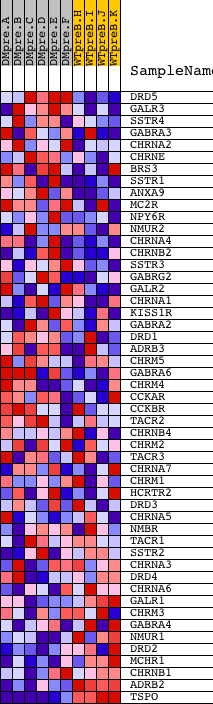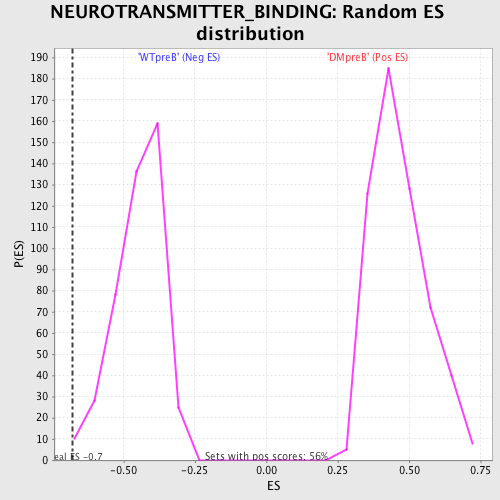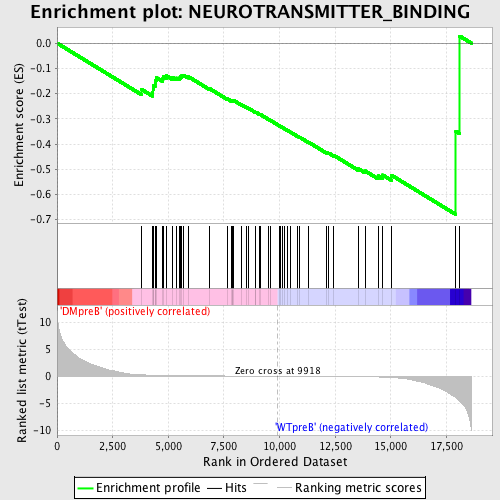
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | NEUROTRANSMITTER_BINDING |
| Enrichment Score (ES) | -0.6794903 |
| Normalized Enrichment Score (NES) | -1.523896 |
| Nominal p-value | 0.0091743115 |
| FDR q-value | 0.30548492 |
| FWER p-Value | 0.99 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DRD5 | 16853 | 3801 | 0.251 | -0.1838 | No | ||
| 2 | GALR3 | 9000 | 4289 | 0.174 | -0.1954 | No | ||
| 3 | SSTR4 | 9836 | 4310 | 0.172 | -0.1822 | No | ||
| 4 | GABRA3 | 24139 | 4334 | 0.168 | -0.1694 | No | ||
| 5 | CHRNA2 | 21978 | 4405 | 0.160 | -0.1598 | No | ||
| 6 | CHRNE | 20368 | 4434 | 0.157 | -0.1482 | No | ||
| 7 | BRS3 | 24333 | 4457 | 0.155 | -0.1364 | No | ||
| 8 | SSTR1 | 21264 | 4753 | 0.128 | -0.1416 | No | ||
| 9 | ANXA9 | 12964 | 4765 | 0.127 | -0.1316 | No | ||
| 10 | MC2R | 23412 | 4905 | 0.118 | -0.1292 | No | ||
| 11 | NPY6R | 23567 | 5192 | 0.102 | -0.1362 | No | ||
| 12 | NMUR2 | 20443 | 5373 | 0.092 | -0.1382 | No | ||
| 13 | CHRNA4 | 8531 4320 | 5504 | 0.086 | -0.1380 | No | ||
| 14 | CHRNB2 | 8532 4321 | 5544 | 0.084 | -0.1330 | No | ||
| 15 | SSTR3 | 22218 | 5607 | 0.081 | -0.1296 | No | ||
| 16 | GABRG2 | 1259 4748 8996 1331 | 5681 | 0.078 | -0.1270 | No | ||
| 17 | GALR2 | 20587 | 5892 | 0.070 | -0.1325 | No | ||
| 18 | CHRNA1 | 4319 14559 8530 | 6841 | 0.046 | -0.1798 | No | ||
| 19 | KISS1R | 19956 | 7647 | 0.030 | -0.2206 | No | ||
| 20 | GABRA2 | 4746 | 7825 | 0.027 | -0.2278 | No | ||
| 21 | DRD1 | 4638 | 7875 | 0.027 | -0.2282 | No | ||
| 22 | ADRB3 | 18901 | 7882 | 0.027 | -0.2263 | No | ||
| 23 | CHRM5 | 10038 | 7936 | 0.026 | -0.2270 | No | ||
| 24 | GABRA6 | 20491 | 8284 | 0.021 | -0.2440 | No | ||
| 25 | CHRM4 | 14945 | 8524 | 0.017 | -0.2554 | No | ||
| 26 | CCKAR | 16531 | 8623 | 0.016 | -0.2593 | No | ||
| 27 | CCKBR | 8706 | 8917 | 0.013 | -0.2741 | No | ||
| 28 | TACR2 | 20004 | 8919 | 0.013 | -0.2731 | No | ||
| 29 | CHRNB4 | 19107 | 9113 | 0.010 | -0.2826 | No | ||
| 30 | CHRM2 | 10840 | 9128 | 0.010 | -0.2825 | No | ||
| 31 | TACR3 | 15415 | 9492 | 0.005 | -0.3016 | No | ||
| 32 | CHRNA7 | 17808 | 9599 | 0.004 | -0.3070 | No | ||
| 33 | CHRM1 | 23943 | 9991 | -0.001 | -0.3280 | No | ||
| 34 | HCRTR2 | 6907 | 10050 | -0.002 | -0.3309 | No | ||
| 35 | DRD3 | 22750 | 10108 | -0.002 | -0.3338 | No | ||
| 36 | CHRNA5 | 8494 | 10219 | -0.004 | -0.3394 | No | ||
| 37 | NMBR | 9466 | 10368 | -0.005 | -0.3469 | No | ||
| 38 | TACR1 | 1082 17403 | 10505 | -0.007 | -0.3537 | No | ||
| 39 | SSTR2 | 20609 | 10814 | -0.012 | -0.3693 | No | ||
| 40 | CHRNA3 | 4291 | 10894 | -0.013 | -0.3725 | No | ||
| 41 | DRD4 | 4639 | 11290 | -0.018 | -0.3922 | No | ||
| 42 | CHRNA6 | 18900 | 12105 | -0.032 | -0.4334 | No | ||
| 43 | GALR1 | 23397 | 12219 | -0.034 | -0.4366 | No | ||
| 44 | CHRM3 | 21547 | 12439 | -0.039 | -0.4451 | No | ||
| 45 | GABRA4 | 16519 | 13536 | -0.075 | -0.4979 | No | ||
| 46 | NMUR1 | 13898 | 13841 | -0.091 | -0.5067 | No | ||
| 47 | DRD2 | 19461 | 14432 | -0.145 | -0.5264 | Yes | ||
| 48 | MCHR1 | 22416 | 14633 | -0.171 | -0.5228 | Yes | ||
| 49 | CHRNB1 | 20384 | 15023 | -0.244 | -0.5234 | Yes | ||
| 50 | ADRB2 | 23422 | 17922 | -3.954 | -0.3492 | Yes | ||
| 51 | TSPO | 2204 22408 | 18098 | -4.628 | 0.0279 | Yes |