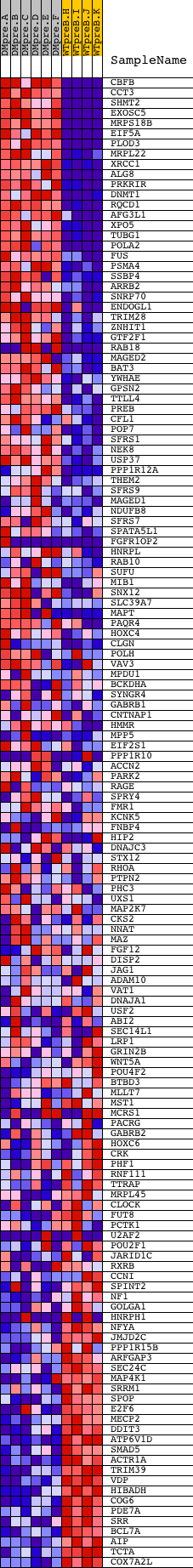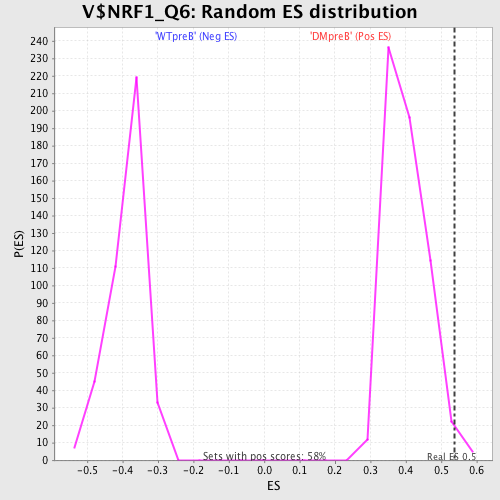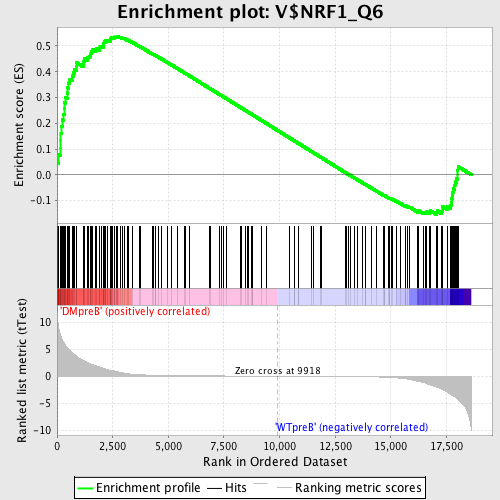
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | DMpreB |
| GeneSet | V$NRF1_Q6 |
| Enrichment Score (ES) | 0.5383303 |
| Normalized Enrichment Score (NES) | 1.3444027 |
| Nominal p-value | 0.011965812 |
| FDR q-value | 0.39130196 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CBFB | 3769 18499 | 0 | 12.040 | 0.0481 | Yes | ||
| 2 | CCT3 | 8709 | 81 | 8.641 | 0.0783 | Yes | ||
| 3 | SHMT2 | 3307 19601 | 169 | 7.348 | 0.1029 | Yes | ||
| 4 | EXOSC5 | 18334 | 171 | 7.337 | 0.1321 | Yes | ||
| 5 | MRPS18B | 7365 | 173 | 7.315 | 0.1613 | Yes | ||
| 6 | EIF5A | 11345 20379 6590 | 188 | 7.145 | 0.1891 | Yes | ||
| 7 | PLOD3 | 16666 | 229 | 6.720 | 0.2138 | Yes | ||
| 8 | MRPL22 | 10111 | 296 | 6.247 | 0.2351 | Yes | ||
| 9 | XRCC1 | 173 | 333 | 5.992 | 0.2571 | Yes | ||
| 10 | ALG8 | 11779 3912 2277 | 337 | 5.985 | 0.2809 | Yes | ||
| 11 | PRKRIR | 7834 | 385 | 5.592 | 0.3006 | Yes | ||
| 12 | DNMT1 | 19217 | 456 | 5.277 | 0.3179 | Yes | ||
| 13 | RQCD1 | 12205 | 475 | 5.201 | 0.3377 | Yes | ||
| 14 | AFG3L1 | 8547 4342 18427 3865 | 503 | 5.047 | 0.3564 | Yes | ||
| 15 | XPO5 | 7802 | 575 | 4.724 | 0.3714 | Yes | ||
| 16 | TUBG1 | 20662 | 674 | 4.381 | 0.3836 | Yes | ||
| 17 | POLA2 | 23988 | 733 | 4.174 | 0.3972 | Yes | ||
| 18 | FUS | 6135 | 784 | 3.989 | 0.4104 | Yes | ||
| 19 | PSMA4 | 11179 | 882 | 3.710 | 0.4200 | Yes | ||
| 20 | SSBP4 | 13368 | 886 | 3.699 | 0.4346 | Yes | ||
| 21 | ARRB2 | 20806 | 1172 | 2.927 | 0.4308 | Yes | ||
| 22 | SNRP70 | 1186 | 1194 | 2.877 | 0.4412 | Yes | ||
| 23 | ENDOGL1 | 3115 19275 | 1238 | 2.779 | 0.4499 | Yes | ||
| 24 | TRIM28 | 18389 | 1356 | 2.510 | 0.4536 | Yes | ||
| 25 | ZNHIT1 | 16334 | 1432 | 2.369 | 0.4590 | Yes | ||
| 26 | GTF2F1 | 13599 | 1480 | 2.279 | 0.4656 | Yes | ||
| 27 | RAB18 | 1972 5342 | 1519 | 2.233 | 0.4725 | Yes | ||
| 28 | MAGED2 | 24034 | 1539 | 2.220 | 0.4803 | Yes | ||
| 29 | BAT3 | 5896 | 1608 | 2.115 | 0.4851 | Yes | ||
| 30 | YWHAE | 20776 | 1727 | 1.923 | 0.4863 | Yes | ||
| 31 | GPSN2 | 18823 3868 | 1774 | 1.862 | 0.4913 | Yes | ||
| 32 | TTLL4 | 328 | 1915 | 1.654 | 0.4903 | Yes | ||
| 33 | PREB | 6948 3628 | 1926 | 1.631 | 0.4963 | Yes | ||
| 34 | CFL1 | 4516 | 1984 | 1.573 | 0.4995 | Yes | ||
| 35 | POP7 | 16329 | 2069 | 1.468 | 0.5008 | Yes | ||
| 36 | SFRS1 | 8492 | 2072 | 1.466 | 0.5065 | Yes | ||
| 37 | NEK8 | 20338 | 2090 | 1.449 | 0.5114 | Yes | ||
| 38 | USP37 | 13923 | 2107 | 1.433 | 0.5163 | Yes | ||
| 39 | PPP1R12A | 3354 19886 | 2187 | 1.326 | 0.5173 | Yes | ||
| 40 | THEM2 | 21512 | 2188 | 1.325 | 0.5226 | Yes | ||
| 41 | SFRS9 | 16731 | 2268 | 1.221 | 0.5232 | Yes | ||
| 42 | MAGED1 | 24108 | 2386 | 1.074 | 0.5211 | Yes | ||
| 43 | NDUFB8 | 7418 | 2404 | 1.062 | 0.5245 | Yes | ||
| 44 | SFRS7 | 22889 | 2416 | 1.053 | 0.5281 | Yes | ||
| 45 | SPATA5L1 | 10062 | 2429 | 1.042 | 0.5316 | Yes | ||
| 46 | FGFR1OP2 | 12544 12543 7457 | 2474 | 1.006 | 0.5332 | Yes | ||
| 47 | HNRPL | 18315 | 2564 | 0.961 | 0.5322 | Yes | ||
| 48 | RAB10 | 2090 21119 | 2585 | 0.939 | 0.5349 | Yes | ||
| 49 | SUFU | 6283 | 2651 | 0.860 | 0.5348 | Yes | ||
| 50 | MIB1 | 5908 | 2705 | 0.812 | 0.5352 | Yes | ||
| 51 | SNX12 | 7061 | 2708 | 0.811 | 0.5383 | Yes | ||
| 52 | SLC39A7 | 23022 | 2832 | 0.703 | 0.5345 | No | ||
| 53 | MAPT | 9420 1347 | 2946 | 0.604 | 0.5308 | No | ||
| 54 | PAQR4 | 23105 | 3014 | 0.549 | 0.5293 | No | ||
| 55 | HOXC4 | 4864 | 3152 | 0.460 | 0.5238 | No | ||
| 56 | CLGN | 4528 | 3211 | 0.437 | 0.5224 | No | ||
| 57 | POLH | 22966 | 3377 | 0.361 | 0.5149 | No | ||
| 58 | VAV3 | 1774 1848 15443 | 3695 | 0.275 | 0.4988 | No | ||
| 59 | MPDU1 | 20386 | 3762 | 0.258 | 0.4963 | No | ||
| 60 | BCKDHA | 17925 | 4286 | 0.175 | 0.4686 | No | ||
| 61 | SYNGR4 | 17825 | 4350 | 0.166 | 0.4659 | No | ||
| 62 | GABRB1 | 16832 | 4410 | 0.160 | 0.4633 | No | ||
| 63 | CNTNAP1 | 11987 | 4429 | 0.158 | 0.4630 | No | ||
| 64 | HMMR | 20493 1243 | 4548 | 0.146 | 0.4572 | No | ||
| 65 | MPP5 | 21230 7078 | 4682 | 0.134 | 0.4505 | No | ||
| 66 | EIF2S1 | 4658 | 4969 | 0.114 | 0.4355 | No | ||
| 67 | PPP1R10 | 6972 11960 6971 | 5145 | 0.105 | 0.4264 | No | ||
| 68 | ACCN2 | 22363 | 5425 | 0.089 | 0.4117 | No | ||
| 69 | PARK2 | 6944 11939 1499 23385 | 5744 | 0.075 | 0.3948 | No | ||
| 70 | RAGE | 6465 2056 2165 | 5782 | 0.074 | 0.3931 | No | ||
| 71 | SPRY4 | 23448 | 5960 | 0.068 | 0.3838 | No | ||
| 72 | FMR1 | 24321 | 6848 | 0.046 | 0.3359 | No | ||
| 73 | KCNK5 | 4946 9212 | 6873 | 0.045 | 0.3348 | No | ||
| 74 | FNBP4 | 14956 2671 | 7287 | 0.037 | 0.3126 | No | ||
| 75 | HIP2 | 3626 3485 16838 | 7403 | 0.035 | 0.3065 | No | ||
| 76 | DNAJC3 | 21934 | 7477 | 0.033 | 0.3027 | No | ||
| 77 | STX12 | 4140 | 7622 | 0.031 | 0.2950 | No | ||
| 78 | RHOA | 8624 4409 4410 | 8261 | 0.021 | 0.2605 | No | ||
| 79 | PTPN2 | 23414 | 8306 | 0.020 | 0.2582 | No | ||
| 80 | PHC3 | 6313 | 8445 | 0.019 | 0.2508 | No | ||
| 81 | UXS1 | 13965 | 8561 | 0.017 | 0.2447 | No | ||
| 82 | MAP2K7 | 6453 | 8613 | 0.016 | 0.2420 | No | ||
| 83 | CKS2 | 7264 | 8727 | 0.015 | 0.2359 | No | ||
| 84 | NNAT | 2764 5180 | 8773 | 0.014 | 0.2336 | No | ||
| 85 | MAZ | 1327 17623 | 8787 | 0.014 | 0.2329 | No | ||
| 86 | FGF12 | 1723 22621 | 9206 | 0.009 | 0.2103 | No | ||
| 87 | DISP2 | 5652 | 9389 | 0.007 | 0.2005 | No | ||
| 88 | JAG1 | 14415 | 10441 | -0.006 | 0.1436 | No | ||
| 89 | ADAM10 | 4336 | 10680 | -0.010 | 0.1308 | No | ||
| 90 | VAT1 | 11253 | 10688 | -0.010 | 0.1304 | No | ||
| 91 | DNAJA1 | 4878 16242 2345 | 10832 | -0.012 | 0.1227 | No | ||
| 92 | USF2 | 10262 5833 10263 | 10845 | -0.012 | 0.1221 | No | ||
| 93 | ABI2 | 4037 6830 | 11434 | -0.020 | 0.0904 | No | ||
| 94 | SEC14L1 | 1212 20583 | 11529 | -0.022 | 0.0854 | No | ||
| 95 | LRP1 | 9284 | 11848 | -0.027 | 0.0683 | No | ||
| 96 | GRIN2B | 16957 | 11902 | -0.028 | 0.0655 | No | ||
| 97 | WNT5A | 22066 5880 | 12962 | -0.053 | 0.0084 | No | ||
| 98 | POU4F2 | 9603 5276 | 13022 | -0.055 | 0.0054 | No | ||
| 99 | BTBD3 | 10457 2975 6007 6006 | 13116 | -0.058 | 0.0006 | No | ||
| 100 | MLLT7 | 24278 | 13187 | -0.061 | -0.0029 | No | ||
| 101 | MST1 | 9089 | 13347 | -0.067 | -0.0113 | No | ||
| 102 | MCRS1 | 6961 6960 6962 | 13486 | -0.072 | -0.0185 | No | ||
| 103 | PACRG | 23127 | 13713 | -0.084 | -0.0304 | No | ||
| 104 | GABRB2 | 20922 | 13846 | -0.091 | -0.0371 | No | ||
| 105 | HOXC6 | 22339 2302 | 13878 | -0.093 | -0.0384 | No | ||
| 106 | CRK | 4559 1249 | 14137 | -0.112 | -0.0520 | No | ||
| 107 | PHF1 | 23324 1551 | 14350 | -0.135 | -0.0629 | No | ||
| 108 | RNF111 | 13555 8218 13556 | 14676 | -0.179 | -0.0798 | No | ||
| 109 | TTRAP | 21692 | 14698 | -0.182 | -0.0802 | No | ||
| 110 | MRPL45 | 20680 | 14727 | -0.188 | -0.0810 | No | ||
| 111 | CLOCK | 8749 16507 4530 | 14905 | -0.220 | -0.0897 | No | ||
| 112 | FUT8 | 2163 21233 | 14949 | -0.227 | -0.0911 | No | ||
| 113 | PCTK1 | 24369 | 14960 | -0.230 | -0.0907 | No | ||
| 114 | U2AF2 | 5813 5812 5814 | 15014 | -0.242 | -0.0926 | No | ||
| 115 | POU2F1 | 5275 3989 4065 4010 | 15082 | -0.259 | -0.0952 | No | ||
| 116 | JARID1C | 5460 2563 | 15264 | -0.308 | -0.1038 | No | ||
| 117 | RXRB | 23285 9768 | 15412 | -0.356 | -0.1103 | No | ||
| 118 | CCNI | 4493 | 15639 | -0.460 | -0.1207 | No | ||
| 119 | SPINT2 | 4128 17899 | 15643 | -0.462 | -0.1190 | No | ||
| 120 | NF1 | 5165 | 15767 | -0.530 | -0.1236 | No | ||
| 121 | GOLGA1 | 14595 | 15843 | -0.582 | -0.1253 | No | ||
| 122 | HNRPH1 | 12222 1486 1220 7202 7201 | 16220 | -0.944 | -0.1419 | No | ||
| 123 | NFYA | 5172 | 16265 | -0.988 | -0.1403 | No | ||
| 124 | JMJD2C | 16188 | 16475 | -1.189 | -0.1469 | No | ||
| 125 | PPP1R15B | 4254 14135 | 16553 | -1.276 | -0.1460 | No | ||
| 126 | ARFGAP3 | 22183 | 16623 | -1.394 | -0.1441 | No | ||
| 127 | SEC24C | 22086 | 16754 | -1.591 | -0.1448 | No | ||
| 128 | MAP4K1 | 18313 | 16775 | -1.624 | -0.1394 | No | ||
| 129 | SRRM1 | 6958 | 17048 | -1.978 | -0.1462 | No | ||
| 130 | SPOP | 5498 | 17077 | -2.024 | -0.1397 | No | ||
| 131 | E2F6 | 6925 6926 11920 | 17291 | -2.401 | -0.1416 | No | ||
| 132 | MECP2 | 5088 | 17315 | -2.463 | -0.1330 | No | ||
| 133 | DDIT3 | 19857 | 17317 | -2.466 | -0.1232 | No | ||
| 134 | ATP6V1D | 7872 | 17533 | -2.931 | -0.1232 | No | ||
| 135 | SMAD5 | 21621 | 17677 | -3.317 | -0.1177 | No | ||
| 136 | ACTR1A | 23653 | 17730 | -3.449 | -0.1067 | No | ||
| 137 | TRIM39 | 22994 | 17740 | -3.476 | -0.0933 | No | ||
| 138 | VDP | 16794 | 17756 | -3.510 | -0.0801 | No | ||
| 139 | HIBADH | 17144 | 17782 | -3.569 | -0.0672 | No | ||
| 140 | COG6 | 15345 | 17814 | -3.654 | -0.0543 | No | ||
| 141 | PDE7A | 9544 1788 | 17861 | -3.771 | -0.0417 | No | ||
| 142 | SRR | 6566 11328 | 17890 | -3.852 | -0.0278 | No | ||
| 143 | BCL7A | 8063 | 17938 | -4.024 | -0.0143 | No | ||
| 144 | AIP | 23955 | 18000 | -4.223 | -0.0008 | No | ||
| 145 | TCTA | 4167 | 18001 | -4.223 | 0.0161 | No | ||
| 146 | COX7A2L | 9819 | 18019 | -4.290 | 0.0323 | No |