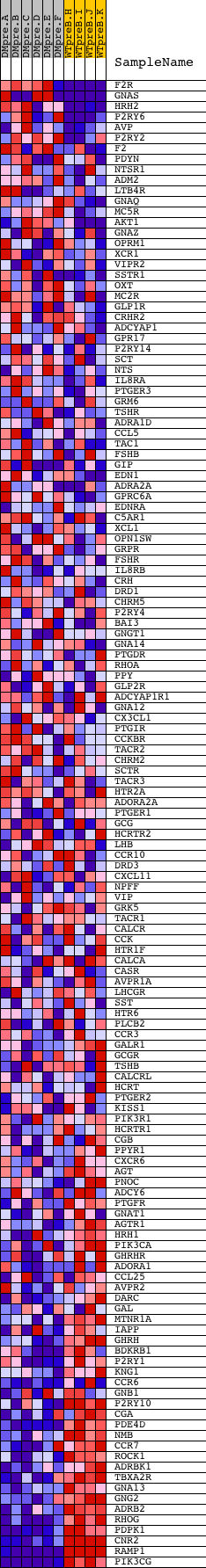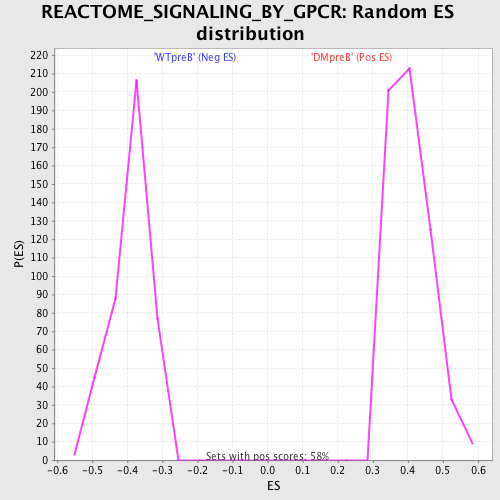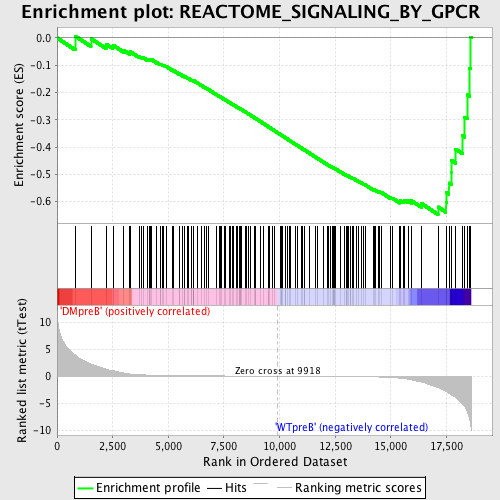
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | REACTOME_SIGNALING_BY_GPCR |
| Enrichment Score (ES) | -0.64731777 |
| Normalized Enrichment Score (NES) | -1.666453 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.015890632 |
| FWER p-Value | 0.21 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | F2R | 21386 | 815 | 3.892 | 0.0066 | No | ||
| 2 | GNAS | 9025 2963 2752 | 1524 | 2.229 | -0.0027 | No | ||
| 3 | HRH2 | 9120 | 2222 | 1.281 | -0.0238 | No | ||
| 4 | P2RY6 | 10595 | 2512 | 0.996 | -0.0265 | No | ||
| 5 | AVP | 14428 | 2994 | 0.560 | -0.0452 | No | ||
| 6 | P2RY2 | 17736 1279 | 3269 | 0.406 | -0.0547 | No | ||
| 7 | F2 | 14524 | 3282 | 0.398 | -0.0502 | No | ||
| 8 | PDYN | 14429 | 3709 | 0.271 | -0.0697 | No | ||
| 9 | NTSR1 | 14710 | 3786 | 0.253 | -0.0706 | No | ||
| 10 | ADM2 | 22388 | 3894 | 0.231 | -0.0733 | No | ||
| 11 | LTB4R | 22003 | 4079 | 0.203 | -0.0807 | No | ||
| 12 | GNAQ | 4786 23909 3685 | 4136 | 0.194 | -0.0812 | No | ||
| 13 | MC5R | 23524 | 4171 | 0.189 | -0.0805 | No | ||
| 14 | AKT1 | 8568 | 4218 | 0.183 | -0.0806 | No | ||
| 15 | GNAZ | 71 | 4228 | 0.182 | -0.0788 | No | ||
| 16 | OPRM1 | 5212 | 4483 | 0.153 | -0.0905 | No | ||
| 17 | XCR1 | 18953 | 4639 | 0.137 | -0.0971 | No | ||
| 18 | VIPR2 | 21125 | 4659 | 0.135 | -0.0964 | No | ||
| 19 | SSTR1 | 21264 | 4753 | 0.128 | -0.0998 | No | ||
| 20 | OXT | 14843 | 4787 | 0.126 | -0.0999 | No | ||
| 21 | MC2R | 23412 | 4905 | 0.118 | -0.1047 | No | ||
| 22 | GLP1R | 23304 | 5165 | 0.103 | -0.1174 | No | ||
| 23 | CRHR2 | 17140 | 5244 | 0.099 | -0.1203 | No | ||
| 24 | ADCYAP1 | 4348 | 5497 | 0.086 | -0.1328 | No | ||
| 25 | GPR17 | 399 | 5502 | 0.086 | -0.1319 | No | ||
| 26 | P2RY14 | 8947 | 5655 | 0.079 | -0.1391 | No | ||
| 27 | SCT | 17560 | 5737 | 0.075 | -0.1425 | No | ||
| 28 | NTS | 19635 3297 | 5746 | 0.075 | -0.1420 | No | ||
| 29 | IL8RA | 13925 | 5863 | 0.071 | -0.1473 | No | ||
| 30 | PTGER3 | 15384 1803 | 5910 | 0.069 | -0.1489 | No | ||
| 31 | GRM6 | 45 | 6038 | 0.065 | -0.1549 | No | ||
| 32 | TSHR | 5801 10223 | 6062 | 0.064 | -0.1554 | No | ||
| 33 | ADRA1D | 20737 8561 | 6115 | 0.063 | -0.1573 | No | ||
| 34 | CCL5 | 20320 | 6118 | 0.063 | -0.1566 | No | ||
| 35 | TAC1 | 17526 19853 | 6150 | 0.062 | -0.1575 | No | ||
| 36 | FSHB | 14500 | 6302 | 0.058 | -0.1649 | No | ||
| 37 | GIP | 76 | 6476 | 0.054 | -0.1736 | No | ||
| 38 | EDN1 | 21658 | 6638 | 0.050 | -0.1817 | No | ||
| 39 | ADRA2A | 4355 | 6721 | 0.048 | -0.1855 | No | ||
| 40 | GPRC6A | 19773 | 6820 | 0.046 | -0.1902 | No | ||
| 41 | EDNRA | 18834 | 7164 | 0.039 | -0.2082 | No | ||
| 42 | C5AR1 | 17957 | 7290 | 0.037 | -0.2145 | No | ||
| 43 | XCL1 | 13781 | 7333 | 0.036 | -0.2163 | No | ||
| 44 | OPN1SW | 17196 | 7381 | 0.035 | -0.2184 | No | ||
| 45 | GRPR | 24015 | 7503 | 0.033 | -0.2245 | No | ||
| 46 | FSHR | 1492 22876 | 7567 | 0.032 | -0.2275 | No | ||
| 47 | IL8RB | 8752 4533 | 7743 | 0.029 | -0.2366 | No | ||
| 48 | CRH | 8784 | 7793 | 0.028 | -0.2389 | No | ||
| 49 | DRD1 | 4638 | 7875 | 0.027 | -0.2430 | No | ||
| 50 | CHRM5 | 10038 | 7936 | 0.026 | -0.2459 | No | ||
| 51 | P2RY4 | 24100 | 8083 | 0.024 | -0.2535 | No | ||
| 52 | BAI3 | 5580 | 8112 | 0.023 | -0.2547 | No | ||
| 53 | GNGT1 | 17533 | 8192 | 0.022 | -0.2587 | No | ||
| 54 | GNA14 | 23907 | 8214 | 0.022 | -0.2595 | No | ||
| 55 | PTGDR | 21866 | 8235 | 0.021 | -0.2603 | No | ||
| 56 | RHOA | 8624 4409 4410 | 8261 | 0.021 | -0.2614 | No | ||
| 57 | PPY | 20208 | 8279 | 0.021 | -0.2621 | No | ||
| 58 | GLP2R | 8227 13563 13564 | 8447 | 0.018 | -0.2708 | No | ||
| 59 | ADCYAP1R1 | 8553 1192 4349 | 8508 | 0.018 | -0.2739 | No | ||
| 60 | GNA12 | 9023 | 8597 | 0.016 | -0.2784 | No | ||
| 61 | CX3CL1 | 9791 | 8684 | 0.015 | -0.2829 | No | ||
| 62 | PTGIR | 18376 | 8862 | 0.013 | -0.2923 | No | ||
| 63 | CCKBR | 8706 | 8917 | 0.013 | -0.2950 | No | ||
| 64 | TACR2 | 20004 | 8919 | 0.013 | -0.2949 | No | ||
| 65 | CHRM2 | 10840 | 9128 | 0.010 | -0.3060 | No | ||
| 66 | SCTR | 14157 | 9272 | 0.008 | -0.3137 | No | ||
| 67 | TACR3 | 15415 | 9492 | 0.005 | -0.3255 | No | ||
| 68 | HTR2A | 9135 21961 | 9500 | 0.005 | -0.3258 | No | ||
| 69 | ADORA2A | 8557 | 9558 | 0.004 | -0.3288 | No | ||
| 70 | PTGER1 | 5309 | 9676 | 0.003 | -0.3351 | No | ||
| 71 | GCG | 14578 | 9789 | 0.002 | -0.3411 | No | ||
| 72 | HCRTR2 | 6907 | 10050 | -0.002 | -0.3552 | No | ||
| 73 | LHB | 4993 | 10102 | -0.002 | -0.3579 | No | ||
| 74 | CCR10 | 20218 | 10107 | -0.002 | -0.3581 | No | ||
| 75 | DRD3 | 22750 | 10108 | -0.002 | -0.3581 | No | ||
| 76 | CXCL11 | 16478 | 10254 | -0.004 | -0.3659 | No | ||
| 77 | NPFF | 12046 | 10264 | -0.004 | -0.3663 | No | ||
| 78 | VIP | 20096 | 10374 | -0.005 | -0.3721 | No | ||
| 79 | GRK5 | 9036 23814 | 10447 | -0.006 | -0.3759 | No | ||
| 80 | TACR1 | 1082 17403 | 10505 | -0.007 | -0.3789 | No | ||
| 81 | CALCR | 17229 | 10722 | -0.010 | -0.3905 | No | ||
| 82 | CCK | 18964 | 10805 | -0.011 | -0.3948 | No | ||
| 83 | HTR1F | 22562 | 10965 | -0.014 | -0.4032 | No | ||
| 84 | CALCA | 4470 | 11037 | -0.014 | -0.4069 | No | ||
| 85 | CASR | 22602 | 11097 | -0.015 | -0.4099 | No | ||
| 86 | AVPR1A | 19863 | 11139 | -0.016 | -0.4119 | No | ||
| 87 | LHCGR | 9273 4994 | 11327 | -0.019 | -0.4217 | No | ||
| 88 | SST | 22625 | 11626 | -0.024 | -0.4376 | No | ||
| 89 | HTR6 | 15698 | 11722 | -0.025 | -0.4424 | No | ||
| 90 | PLCB2 | 5262 | 11963 | -0.029 | -0.4550 | No | ||
| 91 | CCR3 | 19251 | 12158 | -0.033 | -0.4651 | No | ||
| 92 | GALR1 | 23397 | 12219 | -0.034 | -0.4679 | No | ||
| 93 | GCGR | 20566 | 12267 | -0.036 | -0.4699 | No | ||
| 94 | TSHB | 15218 | 12384 | -0.038 | -0.4757 | No | ||
| 95 | CALCRL | 12041 | 12419 | -0.039 | -0.4770 | No | ||
| 96 | HCRT | 20224 | 12426 | -0.039 | -0.4769 | No | ||
| 97 | PTGER2 | 22039 | 12464 | -0.040 | -0.4783 | No | ||
| 98 | KISS1 | 6603 | 12482 | -0.041 | -0.4787 | No | ||
| 99 | PIK3R1 | 3170 | 12502 | -0.041 | -0.4792 | No | ||
| 100 | HCRTR1 | 15741 | 12727 | -0.047 | -0.4907 | No | ||
| 101 | CGB | 8253 | 12917 | -0.052 | -0.5003 | No | ||
| 102 | PPYR1 | 21881 | 12986 | -0.054 | -0.5033 | No | ||
| 103 | CXCR6 | 19252 | 12992 | -0.054 | -0.5028 | No | ||
| 104 | AGT | 18711 | 13040 | -0.056 | -0.5046 | No | ||
| 105 | PNOC | 21783 | 13078 | -0.057 | -0.5059 | No | ||
| 106 | ADCY6 | 22139 2283 8551 | 13201 | -0.061 | -0.5117 | No | ||
| 107 | PTGFR | 15138 | 13203 | -0.061 | -0.5110 | No | ||
| 108 | GNAT1 | 3027 19000 | 13275 | -0.064 | -0.5140 | No | ||
| 109 | AGTR1 | 21681 15369 | 13333 | -0.066 | -0.5162 | No | ||
| 110 | HRH1 | 17321 | 13464 | -0.071 | -0.5223 | No | ||
| 111 | PIK3CA | 9562 | 13540 | -0.075 | -0.5254 | No | ||
| 112 | GHRHR | 1007 17436 | 13660 | -0.081 | -0.5308 | No | ||
| 113 | ADORA1 | 13831 | 13775 | -0.088 | -0.5358 | No | ||
| 114 | CCL25 | 9789 3848 | 13851 | -0.092 | -0.5387 | No | ||
| 115 | AVPR2 | 8646 | 14223 | -0.121 | -0.5572 | No | ||
| 116 | DARC | 13747 | 14252 | -0.124 | -0.5571 | No | ||
| 117 | GAL | 23946 | 14313 | -0.131 | -0.5586 | No | ||
| 118 | MTNR1A | 18626 | 14456 | -0.147 | -0.5644 | No | ||
| 119 | IAPP | 17249 | 14482 | -0.150 | -0.5638 | No | ||
| 120 | GHRH | 14372 | 14572 | -0.160 | -0.5665 | No | ||
| 121 | BDKRB1 | 19418 | 14984 | -0.237 | -0.5857 | No | ||
| 122 | P2RY1 | 15581 | 15071 | -0.254 | -0.5870 | No | ||
| 123 | KNG1 | 9244 22809 | 15410 | -0.355 | -0.6007 | No | ||
| 124 | CCR6 | 1505 23390 | 15424 | -0.361 | -0.5967 | No | ||
| 125 | GNB1 | 15967 | 15565 | -0.428 | -0.5987 | No | ||
| 126 | P2RY10 | 2613 24269 | 15601 | -0.442 | -0.5949 | No | ||
| 127 | CGA | 16246 | 15780 | -0.538 | -0.5975 | No | ||
| 128 | PDE4D | 10722 6235 | 15934 | -0.668 | -0.5971 | No | ||
| 129 | NMB | 12628 | 16373 | -1.075 | -0.6068 | No | ||
| 130 | CCR7 | 20254 | 17123 | -2.114 | -0.6198 | Yes | ||
| 131 | ROCK1 | 5386 | 17480 | -2.824 | -0.6023 | Yes | ||
| 132 | ADRBK1 | 23960 | 17490 | -2.840 | -0.5658 | Yes | ||
| 133 | TBXA2R | 3392 19928 | 17614 | -3.119 | -0.5319 | Yes | ||
| 134 | GNA13 | 20617 | 17715 | -3.402 | -0.4930 | Yes | ||
| 135 | GNG2 | 4790 | 17719 | -3.419 | -0.4486 | Yes | ||
| 136 | ADRB2 | 23422 | 17922 | -3.954 | -0.4081 | Yes | ||
| 137 | RHOG | 17727 | 18209 | -5.063 | -0.3576 | Yes | ||
| 138 | PDPK1 | 23097 | 18324 | -5.661 | -0.2901 | Yes | ||
| 139 | CNR2 | 16035 | 18457 | -6.769 | -0.2091 | Yes | ||
| 140 | RAMP1 | 14189 | 18542 | -7.870 | -0.1112 | Yes | ||
| 141 | PIK3CG | 6635 | 18583 | -8.846 | 0.0018 | Yes |