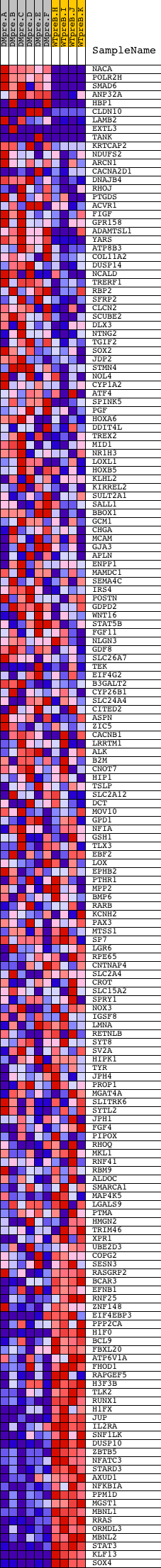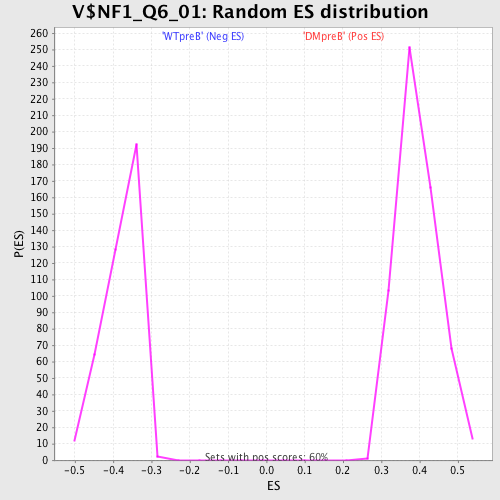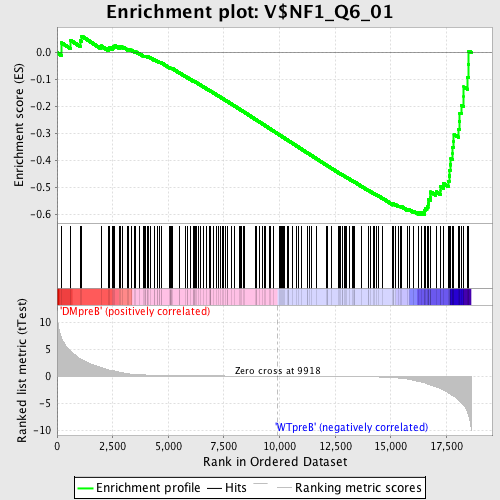
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | V$NF1_Q6_01 |
| Enrichment Score (ES) | -0.6001914 |
| Normalized Enrichment Score (NES) | -1.5861269 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04739025 |
| FWER p-Value | 0.35 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NACA | 9444 5147 | 205 | 6.962 | 0.0362 | No | ||
| 2 | POLR2H | 10888 | 611 | 4.591 | 0.0454 | No | ||
| 3 | SMAD6 | 19083 | 1038 | 3.214 | 0.0441 | No | ||
| 4 | ANP32A | 8592 4387 19415 | 1107 | 3.050 | 0.0612 | No | ||
| 5 | HBP1 | 2048 2162 21086 | 1975 | 1.578 | 0.0249 | No | ||
| 6 | CLDN10 | 21935 2785 2601 2720 7188 | 2307 | 1.167 | 0.0148 | No | ||
| 7 | LAMB2 | 9265 | 2342 | 1.125 | 0.0206 | No | ||
| 8 | EXTL3 | 21785 21786 3088 | 2505 | 0.998 | 0.0186 | No | ||
| 9 | TANK | 2839 15006 2729 | 2541 | 0.980 | 0.0234 | No | ||
| 10 | KRTCAP2 | 15541 | 2600 | 0.920 | 0.0265 | No | ||
| 11 | NDUFS2 | 4089 13760 | 2812 | 0.719 | 0.0199 | No | ||
| 12 | ARCN1 | 19143 | 2833 | 0.701 | 0.0236 | No | ||
| 13 | CACNA2D1 | 8676 | 2958 | 0.591 | 0.0209 | No | ||
| 14 | DNAJB4 | 12451 1916 1832 | 3178 | 0.451 | 0.0121 | No | ||
| 15 | RHOJ | 21241 | 3212 | 0.436 | 0.0133 | No | ||
| 16 | PTGDS | 14658 | 3326 | 0.381 | 0.0097 | No | ||
| 17 | ACVR1 | 4334 | 3479 | 0.334 | 0.0037 | No | ||
| 18 | FIGF | 24212 | 3516 | 0.322 | 0.0040 | No | ||
| 19 | GPR158 | 149 | 3693 | 0.275 | -0.0037 | No | ||
| 20 | ADAMTSL1 | 16181 | 3896 | 0.231 | -0.0131 | No | ||
| 21 | YARS | 16071 | 3945 | 0.223 | -0.0142 | No | ||
| 22 | ATP8B3 | 12513 7426 19693 | 3977 | 0.217 | -0.0144 | No | ||
| 23 | COL11A2 | 23286 | 3982 | 0.216 | -0.0131 | No | ||
| 24 | DUSP14 | 12124 | 4042 | 0.208 | -0.0149 | No | ||
| 25 | NCALD | 22310 | 4083 | 0.202 | -0.0157 | No | ||
| 26 | TRERF1 | 1546 23213 | 4124 | 0.196 | -0.0166 | No | ||
| 27 | RBP2 | 19347 | 4187 | 0.188 | -0.0186 | No | ||
| 28 | SFRP2 | 15564 | 4360 | 0.164 | -0.0269 | No | ||
| 29 | CLCN2 | 22637 | 4385 | 0.162 | -0.0271 | No | ||
| 30 | SCUBE2 | 1879 17678 | 4512 | 0.149 | -0.0329 | No | ||
| 31 | DLX3 | 20697 | 4533 | 0.147 | -0.0330 | No | ||
| 32 | NTNG2 | 2731 9330 | 4608 | 0.140 | -0.0360 | No | ||
| 33 | TGIF2 | 6013 6012 | 4620 | 0.139 | -0.0357 | No | ||
| 34 | SOX2 | 9849 15612 | 4683 | 0.134 | -0.0381 | No | ||
| 35 | JDP2 | 21201 | 4693 | 0.133 | -0.0377 | No | ||
| 36 | STMN4 | 21977 | 5048 | 0.110 | -0.0562 | No | ||
| 37 | NOL4 | 2028 11432 | 5084 | 0.108 | -0.0573 | No | ||
| 38 | CYP1A2 | 19097 | 5092 | 0.107 | -0.0570 | No | ||
| 39 | ATF4 | 22417 2239 | 5109 | 0.106 | -0.0571 | No | ||
| 40 | SPINK5 | 23569 | 5150 | 0.105 | -0.0586 | No | ||
| 41 | PGF | 21020 | 5155 | 0.104 | -0.0581 | No | ||
| 42 | HOXA6 | 17150 | 5207 | 0.101 | -0.0602 | No | ||
| 43 | DDIT4L | 15410 | 5481 | 0.087 | -0.0744 | No | ||
| 44 | TREX2 | 24137 | 5519 | 0.085 | -0.0758 | No | ||
| 45 | MID1 | 5097 5098 | 5763 | 0.074 | -0.0885 | No | ||
| 46 | NR1H3 | 10258 2787 | 5876 | 0.070 | -0.0941 | No | ||
| 47 | LOXL1 | 5002 | 5877 | 0.070 | -0.0936 | No | ||
| 48 | HOXB5 | 20687 | 5990 | 0.067 | -0.0992 | No | ||
| 49 | KLHL2 | 13381 | 6112 | 0.063 | -0.1054 | No | ||
| 50 | KIRREL2 | 17889 3922 | 6116 | 0.063 | -0.1051 | No | ||
| 51 | SULT2A1 | 5527 | 6165 | 0.062 | -0.1073 | No | ||
| 52 | SALL1 | 18796 12207 | 6190 | 0.061 | -0.1082 | No | ||
| 53 | BBOX1 | 5010 9290 | 6199 | 0.060 | -0.1082 | No | ||
| 54 | GCM1 | 19372 | 6273 | 0.058 | -0.1118 | No | ||
| 55 | CHGA | 21178 2151 | 6339 | 0.057 | -0.1149 | No | ||
| 56 | MCAM | 3109 19479 | 6452 | 0.054 | -0.1206 | No | ||
| 57 | GJA3 | 21807 | 6598 | 0.051 | -0.1281 | No | ||
| 58 | APLN | 24164 | 6724 | 0.048 | -0.1346 | No | ||
| 59 | ENPP1 | 19804 | 6838 | 0.046 | -0.1404 | No | ||
| 60 | MAMDC1 | 21054 6798 | 6880 | 0.045 | -0.1423 | No | ||
| 61 | SEMA4C | 9799 | 6892 | 0.045 | -0.1426 | No | ||
| 62 | IRS4 | 9183 4926 | 6907 | 0.044 | -0.1431 | No | ||
| 63 | POSTN | 1801 11927 | 7047 | 0.041 | -0.1503 | No | ||
| 64 | GDPD2 | 24280 | 7172 | 0.039 | -0.1568 | No | ||
| 65 | WNT16 | 17512 1152 | 7261 | 0.037 | -0.1613 | No | ||
| 66 | STAT5B | 20222 | 7325 | 0.036 | -0.1645 | No | ||
| 67 | FGF11 | 20383 | 7450 | 0.034 | -0.1710 | No | ||
| 68 | NLGN3 | 10878 | 7466 | 0.033 | -0.1715 | No | ||
| 69 | GDF8 | 9411 | 7555 | 0.032 | -0.1761 | No | ||
| 70 | SLC26A7 | 5538 2372 | 7638 | 0.030 | -0.1803 | No | ||
| 71 | TEK | 16175 | 7666 | 0.030 | -0.1816 | No | ||
| 72 | EIF4G2 | 1908 8892 | 7830 | 0.027 | -0.1903 | No | ||
| 73 | B3GALT2 | 6498 | 7968 | 0.025 | -0.1975 | No | ||
| 74 | CYP26B1 | 17093 | 8191 | 0.022 | -0.2094 | No | ||
| 75 | SLC24A4 | 6223 | 8245 | 0.021 | -0.2121 | No | ||
| 76 | CITED2 | 5118 14477 | 8285 | 0.021 | -0.2141 | No | ||
| 77 | ASPN | 21644 | 8376 | 0.019 | -0.2189 | No | ||
| 78 | ZIC5 | 12268 | 8443 | 0.019 | -0.2223 | No | ||
| 79 | CACNB1 | 1443 1304 | 8932 | 0.012 | -0.2487 | No | ||
| 80 | LRRTM1 | 17408 | 8945 | 0.012 | -0.2492 | No | ||
| 81 | ALK | 8574 | 8962 | 0.012 | -0.2500 | No | ||
| 82 | B2M | 72 14882 | 9089 | 0.011 | -0.2568 | No | ||
| 83 | CNOT7 | 3787 9600 | 9239 | 0.009 | -0.2648 | No | ||
| 84 | HIP1 | 5676 | 9311 | 0.008 | -0.2686 | No | ||
| 85 | TSLP | 23604 1955 | 9354 | 0.007 | -0.2708 | No | ||
| 86 | SLC2A12 | 20073 | 9543 | 0.005 | -0.2810 | No | ||
| 87 | DCT | 4603 21723 | 9579 | 0.004 | -0.2829 | No | ||
| 88 | MOV10 | 9403 9402 5114 | 9711 | 0.003 | -0.2900 | No | ||
| 89 | GPD1 | 22361 | 10004 | -0.001 | -0.3058 | No | ||
| 90 | NFIA | 16172 5170 | 10033 | -0.002 | -0.3073 | No | ||
| 91 | GSH1 | 16622 | 10075 | -0.002 | -0.3095 | No | ||
| 92 | TLX3 | 20501 | 10139 | -0.003 | -0.3129 | No | ||
| 93 | EBF2 | 21975 | 10186 | -0.003 | -0.3154 | No | ||
| 94 | LOX | 23438 | 10225 | -0.004 | -0.3174 | No | ||
| 95 | EPHB2 | 4675 2440 8910 | 10347 | -0.005 | -0.3239 | No | ||
| 96 | PTHR1 | 5318 18985 | 10406 | -0.006 | -0.3270 | No | ||
| 97 | MPP2 | 20209 | 10408 | -0.006 | -0.3271 | No | ||
| 98 | BMP6 | 8659 | 10582 | -0.008 | -0.3364 | No | ||
| 99 | RARB | 3011 21915 | 10781 | -0.011 | -0.3470 | No | ||
| 100 | KCNH2 | 16592 | 10827 | -0.012 | -0.3494 | No | ||
| 101 | PAX3 | 13910 | 10987 | -0.014 | -0.3579 | No | ||
| 102 | MTSS1 | 5585 22285 | 11237 | -0.018 | -0.3713 | No | ||
| 103 | SP7 | 22110 | 11247 | -0.018 | -0.3717 | No | ||
| 104 | LGR6 | 11623 977 | 11332 | -0.019 | -0.3761 | No | ||
| 105 | RPE65 | 15382 | 11440 | -0.020 | -0.3818 | No | ||
| 106 | CNTNAP4 | 18462 3857 | 11666 | -0.024 | -0.3938 | No | ||
| 107 | SLC2A4 | 20380 | 12102 | -0.032 | -0.4172 | No | ||
| 108 | CROT | 13165 | 12151 | -0.033 | -0.4196 | No | ||
| 109 | SLC15A2 | 22600 | 12152 | -0.033 | -0.4193 | No | ||
| 110 | SPRY1 | 15606 1901 | 12336 | -0.037 | -0.4290 | No | ||
| 111 | NOX3 | 10322 | 12632 | -0.045 | -0.4447 | No | ||
| 112 | IGSF8 | 14048 | 12706 | -0.047 | -0.4483 | No | ||
| 113 | LMNA | 4999 15290 1778 9280 | 12737 | -0.047 | -0.4496 | No | ||
| 114 | RETNLB | 12178 | 12817 | -0.049 | -0.4536 | No | ||
| 115 | SYT8 | 18006 | 12820 | -0.049 | -0.4534 | No | ||
| 116 | SV2A | 15498 12250 | 12915 | -0.052 | -0.4581 | No | ||
| 117 | HIPK1 | 4851 | 12960 | -0.053 | -0.4601 | No | ||
| 118 | TYR | 17763 | 13007 | -0.054 | -0.4623 | No | ||
| 119 | JPH4 | 21826 | 13126 | -0.058 | -0.4683 | No | ||
| 120 | PROP1 | 20472 | 13289 | -0.064 | -0.4766 | No | ||
| 121 | MGAT4A | 13977 | 13305 | -0.065 | -0.4770 | No | ||
| 122 | SLITRK6 | 21724 | 13367 | -0.067 | -0.4798 | No | ||
| 123 | SYTL2 | 18190 1539 3730 | 13377 | -0.068 | -0.4799 | No | ||
| 124 | JPH1 | 13994 | 13674 | -0.082 | -0.4954 | No | ||
| 125 | FGF4 | 8964 | 13993 | -0.101 | -0.5119 | No | ||
| 126 | PIPOX | 20340 | 14001 | -0.102 | -0.5116 | No | ||
| 127 | RHOQ | 23142 | 14100 | -0.110 | -0.5162 | No | ||
| 128 | MKL1 | 5864 | 14199 | -0.119 | -0.5207 | No | ||
| 129 | RNF41 | 12553 7464 | 14282 | -0.127 | -0.5243 | No | ||
| 130 | RBM9 | 2276 13528 | 14358 | -0.136 | -0.5274 | No | ||
| 131 | ALDOC | 20759 311 | 14449 | -0.146 | -0.5313 | No | ||
| 132 | SMARCA1 | 24165 | 14638 | -0.172 | -0.5403 | No | ||
| 133 | MAP4K5 | 11853 21048 2101 2142 | 15096 | -0.263 | -0.5633 | No | ||
| 134 | LGALS9 | 1441 20332 1200 | 15103 | -0.265 | -0.5618 | No | ||
| 135 | PTMA | 9657 | 15118 | -0.268 | -0.5608 | No | ||
| 136 | HMGN2 | 9095 | 15229 | -0.297 | -0.5647 | No | ||
| 137 | TRIM46 | 1779 15281 | 15356 | -0.337 | -0.5693 | No | ||
| 138 | XPR1 | 5383 | 15417 | -0.358 | -0.5701 | No | ||
| 139 | UBE2D3 | 7253 | 15487 | -0.387 | -0.5712 | No | ||
| 140 | COPG2 | 17193 | 15727 | -0.506 | -0.5807 | No | ||
| 141 | SESN3 | 19561 | 15821 | -0.567 | -0.5819 | No | ||
| 142 | RASGRP2 | 9693 | 16026 | -0.747 | -0.5879 | No | ||
| 143 | BCAR3 | 15432 | 16234 | -0.966 | -0.5926 | Yes | ||
| 144 | EFNB1 | 24285 | 16362 | -1.056 | -0.5923 | Yes | ||
| 145 | RNF25 | 13922 3929 | 16509 | -1.222 | -0.5919 | Yes | ||
| 146 | ZNF148 | 5949 | 16515 | -1.233 | -0.5838 | Yes | ||
| 147 | EIF4EBP3 | 8429 | 16544 | -1.267 | -0.5767 | Yes | ||
| 148 | PPP2CA | 20890 | 16627 | -1.403 | -0.5716 | Yes | ||
| 149 | H1F0 | 22428 | 16674 | -1.457 | -0.5642 | Yes | ||
| 150 | BCL9 | 15232 | 16684 | -1.483 | -0.5546 | Yes | ||
| 151 | FBXL20 | 13010 | 16688 | -1.488 | -0.5447 | Yes | ||
| 152 | ATP6V1A | 8638 | 16784 | -1.640 | -0.5387 | Yes | ||
| 153 | FHOD1 | 18772 | 16792 | -1.656 | -0.5278 | Yes | ||
| 154 | RAPGEF5 | 5739 10155 21123 | 16794 | -1.661 | -0.5166 | Yes | ||
| 155 | H3F3B | 4839 | 17030 | -1.944 | -0.5161 | Yes | ||
| 156 | TLK2 | 20628 | 17231 | -2.287 | -0.5115 | Yes | ||
| 157 | RUNX1 | 4481 | 17234 | -2.291 | -0.4960 | Yes | ||
| 158 | H1FX | 10835 | 17374 | -2.596 | -0.4859 | Yes | ||
| 159 | JUP | 4937 | 17591 | -3.058 | -0.4769 | Yes | ||
| 160 | IL2RA | 4918 | 17622 | -3.145 | -0.4571 | Yes | ||
| 161 | SNF1LK | 23033 | 17648 | -3.225 | -0.4366 | Yes | ||
| 162 | DUSP10 | 4003 14016 | 17669 | -3.295 | -0.4153 | Yes | ||
| 163 | ZBTB5 | 15891 | 17699 | -3.357 | -0.3940 | Yes | ||
| 164 | NFATC3 | 5169 | 17772 | -3.544 | -0.3739 | Yes | ||
| 165 | STARD3 | 20675 | 17778 | -3.553 | -0.3500 | Yes | ||
| 166 | AXUD1 | 18967 | 17830 | -3.686 | -0.3277 | Yes | ||
| 167 | NFKBIA | 21065 | 17838 | -3.699 | -0.3030 | Yes | ||
| 168 | PPM1D | 20721 | 18053 | -4.474 | -0.2842 | Yes | ||
| 169 | MGST1 | 17254 | 18083 | -4.575 | -0.2547 | Yes | ||
| 170 | MBNL1 | 1921 15582 | 18100 | -4.631 | -0.2241 | Yes | ||
| 171 | RRAS | 18256 | 18165 | -4.858 | -0.1946 | Yes | ||
| 172 | ORMDL3 | 12386 | 18280 | -5.413 | -0.1640 | Yes | ||
| 173 | MBNL2 | 21932 | 18284 | -5.431 | -0.1273 | Yes | ||
| 174 | STAT3 | 5525 9906 | 18455 | -6.760 | -0.0906 | Yes | ||
| 175 | KLF13 | 17807 | 18488 | -7.203 | -0.0434 | Yes | ||
| 176 | SOX4 | 5479 | 18506 | -7.403 | 0.0060 | Yes |