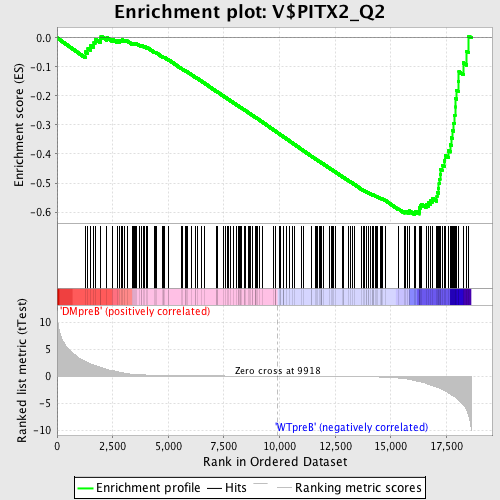
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | V$PITX2_Q2 |
| Enrichment Score (ES) | -0.6084669 |
| Normalized Enrichment Score (NES) | -1.6037179 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.055978063 |
| FWER p-Value | 0.294 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | BRCA2 | 16612 | 1274 | 2.691 | -0.0483 | No | ||
| 2 | NDRG2 | 21843 | 1385 | 2.447 | -0.0353 | No | ||
| 3 | TRPS1 | 8195 | 1515 | 2.233 | -0.0250 | No | ||
| 4 | BMI1 | 4448 15113 | 1655 | 2.046 | -0.0167 | No | ||
| 5 | EIF2B4 | 16574 | 1706 | 1.959 | -0.0043 | No | ||
| 6 | HESX1 | 22070 | 1949 | 1.598 | -0.0050 | No | ||
| 7 | ICT1 | 20598 | 1970 | 1.581 | 0.0061 | No | ||
| 8 | HSD3B7 | 4158 | 2240 | 1.254 | 0.0012 | No | ||
| 9 | FAF1 | 2508 16137 8951 | 2490 | 1.001 | -0.0045 | No | ||
| 10 | DPYSL5 | 92 | 2697 | 0.821 | -0.0094 | No | ||
| 11 | NDUFS2 | 4089 13760 | 2812 | 0.719 | -0.0100 | No | ||
| 12 | LRP6 | 9286 | 2877 | 0.656 | -0.0084 | No | ||
| 13 | CACNA2D3 | 8677 | 2930 | 0.616 | -0.0064 | No | ||
| 14 | PPM2C | 491 | 3034 | 0.536 | -0.0079 | No | ||
| 15 | NCOA2 | 5155 4034 | 3150 | 0.463 | -0.0105 | No | ||
| 16 | TWIST1 | 21291 | 3381 | 0.360 | -0.0202 | No | ||
| 17 | NR2E1 | 19780 | 3420 | 0.348 | -0.0196 | No | ||
| 18 | ACVR1 | 4334 | 3479 | 0.334 | -0.0201 | No | ||
| 19 | NFIX | 9458 | 3501 | 0.326 | -0.0187 | No | ||
| 20 | LTBP1 | 1536 1604 6503 1559 1564 1518 | 3562 | 0.310 | -0.0196 | No | ||
| 21 | GPR158 | 149 | 3693 | 0.275 | -0.0245 | No | ||
| 22 | TRIB1 | 22467 | 3774 | 0.256 | -0.0269 | No | ||
| 23 | APP | 4402 | 3787 | 0.253 | -0.0256 | No | ||
| 24 | ADAMTSL1 | 16181 | 3896 | 0.231 | -0.0296 | No | ||
| 25 | FOXP2 | 4312 17523 8523 | 3935 | 0.225 | -0.0300 | No | ||
| 26 | ORC4L | 11172 6460 | 4035 | 0.208 | -0.0337 | No | ||
| 27 | NCALD | 22310 | 4083 | 0.202 | -0.0347 | No | ||
| 28 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 4380 | 0.162 | -0.0495 | No | ||
| 29 | FES | 17779 3949 | 4400 | 0.160 | -0.0493 | No | ||
| 30 | HCN4 | 9080 | 4475 | 0.153 | -0.0521 | No | ||
| 31 | MEOX2 | 21285 | 4757 | 0.128 | -0.0664 | No | ||
| 32 | ODF2 | 9501 5205 2862 2949 | 4798 | 0.125 | -0.0676 | No | ||
| 33 | GAD1 | 14993 2690 | 4799 | 0.125 | -0.0666 | No | ||
| 34 | BARHL1 | 14634 | 4811 | 0.124 | -0.0663 | No | ||
| 35 | NPPC | 13896 | 5000 | 0.112 | -0.0756 | No | ||
| 36 | MIP | 19845 | 5019 | 0.111 | -0.0757 | No | ||
| 37 | NDST4 | 12259 7228 | 5572 | 0.083 | -0.1050 | No | ||
| 38 | FZD6 | 22486 | 5640 | 0.080 | -0.1080 | No | ||
| 39 | AQP5 | 22365 | 5756 | 0.075 | -0.1137 | No | ||
| 40 | ZMYND17 | 21909 | 5777 | 0.074 | -0.1142 | No | ||
| 41 | HOXA2 | 17151 | 5801 | 0.073 | -0.1149 | No | ||
| 42 | KCTD15 | 17864 | 5849 | 0.071 | -0.1169 | No | ||
| 43 | CREB5 | 10551 | 6052 | 0.065 | -0.1273 | No | ||
| 44 | CDH11 | 18780 3861 8725 | 6211 | 0.060 | -0.1354 | No | ||
| 45 | SLITRK5 | 21939 | 6241 | 0.059 | -0.1365 | No | ||
| 46 | HOXA1 | 1005 17152 | 6290 | 0.058 | -0.1387 | No | ||
| 47 | NEFL | 9459 | 6491 | 0.053 | -0.1491 | No | ||
| 48 | DLC1 | 11931 3855 | 6640 | 0.050 | -0.1568 | No | ||
| 49 | GLRA1 | 9021 | 7162 | 0.039 | -0.1847 | No | ||
| 50 | PRDM8 | 8084 | 7184 | 0.039 | -0.1856 | No | ||
| 51 | UBE2N | 8216 19898 | 7213 | 0.038 | -0.1868 | No | ||
| 52 | NTRK2 | 9491 | 7227 | 0.038 | -0.1872 | No | ||
| 53 | HNRPD | 8645 | 7481 | 0.033 | -0.2007 | No | ||
| 54 | AANAT | 20586 | 7557 | 0.032 | -0.2045 | No | ||
| 55 | KCNJ15 | 4943 9208 | 7640 | 0.030 | -0.2087 | No | ||
| 56 | FOSL2 | 4733 8978 16878 | 7715 | 0.029 | -0.2125 | No | ||
| 57 | HIVEP2 | 9091 | 7812 | 0.028 | -0.2175 | No | ||
| 58 | PRKCE | 9575 | 7915 | 0.026 | -0.2228 | No | ||
| 59 | KCNIP4 | 3638 16536 | 8048 | 0.024 | -0.2298 | No | ||
| 60 | RFX4 | 7721 3410 | 8136 | 0.023 | -0.2343 | No | ||
| 61 | PYY | 20207 | 8141 | 0.023 | -0.2344 | No | ||
| 62 | SLC39A2 | 22025 | 8203 | 0.022 | -0.2375 | No | ||
| 63 | PRG4 | 13592 4036 | 8236 | 0.021 | -0.2391 | No | ||
| 64 | FIGN | 14575 | 8243 | 0.021 | -0.2392 | No | ||
| 65 | RXRG | 14063 | 8295 | 0.021 | -0.2418 | No | ||
| 66 | GUCY2F | 24045 | 8435 | 0.019 | -0.2492 | No | ||
| 67 | LRRTM3 | 19744 | 8438 | 0.019 | -0.2492 | No | ||
| 68 | RORA | 3019 5388 | 8471 | 0.018 | -0.2508 | No | ||
| 69 | IL2 | 15354 | 8580 | 0.017 | -0.2565 | No | ||
| 70 | PALM2 | 10817 2368 | 8653 | 0.016 | -0.2603 | No | ||
| 71 | MSX1 | 16547 | 8672 | 0.015 | -0.2612 | No | ||
| 72 | UNC13B | 10255 | 8764 | 0.015 | -0.2660 | No | ||
| 73 | AMELX | 24002 | 8798 | 0.014 | -0.2677 | No | ||
| 74 | EMX2 | 23806 | 8920 | 0.013 | -0.2741 | No | ||
| 75 | PBX1 | 5224 | 8921 | 0.013 | -0.2740 | No | ||
| 76 | SNX6 | 21070 | 8934 | 0.012 | -0.2746 | No | ||
| 77 | GDNF | 22514 2275 | 8981 | 0.012 | -0.2770 | No | ||
| 78 | HOXA3 | 9108 1075 | 9000 | 0.012 | -0.2779 | No | ||
| 79 | SLC37A4 | 8994 | 9107 | 0.010 | -0.2836 | No | ||
| 80 | SEMA5B | 9801 22777 | 9231 | 0.009 | -0.2902 | No | ||
| 81 | IER5 | 13802 | 9744 | 0.002 | -0.3179 | No | ||
| 82 | TUBA8 | 7012 | 9795 | 0.001 | -0.3206 | No | ||
| 83 | COL2A1 | 22145 4542 | 9823 | 0.001 | -0.3221 | No | ||
| 84 | LPHN2 | 13619 | 9829 | 0.001 | -0.3223 | No | ||
| 85 | PRDM1 | 19775 3337 | 9987 | -0.001 | -0.3308 | No | ||
| 86 | KLK9 | 18267 | 10029 | -0.001 | -0.3331 | No | ||
| 87 | ASB11 | 24211 | 10040 | -0.002 | -0.3336 | No | ||
| 88 | CD2AP | 22975 | 10173 | -0.003 | -0.3407 | No | ||
| 89 | PMFBP1 | 18473 | 10182 | -0.003 | -0.3411 | No | ||
| 90 | MYOG | 14129 | 10185 | -0.003 | -0.3412 | No | ||
| 91 | BAMBI | 23629 | 10325 | -0.005 | -0.3487 | No | ||
| 92 | PURA | 9670 | 10425 | -0.006 | -0.3540 | No | ||
| 93 | EYA1 | 4695 4061 | 10600 | -0.009 | -0.3634 | No | ||
| 94 | CRYGS | 22628 | 10666 | -0.010 | -0.3669 | No | ||
| 95 | PITX2 | 15424 1878 | 10992 | -0.014 | -0.3844 | No | ||
| 96 | SLC13A1 | 17205 | 11076 | -0.015 | -0.3888 | No | ||
| 97 | NPTX2 | 16630 | 11425 | -0.020 | -0.4075 | No | ||
| 98 | COL4A3 | 8766 | 11435 | -0.020 | -0.4078 | No | ||
| 99 | FZD4 | 18193 | 11607 | -0.023 | -0.4169 | No | ||
| 100 | ADAMTS4 | 10791 | 11637 | -0.024 | -0.4183 | No | ||
| 101 | MYB | 3344 3355 19806 | 11688 | -0.025 | -0.4208 | No | ||
| 102 | FGF16 | 24270 | 11798 | -0.027 | -0.4265 | No | ||
| 103 | TCF21 | 3437 | 11847 | -0.027 | -0.4289 | No | ||
| 104 | RIN2 | 14817 | 11866 | -0.028 | -0.4297 | No | ||
| 105 | GRIN2B | 16957 | 11902 | -0.028 | -0.4314 | No | ||
| 106 | MITF | 17349 | 11903 | -0.028 | -0.4311 | No | ||
| 107 | GKN1 | 17081 | 11992 | -0.030 | -0.4357 | No | ||
| 108 | PRL | 21690 | 12257 | -0.036 | -0.4497 | No | ||
| 109 | PRICKLE2 | 10837 | 12319 | -0.037 | -0.4527 | No | ||
| 110 | GPR82 | 24376 | 12375 | -0.038 | -0.4554 | No | ||
| 111 | RORB | 10356 | 12400 | -0.039 | -0.4564 | No | ||
| 112 | RBPMS | 5369 | 12530 | -0.042 | -0.4631 | No | ||
| 113 | SULF1 | 6285 | 12809 | -0.049 | -0.4778 | No | ||
| 114 | MFRP | 11151 | 12852 | -0.050 | -0.4797 | No | ||
| 115 | PPT2 | 1617 12034 | 12884 | -0.051 | -0.4810 | No | ||
| 116 | LIN28 | 15723 | 13081 | -0.057 | -0.4912 | No | ||
| 117 | MLLT7 | 24278 | 13187 | -0.061 | -0.4964 | No | ||
| 118 | ASPA | 20354 | 13255 | -0.063 | -0.4995 | No | ||
| 119 | COL4A4 | 8767 | 13388 | -0.068 | -0.5062 | No | ||
| 120 | RAX | 5361 | 13682 | -0.082 | -0.5214 | No | ||
| 121 | ADORA1 | 13831 | 13775 | -0.088 | -0.5257 | No | ||
| 122 | ACTN3 | 23967 8540 | 13826 | -0.090 | -0.5278 | No | ||
| 123 | ZBTB10 | 10468 | 13893 | -0.094 | -0.5306 | No | ||
| 124 | KITLG | 19889 3342 | 13917 | -0.096 | -0.5311 | No | ||
| 125 | ERBB3 | 19593 | 13976 | -0.100 | -0.5335 | No | ||
| 126 | MAB21L1 | 15589 5046 | 14005 | -0.102 | -0.5342 | No | ||
| 127 | BMPR2 | 14241 4081 | 14078 | -0.109 | -0.5373 | No | ||
| 128 | MYLK | 22778 4213 | 14160 | -0.115 | -0.5408 | No | ||
| 129 | SHOX2 | 1802 5432 | 14173 | -0.116 | -0.5405 | No | ||
| 130 | SLC26A9 | 11560 980 4067 | 14207 | -0.120 | -0.5414 | No | ||
| 131 | TNNC1 | 22058 | 14213 | -0.120 | -0.5407 | No | ||
| 132 | OMG | 5209 | 14306 | -0.130 | -0.5447 | No | ||
| 133 | BNC2 | 15850 | 14348 | -0.135 | -0.5459 | No | ||
| 134 | PDGFA | 5237 | 14395 | -0.140 | -0.5473 | No | ||
| 135 | TLE4 | 23720 3697 | 14521 | -0.154 | -0.5529 | No | ||
| 136 | ROBO3 | 9705 | 14560 | -0.159 | -0.5537 | No | ||
| 137 | AKAP2 | 8565 | 14590 | -0.164 | -0.5540 | No | ||
| 138 | PDLIM7 | 7434 | 14639 | -0.172 | -0.5553 | No | ||
| 139 | JMJD1C | 11531 19996 | 14762 | -0.195 | -0.5604 | No | ||
| 140 | BRUNOL4 | 2010 4229 | 15347 | -0.333 | -0.5895 | No | ||
| 141 | CLASP1 | 14159 | 15606 | -0.444 | -0.6001 | No | ||
| 142 | PDE4B | 9543 | 15644 | -0.464 | -0.5985 | No | ||
| 143 | SH3BGRL3 | 15722 | 15737 | -0.511 | -0.5995 | No | ||
| 144 | SESN3 | 19561 | 15821 | -0.567 | -0.5996 | No | ||
| 145 | TNKS1BP1 | 14970 | 15831 | -0.576 | -0.5957 | No | ||
| 146 | SLC35A2 | 24389 | 16068 | -0.792 | -0.6023 | Yes | ||
| 147 | CPNE1 | 11186 | 16094 | -0.816 | -0.5974 | Yes | ||
| 148 | SLC43A3 | 2824 14973 | 16269 | -0.992 | -0.5991 | Yes | ||
| 149 | HMGB1 | 9094 4855 | 16293 | -1.001 | -0.5926 | Yes | ||
| 150 | CASK | 24184 2618 2604 | 16298 | -1.002 | -0.5851 | Yes | ||
| 151 | TAPBP | 5625 | 16326 | -1.019 | -0.5787 | Yes | ||
| 152 | SLC25A28 | 23669 | 16371 | -1.074 | -0.5728 | Yes | ||
| 153 | PPM1L | 10811 6316 | 16587 | -1.330 | -0.5741 | Yes | ||
| 154 | ZDHHC12 | 14632 | 16693 | -1.493 | -0.5683 | Yes | ||
| 155 | THRAP1 | 20309 | 16781 | -1.630 | -0.5604 | Yes | ||
| 156 | TLN1 | 15899 | 16880 | -1.743 | -0.5522 | Yes | ||
| 157 | GADD45G | 21637 | 17072 | -2.018 | -0.5469 | Yes | ||
| 158 | NCOA3 | 5156 | 17080 | -2.031 | -0.5316 | Yes | ||
| 159 | RYBP | 17056 | 17143 | -2.143 | -0.5184 | Yes | ||
| 160 | SLC41A2 | 19666 | 17164 | -2.179 | -0.5026 | Yes | ||
| 161 | LRRC15 | 22617 | 17177 | -2.196 | -0.4863 | Yes | ||
| 162 | N4BP2 | 16837 | 17212 | -2.262 | -0.4706 | Yes | ||
| 163 | RUNX1 | 4481 | 17234 | -2.291 | -0.4540 | Yes | ||
| 164 | CTCF | 18490 | 17307 | -2.434 | -0.4391 | Yes | ||
| 165 | CREB3 | 16231 | 17406 | -2.658 | -0.4238 | Yes | ||
| 166 | CGGBP1 | 22724 | 17435 | -2.703 | -0.4044 | Yes | ||
| 167 | GSK3B | 22761 | 17596 | -3.078 | -0.3893 | Yes | ||
| 168 | DUSP10 | 4003 14016 | 17669 | -3.295 | -0.3677 | Yes | ||
| 169 | RNF122 | 18639 | 17726 | -3.441 | -0.3441 | Yes | ||
| 170 | SNX17 | 3495 16885 | 17760 | -3.518 | -0.3187 | Yes | ||
| 171 | HOXB4 | 20686 | 17827 | -3.678 | -0.2938 | Yes | ||
| 172 | PDE7A | 9544 1788 | 17861 | -3.771 | -0.2664 | Yes | ||
| 173 | DCTN4 | 12560 7474 | 17888 | -3.850 | -0.2380 | Yes | ||
| 174 | PBXIP1 | 6028 | 17892 | -3.863 | -0.2083 | Yes | ||
| 175 | BMP4 | 21863 | 17960 | -4.096 | -0.1802 | Yes | ||
| 176 | BCL11A | 4691 | 18039 | -4.409 | -0.1503 | Yes | ||
| 177 | TIGD2 | 17429 | 18061 | -4.507 | -0.1166 | Yes | ||
| 178 | SLC29A3 | 19758 | 18256 | -5.322 | -0.0859 | Yes | ||
| 179 | LMO4 | 15151 | 18404 | -6.210 | -0.0458 | Yes | ||
| 180 | SOX4 | 5479 | 18506 | -7.403 | 0.0060 | Yes |

