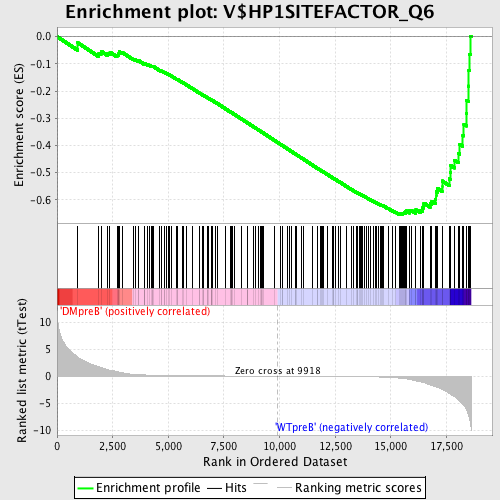
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | V$HP1SITEFACTOR_Q6 |
| Enrichment Score (ES) | -0.65398145 |
| Normalized Enrichment Score (NES) | -1.7295802 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.014807959 |
| FWER p-Value | 0.03 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MRPL28 | 23332 | 929 | 3.542 | -0.0227 | No | ||
| 2 | PRICKLE1 | 8379 | 1877 | 1.705 | -0.0606 | No | ||
| 3 | HBP1 | 2048 2162 21086 | 1975 | 1.578 | -0.0536 | No | ||
| 4 | CDC2L2 | 15964 2419 | 2282 | 1.202 | -0.0607 | No | ||
| 5 | CYFIP2 | 20485 8045 | 2376 | 1.088 | -0.0573 | No | ||
| 6 | EML4 | 8122 | 2695 | 0.823 | -0.0681 | No | ||
| 7 | COL4A3BP | 3221 7522 | 2750 | 0.764 | -0.0650 | No | ||
| 8 | RDH10 | 8272 | 2774 | 0.747 | -0.0604 | No | ||
| 9 | SFRS2 | 9807 20136 | 2789 | 0.733 | -0.0555 | No | ||
| 10 | CACNA2D3 | 8677 | 2930 | 0.616 | -0.0583 | No | ||
| 11 | PAPD1 | 12533 7442 | 3433 | 0.346 | -0.0827 | No | ||
| 12 | NFIX | 9458 | 3501 | 0.326 | -0.0838 | No | ||
| 13 | FBN2 | 23433 | 3654 | 0.286 | -0.0898 | No | ||
| 14 | ACCN4 | 10795 | 3667 | 0.281 | -0.0883 | No | ||
| 15 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 3930 | 0.226 | -0.1007 | No | ||
| 16 | FOXP2 | 4312 17523 8523 | 3935 | 0.225 | -0.0992 | No | ||
| 17 | SYNCRIP | 3078 3035 3107 | 3939 | 0.225 | -0.0976 | No | ||
| 18 | NCALD | 22310 | 4083 | 0.202 | -0.1038 | No | ||
| 19 | PHF16 | 11795 6901 | 4160 | 0.191 | -0.1064 | No | ||
| 20 | CHRDL1 | 8180 13504 | 4255 | 0.178 | -0.1101 | No | ||
| 21 | CS | 19839 | 4271 | 0.177 | -0.1095 | No | ||
| 22 | SMAD1 | 18831 922 | 4300 | 0.173 | -0.1097 | No | ||
| 23 | PJA1 | 9572 | 4319 | 0.171 | -0.1093 | No | ||
| 24 | HOXA9 | 17147 9109 1024 | 4614 | 0.139 | -0.1242 | No | ||
| 25 | SOX2 | 9849 15612 | 4683 | 0.134 | -0.1268 | No | ||
| 26 | RBP4 | 23686 | 4690 | 0.134 | -0.1261 | No | ||
| 27 | BARHL1 | 14634 | 4811 | 0.124 | -0.1316 | No | ||
| 28 | HOXB7 | 666 9110 | 4836 | 0.122 | -0.1320 | No | ||
| 29 | HOXB3 | 20685 | 4909 | 0.118 | -0.1350 | No | ||
| 30 | LHX6 | 9275 2914 | 5003 | 0.112 | -0.1391 | No | ||
| 31 | VLDLR | 5858 3763 912 3682 | 5044 | 0.110 | -0.1404 | No | ||
| 32 | DLX1 | 14988 | 5151 | 0.105 | -0.1454 | No | ||
| 33 | XPO1 | 4172 | 5363 | 0.093 | -0.1561 | No | ||
| 34 | NR5A2 | 4105 13817 | 5404 | 0.091 | -0.1575 | No | ||
| 35 | PPP1CB | 5282 3529 3504 | 5421 | 0.090 | -0.1577 | No | ||
| 36 | PROK2 | 17057 | 5630 | 0.080 | -0.1683 | No | ||
| 37 | CNTFR | 2515 15906 | 5689 | 0.077 | -0.1709 | No | ||
| 38 | ADAM28 | 3484 8868 4645 3113 | 5818 | 0.072 | -0.1772 | No | ||
| 39 | STC1 | 21974 | 6065 | 0.064 | -0.1901 | No | ||
| 40 | BHLHB3 | 16935 | 6401 | 0.055 | -0.2078 | No | ||
| 41 | IGF1 | 3352 9156 3409 | 6411 | 0.055 | -0.2079 | No | ||
| 42 | TYRO3 | 5811 | 6547 | 0.052 | -0.2148 | No | ||
| 43 | AFP | 16805 | 6562 | 0.052 | -0.2151 | No | ||
| 44 | ZIC1 | 19040 | 6572 | 0.052 | -0.2152 | No | ||
| 45 | ANPEP | 17784 | 6755 | 0.047 | -0.2247 | No | ||
| 46 | MYH2 | 20838 | 6769 | 0.047 | -0.2250 | No | ||
| 47 | SEZ6 | 20764 | 6822 | 0.046 | -0.2275 | No | ||
| 48 | ATP6V1E2 | 22880 | 6939 | 0.043 | -0.2334 | No | ||
| 49 | DPYSL2 | 3122 4561 8787 | 6945 | 0.043 | -0.2334 | No | ||
| 50 | GPR27 | 13703 | 6953 | 0.043 | -0.2334 | No | ||
| 51 | ADAM11 | 4340 1432 | 6989 | 0.042 | -0.2350 | No | ||
| 52 | MYCBP | 7093 | 7139 | 0.040 | -0.2427 | No | ||
| 53 | FMO3 | 13785 | 7202 | 0.038 | -0.2458 | No | ||
| 54 | GDF8 | 9411 | 7555 | 0.032 | -0.2646 | No | ||
| 55 | HSD17B2 | 18452 | 7814 | 0.028 | -0.2784 | No | ||
| 56 | IBRDC2 | 5747 | 7828 | 0.027 | -0.2789 | No | ||
| 57 | EIF4G2 | 1908 8892 | 7830 | 0.027 | -0.2787 | No | ||
| 58 | SNCAIP | 12589 | 7868 | 0.027 | -0.2805 | No | ||
| 59 | DIO2 | 21014 2159 | 7974 | 0.025 | -0.2860 | No | ||
| 60 | CITED2 | 5118 14477 | 8285 | 0.021 | -0.3026 | No | ||
| 61 | GPC3 | 24151 | 8537 | 0.017 | -0.3161 | No | ||
| 62 | POU4F1 | 21727 | 8564 | 0.017 | -0.3174 | No | ||
| 63 | CA4 | 20723 | 8578 | 0.017 | -0.3180 | No | ||
| 64 | LHX1 | 4995 | 8838 | 0.014 | -0.3319 | No | ||
| 65 | BMP5 | 19376 | 8936 | 0.012 | -0.3371 | No | ||
| 66 | HOXD3 | 9115 | 9063 | 0.011 | -0.3438 | No | ||
| 67 | ATP2B3 | 24308 11557 | 9149 | 0.010 | -0.3483 | No | ||
| 68 | NRXN3 | 2153 5196 | 9166 | 0.010 | -0.3491 | No | ||
| 69 | NPAS3 | 2077 21273 | 9186 | 0.009 | -0.3501 | No | ||
| 70 | USP13 | 13047 | 9253 | 0.008 | -0.3536 | No | ||
| 71 | EVI1 | 4689 8926 | 9266 | 0.008 | -0.3542 | No | ||
| 72 | GPR110 | 23234 | 9749 | 0.002 | -0.3803 | No | ||
| 73 | NOG | 13663 | 10030 | -0.002 | -0.3954 | No | ||
| 74 | DRD3 | 22750 | 10108 | -0.002 | -0.3996 | No | ||
| 75 | CRYGD | 8797 | 10140 | -0.003 | -0.4013 | No | ||
| 76 | FAP | 14577 | 10349 | -0.005 | -0.4125 | No | ||
| 77 | JAG1 | 14415 | 10441 | -0.006 | -0.4174 | No | ||
| 78 | SCHIP1 | 15569 | 10523 | -0.007 | -0.4217 | No | ||
| 79 | HOXA4 | 17148 | 10724 | -0.010 | -0.4325 | No | ||
| 80 | SPARC | 20444 | 10728 | -0.010 | -0.4325 | No | ||
| 81 | GPR156 | 10753 22763 | 10762 | -0.011 | -0.4342 | No | ||
| 82 | PITX2 | 15424 1878 | 10992 | -0.014 | -0.4465 | No | ||
| 83 | PCDHA7 | 8789 | 11054 | -0.015 | -0.4497 | No | ||
| 84 | GCNT2 | 9168 3166 3243 21661 3270 | 11461 | -0.021 | -0.4716 | No | ||
| 85 | FOXD3 | 9088 | 11683 | -0.025 | -0.4833 | No | ||
| 86 | HOXA11 | 17146 | 11692 | -0.025 | -0.4836 | No | ||
| 87 | DEPDC2 | 14295 | 11860 | -0.027 | -0.4924 | No | ||
| 88 | RIN2 | 14817 | 11866 | -0.028 | -0.4925 | No | ||
| 89 | CSF3 | 1394 20671 | 11910 | -0.028 | -0.4946 | No | ||
| 90 | UBXD3 | 15701 | 11966 | -0.029 | -0.4973 | No | ||
| 91 | FLRT1 | 11852 | 12175 | -0.034 | -0.5083 | No | ||
| 92 | TSHB | 15218 | 12384 | -0.038 | -0.5193 | No | ||
| 93 | ESRRG | 6447 | 12406 | -0.039 | -0.5201 | No | ||
| 94 | CALCRL | 12041 | 12419 | -0.039 | -0.5205 | No | ||
| 95 | PIK3R1 | 3170 | 12502 | -0.041 | -0.5246 | No | ||
| 96 | LEMD1 | 10023 537 | 12652 | -0.046 | -0.5323 | No | ||
| 97 | LMNA | 4999 15290 1778 9280 | 12737 | -0.047 | -0.5365 | No | ||
| 98 | POU4F2 | 9603 5276 | 13022 | -0.055 | -0.5515 | No | ||
| 99 | TBX2 | 20720 | 13236 | -0.062 | -0.5625 | No | ||
| 100 | OFCC1 | 21486 | 13329 | -0.066 | -0.5670 | No | ||
| 101 | LRFN5 | 6213 | 13446 | -0.070 | -0.5727 | No | ||
| 102 | AP1S2 | 8423 4228 | 13489 | -0.072 | -0.5744 | No | ||
| 103 | POU3F3 | 14261 | 13516 | -0.073 | -0.5753 | No | ||
| 104 | MAPK10 | 11169 | 13581 | -0.077 | -0.5781 | No | ||
| 105 | ACRV1 | 19521 | 13650 | -0.081 | -0.5812 | No | ||
| 106 | A2BP1 | 11212 1652 1689 | 13688 | -0.083 | -0.5826 | No | ||
| 107 | FOXA2 | 14405 2889 | 13717 | -0.084 | -0.5834 | No | ||
| 108 | HAPLN2 | 15296 | 13814 | -0.089 | -0.5879 | No | ||
| 109 | ZBTB10 | 10468 | 13893 | -0.094 | -0.5914 | No | ||
| 110 | RCOR1 | 23998 | 13998 | -0.101 | -0.5963 | No | ||
| 111 | RHOQ | 23142 | 14100 | -0.110 | -0.6009 | No | ||
| 112 | TNNC1 | 22058 | 14213 | -0.120 | -0.6060 | No | ||
| 113 | NOSIP | 18254 3847 | 14319 | -0.132 | -0.6107 | No | ||
| 114 | BNC2 | 15850 | 14348 | -0.135 | -0.6111 | No | ||
| 115 | AXL | 17922 | 14463 | -0.148 | -0.6161 | No | ||
| 116 | TLE4 | 23720 3697 | 14521 | -0.154 | -0.6180 | No | ||
| 117 | SFXN5 | 17090 | 14566 | -0.160 | -0.6192 | No | ||
| 118 | DMD | 24295 2647 | 14621 | -0.170 | -0.6208 | No | ||
| 119 | FBXL19 | 6134 | 14655 | -0.175 | -0.6212 | No | ||
| 120 | DMTF1 | 887 3662 16928 3530 | 14878 | -0.214 | -0.6315 | No | ||
| 121 | POU2F1 | 5275 3989 4065 4010 | 15082 | -0.259 | -0.6405 | No | ||
| 122 | ELF4 | 24162 | 15221 | -0.294 | -0.6457 | No | ||
| 123 | KIF1B | 15671 2476 4950 15673 | 15370 | -0.341 | -0.6511 | No | ||
| 124 | POLK | 21384 | 15425 | -0.362 | -0.6512 | Yes | ||
| 125 | HIVEP3 | 4973 | 15475 | -0.379 | -0.6508 | Yes | ||
| 126 | AP1G1 | 4392 | 15515 | -0.404 | -0.6498 | Yes | ||
| 127 | HDAC9 | 21084 2131 | 15556 | -0.425 | -0.6486 | Yes | ||
| 128 | P2RY10 | 2613 24269 | 15601 | -0.442 | -0.6476 | Yes | ||
| 129 | PARD6A | 12142 | 15662 | -0.471 | -0.6471 | Yes | ||
| 130 | PUM2 | 2071 8161 | 15663 | -0.471 | -0.6435 | Yes | ||
| 131 | PLS3 | 8332 | 15682 | -0.480 | -0.6407 | Yes | ||
| 132 | TNKS1BP1 | 14970 | 15831 | -0.576 | -0.6442 | Yes | ||
| 133 | MAP4K4 | 11241 | 15849 | -0.588 | -0.6405 | Yes | ||
| 134 | TGFB3 | 10161 | 15913 | -0.649 | -0.6389 | Yes | ||
| 135 | STAG2 | 5521 | 16122 | -0.850 | -0.6435 | Yes | ||
| 136 | ZADH2 | 23501 | 16127 | -0.854 | -0.6370 | Yes | ||
| 137 | TAPBP | 5625 | 16326 | -1.019 | -0.6398 | Yes | ||
| 138 | H2AFY | 21447 | 16415 | -1.119 | -0.6358 | Yes | ||
| 139 | CDKL5 | 24019 | 16438 | -1.139 | -0.6281 | Yes | ||
| 140 | VAMP3 | 15660 | 16482 | -1.197 | -0.6211 | Yes | ||
| 141 | PRKAG1 | 22135 | 16486 | -1.198 | -0.6119 | Yes | ||
| 142 | MTCH2 | 14954 7120 | 16765 | -1.602 | -0.6144 | Yes | ||
| 143 | RAB3IP | 19623 | 16842 | -1.714 | -0.6052 | Yes | ||
| 144 | CASP8 | 8694 | 17013 | -1.927 | -0.5993 | Yes | ||
| 145 | H3F3B | 4839 | 17030 | -1.944 | -0.5850 | Yes | ||
| 146 | NCOR1 | 5402 | 17034 | -1.951 | -0.5699 | Yes | ||
| 147 | MAP2K6 | 20614 1414 | 17087 | -2.043 | -0.5567 | Yes | ||
| 148 | CTCF | 18490 | 17307 | -2.434 | -0.5496 | Yes | ||
| 149 | LUC7L | 47 1520 | 17310 | -2.444 | -0.5306 | Yes | ||
| 150 | PRKACB | 15140 | 17626 | -3.163 | -0.5229 | Yes | ||
| 151 | DUSP10 | 4003 14016 | 17669 | -3.295 | -0.4995 | Yes | ||
| 152 | MEIS1 | 20524 | 17695 | -3.354 | -0.4746 | Yes | ||
| 153 | SCML4 | 20042 | 17854 | -3.739 | -0.4539 | Yes | ||
| 154 | BCL11A | 4691 | 18039 | -4.409 | -0.4295 | Yes | ||
| 155 | MBNL1 | 1921 15582 | 18100 | -4.631 | -0.3965 | Yes | ||
| 156 | CHD2 | 10847 | 18237 | -5.235 | -0.3630 | Yes | ||
| 157 | MBNL2 | 21932 | 18284 | -5.431 | -0.3230 | Yes | ||
| 158 | HHEX | 23872 | 18391 | -6.087 | -0.2812 | Yes | ||
| 159 | LMO4 | 15151 | 18404 | -6.210 | -0.2333 | Yes | ||
| 160 | LRRN3 | 21077 | 18495 | -7.254 | -0.1815 | Yes | ||
| 161 | SOX4 | 5479 | 18506 | -7.403 | -0.1241 | Yes | ||
| 162 | ZFP36L1 | 4453 4454 | 18544 | -7.886 | -0.0645 | Yes | ||
| 163 | BCL6 | 22624 | 18580 | -8.747 | 0.0020 | Yes |

