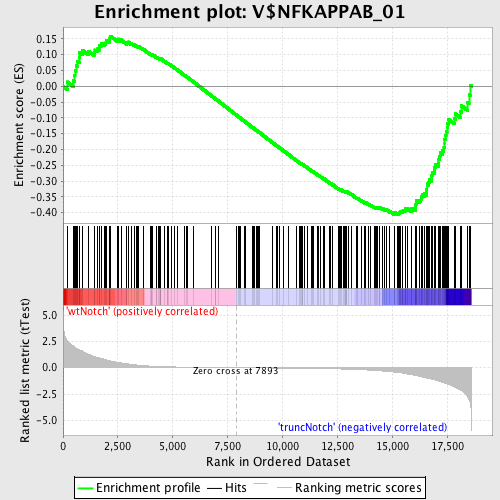
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch_repos |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#wtNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | V$NFKAPPAB_01 |
| Enrichment Score (ES) | -0.4057877 |
| Normalized Enrichment Score (NES) | -1.1979016 |
| Nominal p-value | 0.08203678 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SIN3A | 2190121 | 191 | 2.561 | 0.0138 | No | ||
| 2 | HCFC1 | 2260008 | 463 | 2.041 | 0.0183 | No | ||
| 3 | MSN | 4060044 | 514 | 1.972 | 0.0342 | No | ||
| 4 | UBE2D3 | 3190452 | 542 | 1.915 | 0.0507 | No | ||
| 5 | RAP2C | 1690132 | 588 | 1.868 | 0.0659 | No | ||
| 6 | ARPC2 | 6020484 | 662 | 1.788 | 0.0788 | No | ||
| 7 | UACA | 1190725 | 742 | 1.708 | 0.0906 | No | ||
| 8 | SDC4 | 6370411 | 753 | 1.694 | 0.1060 | No | ||
| 9 | ATP1B1 | 3130594 | 875 | 1.572 | 0.1143 | No | ||
| 10 | CYLD | 6590079 | 1155 | 1.276 | 0.1111 | No | ||
| 11 | BCL3 | 3990440 | 1412 | 1.061 | 0.1073 | No | ||
| 12 | ICAM1 | 6980138 | 1452 | 1.039 | 0.1149 | No | ||
| 13 | PCBP4 | 6350722 | 1563 | 0.991 | 0.1183 | No | ||
| 14 | CDC14A | 4050132 | 1639 | 0.950 | 0.1232 | No | ||
| 15 | RBMS1 | 6400014 | 1665 | 0.929 | 0.1306 | No | ||
| 16 | UBE4B | 780008 3610154 | 1729 | 0.888 | 0.1355 | No | ||
| 17 | NFAT5 | 2510411 5890195 6550152 | 1871 | 0.804 | 0.1355 | No | ||
| 18 | RALGDS | 430463 | 1952 | 0.757 | 0.1382 | No | ||
| 19 | P2RY10 | 6370039 6840204 | 1972 | 0.740 | 0.1442 | No | ||
| 20 | BMF | 610102 | 2098 | 0.672 | 0.1437 | No | ||
| 21 | SMOC1 | 840039 | 2106 | 0.670 | 0.1497 | No | ||
| 22 | TNF | 6650603 | 2136 | 0.652 | 0.1542 | No | ||
| 23 | SEC14L2 | 2640370 | 2163 | 0.638 | 0.1588 | No | ||
| 24 | EGF | 5220154 | 2474 | 0.515 | 0.1469 | No | ||
| 25 | USP52 | 3710215 6380133 | 2509 | 0.502 | 0.1498 | No | ||
| 26 | GNB1 | 2120397 | 2641 | 0.459 | 0.1470 | No | ||
| 27 | CREB1 | 1500717 2230358 3610600 6550601 | 2898 | 0.374 | 0.1366 | No | ||
| 28 | TCEA2 | 6370546 | 2902 | 0.373 | 0.1400 | No | ||
| 29 | SAMSN1 | 2370563 | 2958 | 0.354 | 0.1403 | No | ||
| 30 | FOXJ2 | 2450441 | 3109 | 0.310 | 0.1351 | No | ||
| 31 | SHOX2 | 3190438 6450059 | 3233 | 0.282 | 0.1311 | No | ||
| 32 | BLR1 | 1500300 5420017 | 3341 | 0.256 | 0.1277 | No | ||
| 33 | HIVEP1 | 3850632 | 3407 | 0.245 | 0.1265 | No | ||
| 34 | NLK | 2030010 2450041 | 3455 | 0.233 | 0.1261 | No | ||
| 35 | SESN2 | 2370019 | 3645 | 0.198 | 0.1177 | No | ||
| 36 | ASCL3 | 4570484 | 3989 | 0.149 | 0.1005 | No | ||
| 37 | ERN1 | 2360403 | 4031 | 0.143 | 0.0997 | No | ||
| 38 | JARID2 | 6290538 | 4050 | 0.141 | 0.1000 | No | ||
| 39 | SOX10 | 6200538 | 4094 | 0.137 | 0.0990 | No | ||
| 40 | BIRC3 | 6550411 | 4236 | 0.125 | 0.0925 | No | ||
| 41 | WNT10A | 2100706 | 4269 | 0.122 | 0.0919 | No | ||
| 42 | WNT10B | 510050 | 4346 | 0.114 | 0.0889 | No | ||
| 43 | KLK9 | 4120164 | 4409 | 0.109 | 0.0865 | No | ||
| 44 | TRAF4 | 3060041 4920528 6980286 | 4421 | 0.107 | 0.0869 | No | ||
| 45 | TAL1 | 7040239 | 4446 | 0.106 | 0.0866 | No | ||
| 46 | FGF17 | 3130022 | 4451 | 0.106 | 0.0874 | No | ||
| 47 | CHD4 | 5420059 6130338 6380717 | 4637 | 0.093 | 0.0783 | No | ||
| 48 | UBD | 5570632 | 4753 | 0.086 | 0.0728 | No | ||
| 49 | GNG4 | 2630600 | 4770 | 0.085 | 0.0728 | No | ||
| 50 | GRIN2D | 6620372 | 4791 | 0.083 | 0.0725 | No | ||
| 51 | EDN2 | 6760647 | 4940 | 0.076 | 0.0651 | No | ||
| 52 | KCNN3 | 6520600 6420138 | 5078 | 0.069 | 0.0584 | No | ||
| 53 | CXCL11 | 1090551 | 5210 | 0.063 | 0.0519 | No | ||
| 54 | BAI2 | 380632 630113 6650170 | 5536 | 0.051 | 0.0347 | No | ||
| 55 | NR2F2 | 3170609 3310577 | 5628 | 0.048 | 0.0302 | No | ||
| 56 | UBE2I | 2680056 6350446 | 5662 | 0.047 | 0.0289 | No | ||
| 57 | BDNF | 2940128 3520368 | 5956 | 0.038 | 0.0133 | No | ||
| 58 | GNAO1 | 2510292 6590487 | 6769 | 0.020 | -0.0305 | No | ||
| 59 | SALL1 | 5420020 7050195 | 6955 | 0.016 | -0.0404 | No | ||
| 60 | AP1S2 | 780279 4280707 | 6958 | 0.016 | -0.0404 | No | ||
| 61 | IL23A | 6290044 | 7103 | 0.013 | -0.0481 | No | ||
| 62 | LAMA1 | 1770594 | 7897 | -0.000 | -0.0911 | No | ||
| 63 | IL1RN | 2370333 | 7976 | -0.001 | -0.0953 | No | ||
| 64 | MADCAM1 | 1580010 | 8059 | -0.003 | -0.0997 | No | ||
| 65 | KCNS3 | 4150039 | 8093 | -0.003 | -0.1015 | No | ||
| 66 | MSX1 | 2650309 | 8255 | -0.006 | -0.1102 | No | ||
| 67 | RIN2 | 2450184 | 8323 | -0.007 | -0.1137 | No | ||
| 68 | EHF | 4560088 | 8646 | -0.012 | -0.1311 | No | ||
| 69 | ARHGAP5 | 2510619 3360035 | 8655 | -0.013 | -0.1314 | No | ||
| 70 | CHX10 | 3830184 | 8720 | -0.014 | -0.1347 | No | ||
| 71 | FLRT1 | 4540348 | 8797 | -0.015 | -0.1387 | No | ||
| 72 | CXCL10 | 2450408 | 8852 | -0.016 | -0.1415 | No | ||
| 73 | TLX1 | 2630162 | 8885 | -0.016 | -0.1431 | No | ||
| 74 | SLC6A12 | 3170685 | 8952 | -0.018 | -0.1465 | No | ||
| 75 | FKHL18 | 6370603 | 9560 | -0.028 | -0.1792 | No | ||
| 76 | GREM1 | 3940180 | 9722 | -0.030 | -0.1876 | No | ||
| 77 | HOXA11 | 6980133 | 9753 | -0.031 | -0.1890 | No | ||
| 78 | HOXB9 | 2640687 | 9859 | -0.033 | -0.1943 | No | ||
| 79 | GATA4 | 1410215 2690609 | 10033 | -0.037 | -0.2034 | No | ||
| 80 | FGF1 | 4780435 5670601 | 10272 | -0.041 | -0.2159 | No | ||
| 81 | CXCL5 | 6370333 | 10633 | -0.050 | -0.2350 | No | ||
| 82 | ALG6 | 4060131 | 10768 | -0.053 | -0.2417 | No | ||
| 83 | NTN1 | 5700600 | 10809 | -0.054 | -0.2434 | No | ||
| 84 | IL27 | 1990324 | 10879 | -0.056 | -0.2466 | No | ||
| 85 | BNC2 | 4810603 | 10897 | -0.056 | -0.2470 | No | ||
| 86 | TNFSF15 | 360341 | 10898 | -0.056 | -0.2465 | No | ||
| 87 | HSD11B2 | 5900053 | 10987 | -0.058 | -0.2507 | No | ||
| 88 | ATOH1 | 460278 | 11123 | -0.062 | -0.2574 | No | ||
| 89 | ZBTB11 | 6520092 | 11299 | -0.067 | -0.2663 | No | ||
| 90 | TSLP | 730408 1990500 | 11360 | -0.069 | -0.2689 | No | ||
| 91 | MAP3K8 | 2940286 | 11413 | -0.070 | -0.2711 | No | ||
| 92 | IER5 | 2350575 | 11613 | -0.076 | -0.2812 | No | ||
| 93 | GABRB1 | 6760193 | 11624 | -0.076 | -0.2810 | No | ||
| 94 | FTHL17 | 6550193 | 11733 | -0.080 | -0.2861 | No | ||
| 95 | LTA | 1740088 | 11847 | -0.085 | -0.2914 | No | ||
| 96 | C1QL1 | 2030465 | 11892 | -0.086 | -0.2930 | No | ||
| 97 | SP6 | 60484 510452 2690333 | 12130 | -0.096 | -0.3049 | No | ||
| 98 | PUNC | 2470647 | 12188 | -0.098 | -0.3071 | No | ||
| 99 | IFNB1 | 1400142 | 12299 | -0.104 | -0.3121 | No | ||
| 100 | IL1A | 3610056 | 12568 | -0.117 | -0.3255 | No | ||
| 101 | RIMS2 | 670725 | 12595 | -0.119 | -0.3258 | No | ||
| 102 | BCL6B | 60047 | 12648 | -0.121 | -0.3275 | No | ||
| 103 | MOBKL2C | 2640088 3830017 | 12668 | -0.122 | -0.3274 | No | ||
| 104 | SMPD3 | 520164 | 12801 | -0.130 | -0.3333 | No | ||
| 105 | CPD | 5360403 5860685 | 12814 | -0.130 | -0.3328 | No | ||
| 106 | BMP2K | 60541 5570504 | 12863 | -0.133 | -0.3341 | No | ||
| 107 | CTGF | 4540577 | 12887 | -0.135 | -0.3341 | No | ||
| 108 | KCNH3 | 3130100 | 12895 | -0.135 | -0.3332 | No | ||
| 109 | TNFRSF9 | 2510400 6650484 | 12915 | -0.137 | -0.3329 | No | ||
| 110 | HSD3B7 | 1410129 | 13019 | -0.143 | -0.3372 | No | ||
| 111 | DAP3 | 1660528 2120039 5420593 | 13126 | -0.151 | -0.3415 | No | ||
| 112 | NOL4 | 5050446 5360519 | 13386 | -0.171 | -0.3539 | No | ||
| 113 | FUT7 | 2900736 | 13420 | -0.173 | -0.3541 | No | ||
| 114 | GRK5 | 1940348 4670053 | 13601 | -0.190 | -0.3621 | No | ||
| 115 | CXCL2 | 610398 | 13750 | -0.205 | -0.3682 | No | ||
| 116 | SLC12A2 | 4610019 6040672 | 13803 | -0.210 | -0.3690 | No | ||
| 117 | TNFSF18 | 1990458 2850520 | 13926 | -0.222 | -0.3735 | No | ||
| 118 | IL13 | 7040021 | 14026 | -0.235 | -0.3767 | No | ||
| 119 | CXCL9 | 1570673 | 14178 | -0.253 | -0.3825 | No | ||
| 120 | RND1 | 5080300 | 14247 | -0.264 | -0.3837 | No | ||
| 121 | SOX5 | 2370576 2900167 3190128 5050528 | 14287 | -0.268 | -0.3833 | No | ||
| 122 | COL11A2 | 5130341 | 14340 | -0.274 | -0.3835 | No | ||
| 123 | TRIM47 | 5080717 | 14403 | -0.283 | -0.3842 | No | ||
| 124 | PCSK2 | 360017 430528 5900619 | 14430 | -0.287 | -0.3829 | No | ||
| 125 | SLAMF8 | 1230048 | 14557 | -0.310 | -0.3868 | No | ||
| 126 | ZFP36L2 | 510180 | 14662 | -0.328 | -0.3894 | No | ||
| 127 | PRX | 2260400 4200347 | 14716 | -0.338 | -0.3891 | No | ||
| 128 | DCAMKL1 | 540095 2690092 | 14896 | -0.369 | -0.3953 | No | ||
| 129 | DDR1 | 2060044 5220180 | 15087 | -0.411 | -0.4018 | Yes | ||
| 130 | MAG | 2370037 | 15125 | -0.419 | -0.3998 | Yes | ||
| 131 | TP53 | 6130707 | 15236 | -0.442 | -0.4016 | Yes | ||
| 132 | ACTN3 | 3140541 6480598 | 15308 | -0.461 | -0.4011 | Yes | ||
| 133 | CDK6 | 4920253 | 15335 | -0.469 | -0.3981 | Yes | ||
| 134 | MMP9 | 580338 | 15380 | -0.483 | -0.3960 | Yes | ||
| 135 | TNFRSF1B | 3990035 5860372 | 15464 | -0.510 | -0.3957 | Yes | ||
| 136 | TNIP1 | 2510279 5900041 6510435 | 15490 | -0.519 | -0.3921 | Yes | ||
| 137 | ITGB4 | 1740021 3840482 | 15586 | -0.552 | -0.3921 | Yes | ||
| 138 | GPHN | 3290195 3840121 | 15607 | -0.562 | -0.3879 | Yes | ||
| 139 | PPARBP | 4280131 5690022 5860050 | 15688 | -0.591 | -0.3866 | Yes | ||
| 140 | ASH1L | 2510402 5360687 6650685 | 15870 | -0.652 | -0.3903 | Yes | ||
| 141 | KRTAP13-1 | 2600242 | 15894 | -0.662 | -0.3853 | Yes | ||
| 142 | CD83 | 4150242 | 16053 | -0.739 | -0.3869 | Yes | ||
| 143 | NFKBIB | 2260167 | 16064 | -0.743 | -0.3805 | Yes | ||
| 144 | STC2 | 4920601 | 16076 | -0.749 | -0.3740 | Yes | ||
| 145 | LIG1 | 1980438 4540112 5570609 | 16090 | -0.754 | -0.3676 | Yes | ||
| 146 | GADD45B | 2350408 | 16116 | -0.766 | -0.3617 | Yes | ||
| 147 | ZBTB5 | 770279 | 16254 | -0.827 | -0.3614 | Yes | ||
| 148 | VCAM1 | 2900450 | 16311 | -0.852 | -0.3564 | Yes | ||
| 149 | CLCN1 | 840161 | 16324 | -0.858 | -0.3490 | Yes | ||
| 150 | HCST | 2350603 | 16360 | -0.874 | -0.3426 | Yes | ||
| 151 | EDG5 | 2640300 | 16479 | -0.930 | -0.3403 | Yes | ||
| 152 | ENO3 | 5270136 | 16556 | -0.967 | -0.3353 | Yes | ||
| 153 | COL16A1 | 1780520 | 16565 | -0.973 | -0.3265 | Yes | ||
| 154 | PLAU | 6510047 | 16585 | -0.982 | -0.3183 | Yes | ||
| 155 | TAZ | 7100193 | 16586 | -0.983 | -0.3091 | Yes | ||
| 156 | MSC | 5080139 | 16663 | -1.005 | -0.3037 | Yes | ||
| 157 | LTB | 3990672 | 16691 | -1.016 | -0.2956 | Yes | ||
| 158 | NFKB2 | 2320670 | 16799 | -1.069 | -0.2913 | Yes | ||
| 159 | STAT6 | 1190010 5720019 | 16806 | -1.075 | -0.2815 | Yes | ||
| 160 | RRAS | 4150609 | 16850 | -1.094 | -0.2736 | Yes | ||
| 161 | PFN1 | 6130132 | 16915 | -1.144 | -0.2663 | Yes | ||
| 162 | ABHD8 | 6660082 | 16925 | -1.150 | -0.2559 | Yes | ||
| 163 | EBI3 | 5340300 | 16967 | -1.172 | -0.2471 | Yes | ||
| 164 | JAK3 | 70347 3290008 | 17108 | -1.262 | -0.2428 | Yes | ||
| 165 | RPS6KA4 | 2190524 | 17125 | -1.271 | -0.2317 | Yes | ||
| 166 | AGPAT1 | 610056 | 17152 | -1.298 | -0.2209 | Yes | ||
| 167 | TRIB2 | 4120605 | 17181 | -1.321 | -0.2099 | Yes | ||
| 168 | IRF1 | 2340152 3450592 6290121 6980577 | 17303 | -1.388 | -0.2034 | Yes | ||
| 169 | BCKDK | 1940601 3990600 | 17358 | -1.428 | -0.1929 | Yes | ||
| 170 | BFSP1 | 1410347 | 17384 | -1.450 | -0.1806 | Yes | ||
| 171 | CCL5 | 3710397 | 17387 | -1.453 | -0.1670 | Yes | ||
| 172 | MAP4K2 | 4200037 | 17435 | -1.482 | -0.1556 | Yes | ||
| 173 | UBE2H | 1980142 2970079 | 17455 | -1.491 | -0.1426 | Yes | ||
| 174 | RELB | 1400048 | 17499 | -1.525 | -0.1305 | Yes | ||
| 175 | EIF5A | 1500059 5290022 6370026 | 17518 | -1.536 | -0.1170 | Yes | ||
| 176 | IL4I1 | 3190161 | 17569 | -1.578 | -0.1049 | Yes | ||
| 177 | SLC11A2 | 3140603 | 17828 | -1.824 | -0.1017 | Yes | ||
| 178 | IL27RA | 940093 | 17872 | -1.864 | -0.0865 | Yes | ||
| 179 | FBXL12 | 2230040 3520008 | 18094 | -2.115 | -0.0785 | Yes | ||
| 180 | ETV6 | 610524 | 18156 | -2.182 | -0.0613 | Yes | ||
| 181 | IL2RA | 6620450 | 18426 | -2.744 | -0.0500 | Yes | ||
| 182 | CXCL16 | 510278 | 18502 | -2.989 | -0.0259 | Yes | ||
| 183 | ARHGAP8 | 380333 5390487 | 18563 | -3.405 | 0.0029 | Yes |

