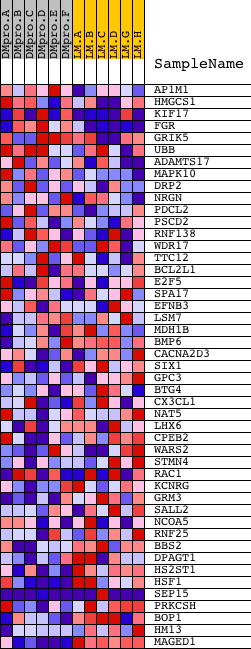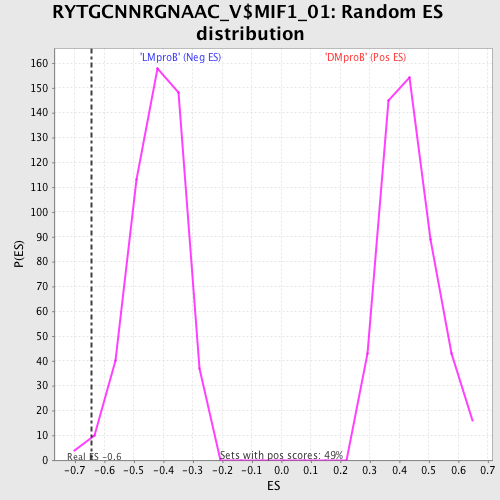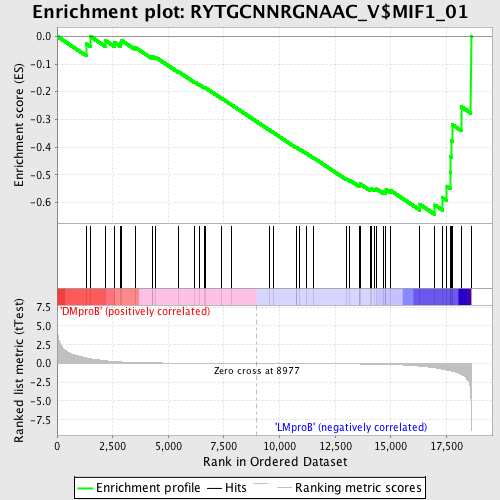
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | RYTGCNNRGNAAC_V$MIF1_01 |
| Enrichment Score (ES) | -0.6434384 |
| Normalized Enrichment Score (NES) | -1.523313 |
| Nominal p-value | 0.015686275 |
| FDR q-value | 0.62346363 |
| FWER p-Value | 0.8 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AP1M1 | 18571 | 1312 | 0.733 | -0.0275 | No | ||
| 2 | HMGCS1 | 9917 | 1501 | 0.626 | -0.0008 | No | ||
| 3 | KIF17 | 9215 2392 | 2157 | 0.364 | -0.0147 | No | ||
| 4 | FGR | 4723 | 2569 | 0.260 | -0.0215 | No | ||
| 5 | GRIK5 | 350 17930 | 2843 | 0.207 | -0.0240 | No | ||
| 6 | UBB | 10240 | 2874 | 0.202 | -0.0137 | No | ||
| 7 | ADAMTS17 | 598 | 3505 | 0.131 | -0.0399 | No | ||
| 8 | MAPK10 | 11169 | 4286 | 0.082 | -0.0771 | No | ||
| 9 | DRP2 | 24254 | 4288 | 0.082 | -0.0724 | No | ||
| 10 | NRGN | 19176 | 4433 | 0.076 | -0.0756 | No | ||
| 11 | PDCL2 | 3604 16508 | 5469 | 0.047 | -0.1286 | No | ||
| 12 | PSCD2 | 17826 | 6192 | 0.034 | -0.1655 | No | ||
| 13 | RNF138 | 2006 23613 2029 2037 | 6421 | 0.030 | -0.1760 | No | ||
| 14 | WDR17 | 3785 18873 | 6621 | 0.027 | -0.1851 | No | ||
| 15 | TTC12 | 19127 | 6665 | 0.026 | -0.1859 | No | ||
| 16 | BCL2L1 | 4440 2930 8652 | 7407 | 0.017 | -0.2248 | No | ||
| 17 | E2F5 | 15633 | 7857 | 0.012 | -0.2482 | No | ||
| 18 | SPA17 | 19175 | 9542 | -0.006 | -0.3386 | No | ||
| 19 | EFNB3 | 20391 | 9705 | -0.007 | -0.3469 | No | ||
| 20 | LSM7 | 12286 | 10746 | -0.018 | -0.4018 | No | ||
| 21 | MDH1B | 13938 | 10775 | -0.018 | -0.4023 | No | ||
| 22 | BMP6 | 8659 | 10897 | -0.020 | -0.4076 | No | ||
| 23 | CACNA2D3 | 8677 | 11201 | -0.023 | -0.4226 | No | ||
| 24 | SIX1 | 9821 | 11522 | -0.027 | -0.4382 | No | ||
| 25 | GPC3 | 24151 | 13022 | -0.051 | -0.5159 | No | ||
| 26 | BTG4 | 19456 | 13155 | -0.054 | -0.5198 | No | ||
| 27 | CX3CL1 | 9791 | 13583 | -0.066 | -0.5389 | No | ||
| 28 | NAT5 | 18601 7496 12597 | 13612 | -0.067 | -0.5365 | No | ||
| 29 | LHX6 | 9275 2914 | 13615 | -0.067 | -0.5326 | No | ||
| 30 | CPEB2 | 6077 | 14064 | -0.084 | -0.5518 | No | ||
| 31 | WARS2 | 12884 7688 | 14109 | -0.086 | -0.5492 | No | ||
| 32 | STMN4 | 21977 | 14267 | -0.092 | -0.5522 | No | ||
| 33 | RAC1 | 16302 | 14356 | -0.096 | -0.5513 | No | ||
| 34 | KCNRG | 495 11601 | 14658 | -0.112 | -0.5609 | No | ||
| 35 | GRM3 | 16929 | 14778 | -0.119 | -0.5604 | No | ||
| 36 | SALL2 | 21840 | 14782 | -0.119 | -0.5535 | No | ||
| 37 | NCOA5 | 14341 | 14977 | -0.134 | -0.5561 | No | ||
| 38 | RNF25 | 13922 3929 | 16309 | -0.348 | -0.6073 | No | ||
| 39 | BBS2 | 18787 | 16981 | -0.582 | -0.6092 | Yes | ||
| 40 | DPAGT1 | 19478 | 17337 | -0.761 | -0.5835 | Yes | ||
| 41 | HS2ST1 | 15150 | 17523 | -0.874 | -0.5420 | Yes | ||
| 42 | HSF1 | 4877 | 17671 | -0.978 | -0.4924 | Yes | ||
| 43 | SEP15 | 13527 1895 15398 | 17691 | -0.992 | -0.4350 | Yes | ||
| 44 | PRKCSH | 19535 | 17722 | -1.001 | -0.3777 | Yes | ||
| 45 | BOP1 | 22244 | 17777 | -1.036 | -0.3196 | Yes | ||
| 46 | HM13 | 2880 2707 14793 | 18174 | -1.470 | -0.2544 | Yes | ||
| 47 | MAGED1 | 24108 | 18603 | -4.726 | 0.0007 | Yes |