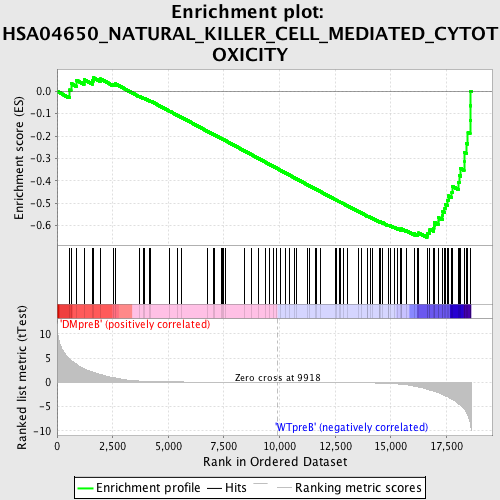
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | HSA04650_NATURAL_KILLER_CELL_MEDIATED_CYTOTOXICITY |
| Enrichment Score (ES) | -0.65362924 |
| Normalized Enrichment Score (NES) | -1.6295093 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.034952547 |
| FWER p-Value | 0.305 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CD48 | 14051 | 548 | 4.836 | 0.0060 | No | ||
| 2 | PIK3CB | 19030 | 626 | 4.520 | 0.0352 | No | ||
| 3 | HCST | 6237 | 866 | 3.779 | 0.0501 | No | ||
| 4 | PIK3R3 | 5248 | 1211 | 2.839 | 0.0524 | No | ||
| 5 | CASP3 | 8693 | 1584 | 2.145 | 0.0481 | No | ||
| 6 | FCER1G | 13759 | 1625 | 2.092 | 0.0614 | No | ||
| 7 | BID | 17017 435 | 1954 | 1.590 | 0.0554 | No | ||
| 8 | PPP3CA | 1863 5284 | 2520 | 0.993 | 0.0322 | No | ||
| 9 | HRAS | 4868 | 2610 | 0.906 | 0.0341 | No | ||
| 10 | VAV3 | 1774 1848 15443 | 3695 | 0.275 | -0.0224 | No | ||
| 11 | PIK3R2 | 18850 | 3888 | 0.232 | -0.0311 | No | ||
| 12 | SHC1 | 9813 9812 5430 | 3936 | 0.225 | -0.0320 | No | ||
| 13 | IFNA1 | 9147 | 4153 | 0.191 | -0.0422 | No | ||
| 14 | TNFRSF10B | 21971 10205 5781 10204 | 4210 | 0.184 | -0.0439 | No | ||
| 15 | FYN | 3375 3395 20052 | 5072 | 0.109 | -0.0896 | No | ||
| 16 | TNFSF10 | 15626 | 5420 | 0.090 | -0.1077 | No | ||
| 17 | IFNA7 | 9152 | 5600 | 0.082 | -0.1167 | No | ||
| 18 | NRAS | 5191 | 6779 | 0.047 | -0.1800 | No | ||
| 19 | IFNA14 | 11917 | 7024 | 0.042 | -0.1929 | No | ||
| 20 | IFNA6 | 17749 | 7067 | 0.041 | -0.1948 | No | ||
| 21 | IFNB1 | 15846 | 7388 | 0.035 | -0.2118 | No | ||
| 22 | KIR3DL1 | 2611 6365 | 7433 | 0.034 | -0.2140 | No | ||
| 23 | IFNA2 | 9149 | 7465 | 0.033 | -0.2154 | No | ||
| 24 | PAK1 | 9527 | 7590 | 0.031 | -0.2219 | No | ||
| 25 | PIK3R5 | 11507 | 8444 | 0.019 | -0.2678 | No | ||
| 26 | SHC3 | 21465 | 8738 | 0.015 | -0.2835 | No | ||
| 27 | CD247 | 4498 8715 | 9036 | 0.011 | -0.2994 | No | ||
| 28 | PTPN11 | 5326 16391 9660 | 9360 | 0.007 | -0.3168 | No | ||
| 29 | KLRC1 | 9242 | 9547 | 0.005 | -0.3268 | No | ||
| 30 | PRF1 | 20011 | 9734 | 0.002 | -0.3369 | No | ||
| 31 | PPP3R2 | 9612 | 9858 | 0.001 | -0.3435 | No | ||
| 32 | SH2D1A | 9810 | 10059 | -0.002 | -0.3543 | No | ||
| 33 | NCR1 | 18409 | 10246 | -0.004 | -0.3643 | No | ||
| 34 | SH2D1B | 14061 | 10433 | -0.006 | -0.3743 | No | ||
| 35 | RAC3 | 20561 | 10656 | -0.009 | -0.3862 | No | ||
| 36 | NFATC4 | 22002 | 10757 | -0.011 | -0.3915 | No | ||
| 37 | IFNA13 | 15844 | 11243 | -0.018 | -0.4176 | No | ||
| 38 | KLRC3 | 16975 | 11351 | -0.019 | -0.4232 | No | ||
| 39 | IFNA4 | 9150 | 11609 | -0.023 | -0.4369 | No | ||
| 40 | PPP3R1 | 9611 489 | 11661 | -0.024 | -0.4395 | No | ||
| 41 | NFATC2 | 5168 2866 | 11816 | -0.027 | -0.4476 | No | ||
| 42 | PIK3R1 | 3170 | 12502 | -0.041 | -0.4843 | No | ||
| 43 | GZMB | 21810 | 12565 | -0.043 | -0.4873 | No | ||
| 44 | PLCG1 | 14753 | 12675 | -0.046 | -0.4929 | No | ||
| 45 | CSF2 | 20454 | 12732 | -0.047 | -0.4956 | No | ||
| 46 | IFNAR1 | 22703 | 12857 | -0.050 | -0.5019 | No | ||
| 47 | SOS1 | 5476 | 13046 | -0.056 | -0.5116 | No | ||
| 48 | PIK3CA | 9562 | 13540 | -0.075 | -0.5377 | No | ||
| 49 | TNF | 23004 | 13668 | -0.082 | -0.5439 | No | ||
| 50 | KLRC2 | 9243 | 13960 | -0.099 | -0.5589 | No | ||
| 51 | NFAT5 | 3921 7037 12036 | 13961 | -0.099 | -0.5582 | No | ||
| 52 | IFNA5 | 9151 | 14080 | -0.109 | -0.5638 | No | ||
| 53 | LAT | 17643 | 14176 | -0.117 | -0.5680 | No | ||
| 54 | LCP2 | 4988 9268 | 14508 | -0.152 | -0.5848 | No | ||
| 55 | FASLG | 13789 | 14511 | -0.153 | -0.5838 | No | ||
| 56 | ZAP70 | 14271 4042 | 14538 | -0.156 | -0.5840 | No | ||
| 57 | KLRK1 | 6544 16976 11281 | 14641 | -0.173 | -0.5883 | No | ||
| 58 | FCGR3A | 8960 | 14882 | -0.215 | -0.5996 | No | ||
| 59 | MAP2K2 | 19933 | 14908 | -0.221 | -0.5994 | No | ||
| 60 | VAV2 | 5848 2670 | 14978 | -0.236 | -0.6013 | No | ||
| 61 | IFNAR2 | 22705 1699 | 15148 | -0.277 | -0.6084 | No | ||
| 62 | MAP2K1 | 19082 | 15318 | -0.325 | -0.6152 | No | ||
| 63 | LCK | 15746 | 15440 | -0.368 | -0.6190 | No | ||
| 64 | ITGAL | 9187 | 15467 | -0.377 | -0.6176 | No | ||
| 65 | IFNG | 19869 | 15478 | -0.380 | -0.6154 | No | ||
| 66 | FAS | 23881 3750 | 15710 | -0.499 | -0.6242 | No | ||
| 67 | ARAF | 24367 | 16058 | -0.778 | -0.6372 | No | ||
| 68 | KLRD1 | 17266 | 16214 | -0.939 | -0.6386 | No | ||
| 69 | TYROBP | 18302 | 16254 | -0.981 | -0.6335 | No | ||
| 70 | IFNGR1 | 20083 | 16628 | -1.403 | -0.6433 | Yes | ||
| 71 | RAC1 | 16302 | 16651 | -1.434 | -0.6339 | Yes | ||
| 72 | PIK3CD | 9563 | 16730 | -1.548 | -0.6267 | Yes | ||
| 73 | ITGB2 | 19978 | 16756 | -1.593 | -0.6163 | Yes | ||
| 74 | MAPK3 | 6458 11170 | 16931 | -1.809 | -0.6124 | Yes | ||
| 75 | PPP3CB | 5285 | 16941 | -1.823 | -0.5995 | Yes | ||
| 76 | PPP3CC | 21763 | 16961 | -1.850 | -0.5869 | Yes | ||
| 77 | ICAM1 | 19545 | 17130 | -2.122 | -0.5803 | Yes | ||
| 78 | RAF1 | 17035 | 17131 | -2.125 | -0.5647 | Yes | ||
| 79 | SOS2 | 21049 | 17328 | -2.499 | -0.5569 | Yes | ||
| 80 | PRKCA | 20174 | 17337 | -2.525 | -0.5387 | Yes | ||
| 81 | IFNGR2 | 9155 22702 | 17428 | -2.696 | -0.5237 | Yes | ||
| 82 | NFATC1 | 23398 1999 5167 9455 1985 1957 | 17449 | -2.738 | -0.5046 | Yes | ||
| 83 | SH3BP2 | 16874 | 17547 | -2.956 | -0.4881 | Yes | ||
| 84 | GRB2 | 20149 | 17590 | -3.055 | -0.4678 | Yes | ||
| 85 | PLCG2 | 18453 | 17743 | -3.480 | -0.4504 | Yes | ||
| 86 | NFATC3 | 5169 | 17772 | -3.544 | -0.4258 | Yes | ||
| 87 | ICAM2 | 20183 | 18035 | -4.385 | -0.4077 | Yes | ||
| 88 | SYK | 21636 | 18074 | -4.548 | -0.3762 | Yes | ||
| 89 | VAV1 | 23173 | 18136 | -4.751 | -0.3445 | Yes | ||
| 90 | PRKCB1 | 1693 9574 | 18292 | -5.486 | -0.3125 | Yes | ||
| 91 | PTK2B | 21776 | 18298 | -5.513 | -0.2722 | Yes | ||
| 92 | KRAS | 9247 | 18396 | -6.150 | -0.2321 | Yes | ||
| 93 | RAC2 | 22217 | 18467 | -6.932 | -0.1848 | Yes | ||
| 94 | PTPN6 | 17002 | 18561 | -8.216 | -0.1293 | Yes | ||
| 95 | PIK3CG | 6635 | 18583 | -8.846 | -0.0653 | Yes | ||
| 96 | MAPK1 | 1642 11167 | 18592 | -9.104 | 0.0013 | Yes |

