 | 0.4150 | No | ||
| 97 | PGLYRP3 | 10812 | 6661 | 0.026 | 0.4136 | No | ||
| 98 | OR2H2 | 17694 | 6748 | 0.025 | 0.4092 | No | ||
| 99 | APOA4 | 4401 | 6770 | 0.025 | 0.4083 | No | ||
| 100 | LALBA | 22141 | 6802 | 0.025 | 0.4069 | No | ||
| 101 | DCDC2 | 9703 | 6836 | 0.024 | 0.4053 | No | ||
| 102 | CCR3 | 19251 | 6963 | 0.023 | 0.3987 | No | ||
| 103 | CCL4 | 1362 20730 | 6983 | 0.022 | 0.3979 | No | ||
| 104 | SELE | 14074 | 7106 | 0.021 | 0.3915 | No | ||
| 105 | KLRC3 | 16975 | 7257 | 0.019 | 0.3835 | No | ||
| 106 | APCS | 13748 | 7261 | 0.019 | 0.3836 | No | ||
| 107 | HDAC9 | 21084 2131 | 7396 | 0.017 | 0.3765 | No | ||
| 108 | TLR7 | 24004 | 7409 | 0.017 | 0.3760 | No | ||
| 109 | CCR4 | 18978 | 7471 | 0.017 | 0.3729 | No | ||
| 110 | PLA2G2D | 16018 | 7490 | 0.016 | 0.3720 | No | ||
| 111 | ORM2 | 16193 | 7528 | 0.016 | 0.3702 | No | ||
| 112 | TACR1 | 1082 17403 | 7576 | 0.015 | 0.3678 | No | ||
| 113 | EDG3 | 21639 | 7696 | 0.014 | 0.3615 | No | ||
| 114 | DMBT1 | 18050 | 7742 | 0.013 | 0.3592 | No | ||
| 115 | CXCL1 | 9044 | 7802 | 0.013 | 0.3561 | No | ||
| 116 | IL20 | 13840 | 7922 | 0.011 | 0.3498 | No | ||
| 117 | ITK | 4934 | 8042 | 0.010 | 0.3434 | No | ||
| 118 | C3AR1 | 8668 | 8273 | 0.007 | 0.3310 | No | ||
| 119 | CD160 | 12023 | 8362 | 0.006 | 0.3263 | No | ||
| 120 | CCR6 | 1505 23390 | 8423 | 0.005 | 0.3231 | No | ||
| 121 | LY75 | 9313 5026 | 8950 | 0.000 | 0.2945 | No | ||
| 122 | ADORA1 | 13831 | 8951 | 0.000 | 0.2945 | No | ||
| 123 | CCL11 | 20742 | 9193 | -0.002 | 0.2814 | No | ||
| 124 | CXCL11 | 16478 | 9294 | -0.003 | 0.2760 | No | ||
| 125 | CHRNA7 | 17808 | 9323 | -0.004 | 0.2745 | No | ||
| 126 | CCR9 | 8753 | 9452 | -0.005 | 0.2676 | No | ||
| 127 | IL17C | 18438 | 9521 | -0.005 | 0.2640 | No | ||
| 128 | IL8RA | 13925 | 9560 | -0.006 | 0.2620 | No | ||
| 129 | TNFAIP6 | 10203 | 9610 | -0.006 | 0.2594 | No | ||
| 130 | IL1RL2 | 4212 | 9668 | -0.007 | 0.2564 | No | ||
| 131 | IL12B | 20918 | 9683 | -0.007 | 0.2557 | No | ||
| 132 | FOSL1 | 23779 | 9707 | -0.007 | 0.2545 | No | ||
| 133 | ALOX15 | 20370 | 9747 | -0.007 | 0.2525 | No | ||
| 134 | WFDC12 | 14356 | 9869 | -0.009 | 0.2460 | No | ||
| 135 | IFNA2 | 9149 | 10274 | -0.013 | 0.2242 | No | ||
| 136 | TFF3 | 23038 | 10355 | -0.014 | 0.2200 | No | ||
| 137 | KLRC2 | 9243 | 10539 | -0.016 | 0.2102 | No | ||
| 138 | BNIP3 | 17581 | 10620 | -0.016 | 0.2060 | No | ||
| 139 | PTX3 | 9668 15575 | 10657 | -0.017 | 0.2042 | No | ||
| 140 | HRH1 | 17321 | 10828 | -0.019 | 0.1952 | No | ||
| 141 | RIPK2 | 2528 15935 | 10837 | -0.019 | 0.1949 | No | ||
| 142 | ANKRD1 | 23689 | 11045 | -0.022 | 0.1839 | No | ||
| 143 | GPR68 | 21004 | 11118 | -0.022 | 0.1802 | No | ||
| 144 | IL4 | 9174 | 11378 | -0.026 | 0.1664 | No | ||
| 145 | UMOD | 17660 | 11457 | -0.027 | 0.1625 | No | ||
| 146 | IL5 | 20884 10220 | 11462 | -0.027 | 0.1625 | No | ||
| 147 | NCR1 | 18409 | 11582 | -0.028 | 0.1564 | No | ||
| 148 | PARP4 | 21999 | 11712 | -0.030 | 0.1497 | No | ||
| 149 | FPRL1 | 8982 | 11997 | -0.034 | 0.1346 | No | ||
| 150 | ADORA2A | 8557 | 12029 | -0.034 | 0.1333 | No | ||
| 151 | AOC3 | 8595 | 12260 | -0.038 | 0.1212 | No | ||
| 152 | IL1A | 4915 | 12300 | -0.038 | 0.1194 | No | ||
| 153 | ADORA3 | 4352 | 12383 | -0.040 | 0.1154 | No | ||
| 154 | FOXN1 | 20337 | 12414 | -0.040 | 0.1142 | No | ||
| 155 | NOX4 | 18196 | 12532 | -0.042 | 0.1083 | No | ||
| 156 | CRTAM | 19160 | 12733 | -0.046 | 0.0979 | No | ||
| 157 | GHRL | 1183 17040 | 12748 | -0.046 | 0.0976 | No | ||
| 158 | DEFB1 | 18663 | 12921 | -0.049 | 0.0888 | No | ||
| 159 | SCG2 | 5410 14425 | 12960 | -0.050 | 0.0872 | No | ||
| 160 | IL28RA | 10823 16038 | 12981 | -0.050 | 0.0866 | No | ||
| 161 | CCR5 | 8754 4534 | 13005 | -0.051 | 0.0859 | No | ||
| 162 | CADM1 | 7057 | 13209 | -0.056 | 0.0755 | No | ||
| 163 | SPACA3 | 13680 | 13290 | -0.058 | 0.0717 | No | ||
| 164 | CX3CL1 | 9791 | 13583 | -0.066 | 0.0565 | No | ||
| 165 | INHBB | 13858 | 13629 | -0.068 | 0.0548 | No | ||
| 166 | MBL2 | 23886 | 13780 | -0.073 | 0.0474 | No | ||
| 167 | CFHR1 | 11926 13815 | 13883 | -0.077 | 0.0427 | No | ||
| 168 | MX2 | 9433 | 14148 | -0.087 | 0.0292 | No | ||
| 169 | STAB1 | 21892 | 14247 | -0.091 | 0.0249 | No | ||
| 170 | MGLL | 17368 1145 | 14328 | -0.095 | 0.0215 | No | ||
| 171 | RAC1 | 16302 | 14356 | -0.096 | 0.0210 | No | ||
| 172 | F11R | 9199 | 14381 | -0.097 | 0.0207 | No | ||
| 173 | CYBB | 4579 | 14461 | -0.101 | 0.0175 | No | ||
| 174 | ELF3 | 8894 13824 | 14483 | -0.103 | 0.0174 | No | ||
| 175 | CCR2 | 19250 | 14494 | -0.103 | 0.0179 | No | ||
| 176 | CD5L | 15560 | 14504 | -0.104 | 0.0185 | No | ||
| 177 | VPS45 | 15242 | 14634 | -0.111 | 0.0127 | No | ||
| 178 | CDO1 | 8731 | 14703 | -0.114 | 0.0102 | No | ||
| 179 | CCL3 | 20317 | 14822 | -0.122 | 0.0050 | No | ||
| 180 | CSF3R | 8804 8803 2326 4565 | 14921 | -0.129 | 0.0010 | No | ||
| 181 | IL17RB | 21898 | 15025 | -0.138 | -0.0031 | No | ||
| 182 | IL1RAP | 1653 1676 4917 1748 1712 9173 | 15085 | -0.143 | -0.0049 | No | ||
| 183 | KLRG1 | 17014 | 15278 | -0.162 | -0.0136 | No | ||
| 184 | AHSG | 22812 | 15313 | -0.165 | -0.0138 | No | ||
| 185 | IL13 | 20461 | 15361 | -0.170 | -0.0145 | No | ||
| 186 | GATA3 | 9004 | 15431 | -0.179 | -0.0164 | No | ||
| 187 | PGLYRP4 | 11810 | 15481 | -0.185 | -0.0172 | No | ||
| 188 | DEFB127 | 622 | 15499 | -0.188 | -0.0162 | No | ||
| 189 | CAMLG | 21627 | 15582 | -0.201 | -0.0185 | No | ||
| 190 | PTPRC | 5327 9662 | 15705 | -0.217 | -0.0229 | No | ||
| 191 | SOCS6 | 12042 7044 | 15755 | -0.226 | -0.0232 | No | ||
| 192 | SPN | 452 5493 | 15968 | -0.264 | -0.0320 | No | ||
| 193 | RELA | 23783 | 16057 | -0.286 | -0.0338 | No | ||
| 194 | TGFB1 | 18332 | 16317 | -0.350 | -0.0442 | No | ||
| 195 | IL8RB | 8752 4533 | 16608 | -0.430 | -0.0555 | No | ||
| 196 | HP | 3805 18750 | 16701 | -0.467 | -0.0557 | No | ||
| 197 | MPO | 5116 20710 1324 | 17023 | -0.599 | -0.0669 | No | ||
| 198 | LY96 | 14292 | 17129 | -0.646 | -0.0659 | No | ||
| 199 | ABCF1 | 22996 10329 5897 | 17182 | -0.670 | -0.0618 | No | ||
| 200 | PTPRCAP | 23767 | 17340 | -0.763 | -0.0624 | No | ||
| 201 | SCYE1 | 8897 | 17457 | -0.844 | -0.0600 | No | ||
| 202 | BNIP3L | 21775 | 17537 | -0.884 | -0.0551 | No | ||
| 203 | NCF2 | 14098 5151 9446 | 17554 | -0.893 | -0.0467 | No | ||
| 204 | LGALS3BP | 20129 | 17602 | -0.927 | -0.0396 | No | ||
| 205 | TARBP2 | 10033 10034 5626 | 17685 | -0.988 | -0.0338 | No | ||
| 206 | BLNK | 23681 3691 | 17772 | -1.030 | -0.0278 | No | ||
| 207 | IL10RB | 9169 | 17888 | -1.114 | -0.0225 | No | ||
| 208 | FOS | 21202 | 17967 | -1.183 | -0.0145 | No | ||
| 209 | HDAC5 | 1480 20205 | 18107 | -1.357 | -0.0079 | No | ||
| 210 | RNASE6 | 13439 | 18142 | -1.414 | 0.0049 | No | ||
| 211 | CCL5 | 20320 | 18406 | -2.006 | 0.0114 | No |

